Stem Cell Markers
Contents
- Embryonic Stem Cell Markers
- Hematopoietic Stem Cell Markers
- Mesenchymal/Stromal Stem Cell Markers
- Neural Stem Cell Markers
- References
While stem cells are best defined functionally, a number of molecular markers have been used to characterize various stem cell populations.
Although functions have yet to be ascertained for many of these early markers, their unique expression pattern and timing provide a useful tool for scientists to initially identify as well as isolate stem cells. This mini-review summarizes evidence regarding the roles of specific markers in defining embryonic, hematopoietic, mesenchymal/stromal, and neural stem cell populations. For most of the molecules discussed, studies performed both in vitro and in vivo support their significant role in characterizing stem cells. Until more is known about the novel marker-negative stem cell population, however, uncertainty still exists regarding the benefits of using these markers alone or in various combinations when identifying and isolating cells for stem cell research.
Embryonic Stem Cell Markers
Oct-4: Oct-4 (also termed Oct-3 or Oct-3/4), one of the POU transcription factors, was originally identified as a DNA-binding protein that activates gene transcription via a cis-element containing octamer motif.1 It is expressed in totipotent embryonic stem and germ cells.2, 3 A critical level of Oct-4 expression is required to sustain stem cell self-renewal and pluripotency.4 Differentiation of embryonic stem (ES) cells results in down- regulation of Oct-4, an event essential for a proper and divergent developmental program.5 Oct-4 is not only a master regulator of pluripotency that controls lineage commitment, but is also the first and most recognized marker used for the identification of totipotent ES cells.
SSEAs (Stage Specific Embryonic Antigens): SSEAs were originally identified by three monoclonal antibodies (Abs) recognizing defined carbohydrate epitopes associated with lacto- and globo-series glycolipids, SSEA-1, -3 and - 4.6 SSEA-1 is expressed on the surface of preimplantation-stage murine embryos (i.e. at the eight cell stage) and has been found on the surface of teratocarcinoma stem cells, but not on their differentiated derivatives.7, 8 The oviduct epithelium, endometrium and epididymis, as well as some areas of the brain and kidney tubules in adult mice have also been shown to be reactive with SSEA-1 Abs.9 SSEA-3and SSEA-4 are synthesized during oogenesis and are present in the membranes of oocytes, zygotes and early cleavage-stage embryos.10, 11 Biological roles of these carbohydrate-associated molecules have been suggested in controlling cell surface interactions during development.6 Undifferentiated primate ES cells, human EC and ES cells express SSEA-3 and SSEA-4, but not SSEA-1. Undifferentiated mouse ES cells express SSEA-1, but not SSEA-3 or SSEA-4.12, 13
Hematopoietic Stem Cell Markers
CD34: The cell surface sialomucin, CD34 has been a focus of interest ever since it was found expressed on a small fraction of human bone marrow cells.14 The CD34+-enriched cell population from marrow or mobilized peripheral blood appears responsible for most of the hematopoietic activity.14, 15, 16, 17, 18, 19, 20, 21CD34 has therefore been considered to be the most critical marker for hematopoietic stem cells (HSCs). CD34 expression on primitive cells is down-regulated as they differentiate into mature cells.22 It is also found on clonogenic progenitors, however, and some lineage-committed cells.23 Although its precise function is still unknown, the pattern of expression of CD34 suggests that it plays a significant role in early hematopoiesis.22 The theory of CD34 being the most primitive HSC marker, however, has recently been challenged. Osawa et al. first demonstrated that murine HSCs could be CD34 negative.24 In addition, a low level of engraftment and hematopoietic capacity has been demonstrated in human CD34- cells.25 Transplantation studies also showed repopulating activity in a CD34- cell population in fetal sheep.26 Additionally, studies have shown that both murine and human CD34+ cells may be derived from CD34- cells.27, 28 Collectively, these reports suggest the possibility that HSCs may be CD34+ or CD34- and that selection of cells expressing CD34 might result in exclusion of more primitive stem cells. Nevertheless, almost all clinical and experimental protocols including ex vivo culture, gene therapy, and HSC transplantation are currently designed for cell populations enriched for CD34+ cells.
CD133: CD133, a 120 kDa, glycosylated protein containing five transmembrane domains (Figure 1), was identified initially by the AC133 monoclonal Ab, which recognizes a CD34+ subset of human HSCs.29, 30 A CD133 isoform, AC133-2, has been recently cloned and identified as the original surface antigen recognized by the AC133 Ab.31 CD133 may provide an alternative to CD34 for HSC selection and ex vivo expansion. A CD133+ enriched subset can be expanded in a similar manner as a CD34+ enriched subset, retaining its multilineage capacity.32 Recent studies have offered evidence that CD133 expression is not limited to primitive blood cells, but defines unique cell populations in non-hematopoietic tissues as well. CD133+ progenitor cells from peripheral blood can be induced to differentiate into endothelial cells in vitro.33 In addition, human neural stem cells can be directly isolated by using an anti-CD133 Ab.34
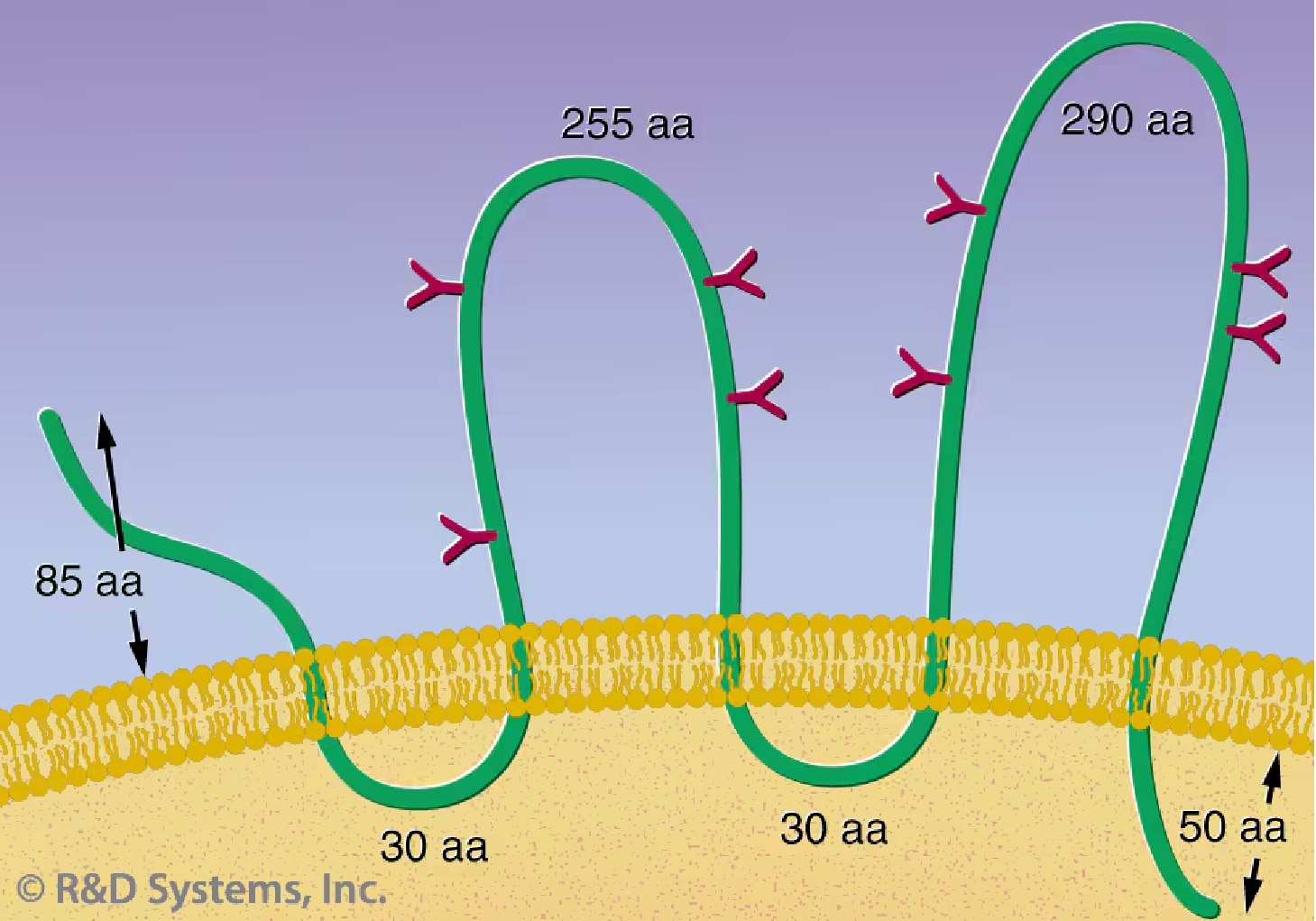
Figure 1. A structure model of CD133 proposed by Miraglia S. et al.30 This protein has an extracellular N-terminus, 5 hydrophobic transmembrane domains, 2 small cytoplasmic loops, 2 large extracellular loops and a cytoplasmic C-termus.
ABCG2: ABCG2 (ATP-binding cassette superfamily G member 2) is a determinant of the Hoechst-negative phenotype of side population (SP) cells and found in a wide variety of stem cells, including HSC.35, 36 ABCG2 is a member of the family of ABC transporters and was first identified in a breast cancer cell line.37 It belongs to the half-transporter group and is unique as it is localized to the plasma membrane (Figure 2).38 The expression of ABCG2 appears greatest on CD34-cells and is down-regulated with the acquisition of CD34 on the cell surface.35 Down-regulation in ABCG2 expression is also observed in various committed hematopoietic progenitors.39 ABCG2 may therefore serve as a more promising marker than CD34 for primitive HSC isolation and characterization. The expression pattern of ABCG2, however, is not limited to HSC. ABCG2 expression exclusively characterizes the Hoechst SP phenotype in cells from diverse sources, including monkey bone marrow, mouse skeletal muscle and ES cells.35 The potential plasticity of SP cells has been demonstrated by studies showing that cardiomyocytes and muscle can be regenerated from transplanted bone marrow-derived SP cells.40, 41 Exclusive expression of ABCG2 on SP cells suggests that ABCG2 may be a potential marker for positive selection of pluripotent stem cells from various adult sources. ABCG2 has been implicated in playing a functional role in developmental stem cell biology (see reference 42 for a review).
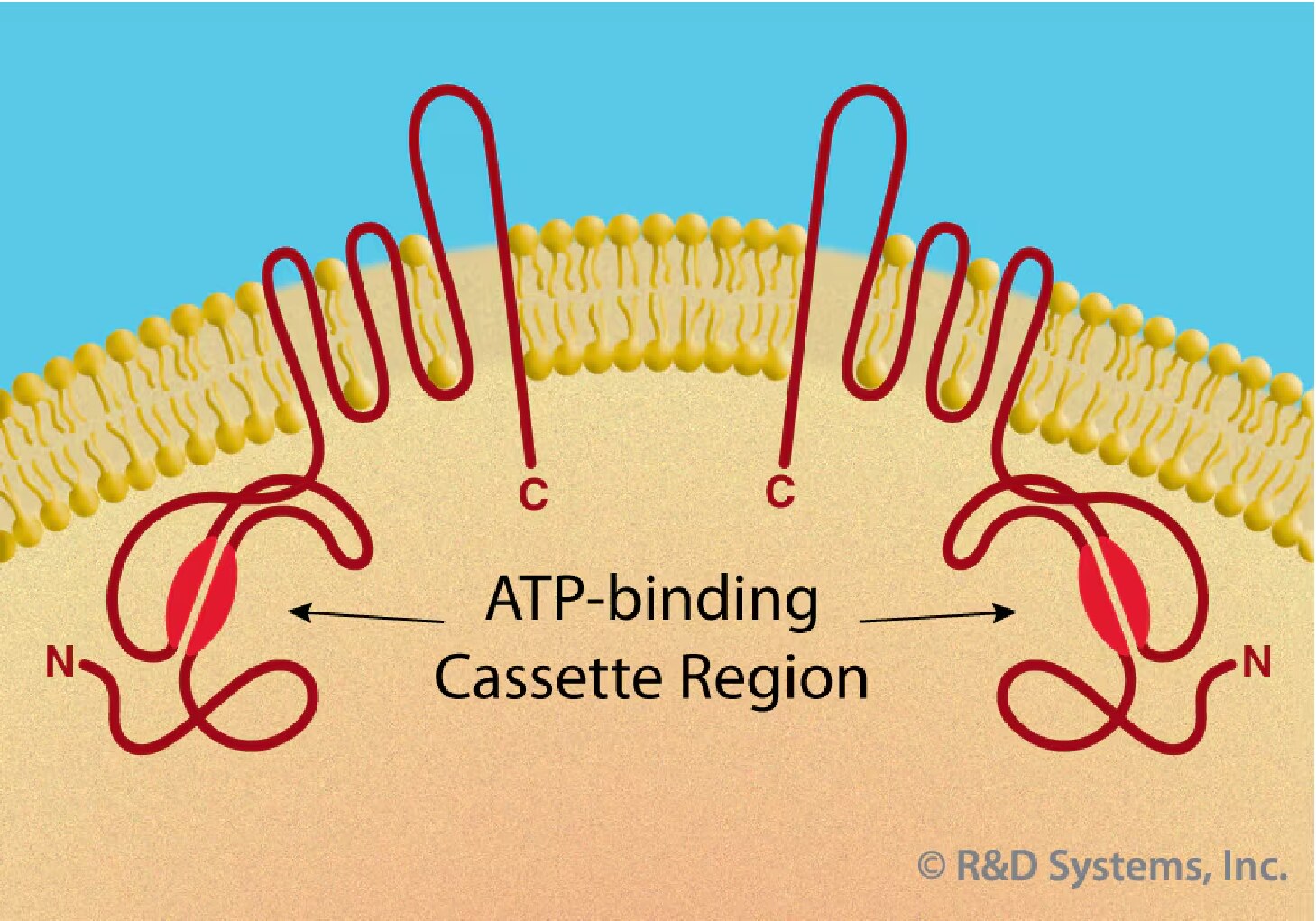
Figure 2. The family of ABC transporters is characterized by the presence of an ATP-binding cassette region, which hydrolyzes ATP to support energy- dependent substrate exportation from the intracellular cytoplasm to the extracellular space. Full-length transporters contain two mirror image halves that are separated by a flexible linker region (not shown). Half-transporters, e.g. ABCG2, function as homo- or heterodimers and may be localized to the plasma membrane.
Sca-1: Sca-1 (stem cell antigen 1, Ly-6A/E), an 18 kDa phosphatidylinositol-anchored protein, is a member of the Ly-6 antigen family.43 Sca-1 is the most recognized HSC marker in mice with both Ly-6 haplotypes as it is expressed on multipotent HSCs.44, 45 An anti-Sca-1 Ab is frequently used in combination with negative selection for expression of a number of cell surface markers characteristic of differentiated cells of hematolymphoid lineages (Lin-) to identify and isolate murine HSCs. Sca-1+ HSCs can be found in the adult bone marrow, fetal liver and mobilized peripheral blood and spleen within the adult animal.44, 45, 46, 47, 48, 49Sca-1 has also been discovered in several non-hematopoietic tissues,43 however, and can be used to enrich progenitor cell populations other than HSCs.50 Sca-1 may be involved in regulating both B and T cell activation.51, 52, 53, 54
Mesenchymal/Stromal Stem Cell Markers
STRO-1: The murine IgM monoclonal Ab STRO-1, produced from an immunization with a population of human CD34+ bone marrow cells, can identify a cell surface antigen expressed by stromal elements in human bone marrow.55 From bone marrow cells, the frequency of fibroblast colony-forming cells (CFU-F) is enriched approximately 100-fold in the STRO-1+/Glycophorin A- population than in the STRO-1+/Glycophorin A+ population.55 A STRO-1+ enriched subset of marrow cells is capable of differentiating into multiple mesenchymal lineages including hematopoiesis-supportive stromal cells with a vascular smooth muscle-like phenotype, adipocytes, osteoblasts and chondrocytes.56, 57, 58, 59 STRO-1 is a valuable Ab for the identification, isolation and functional characterization of human bone marrow stromal cell precursors, which are quite distinct from those of primitive HSCs.
Neural Stem Cell Markers
Nestin: Nestin is a class VI intermediate filament protein.60,61 Although it is expressed predominantly in stem cells of the central nervous system (CNS),62 its expression is absent from nearly all mature CNS cells.63 Nestin has been the most extensively used marker to identify CNS stem cells within various areas of the developing nervous system and in cultured cells in vitro.34, 64, 65, 66, 67,68 The role of nestin in CNS stem cell biology, however, remains undefined. Although nestin does not form intermediate filaments by itself in vitro it does co-assemble with vimentin or alpha-internexin to form and heterodimer, coiled-coil complexes that may then form intermediate filaments.69 Its transient expression has been suggested to be a major step in the neural differentiation pathway.61 Nestin expression has also been discovered in non-neural stem cell populations, such as pancreatic islet progenitors70, 71, 72 as well as hematopoietic progenitors.73
PSA-NCAM (Polysialic acid-neural cell adhesion molecule): The regulated expression of neural cell adhesion molecule (NCAM) isoforms in the brain is critical for many neural developmental processes. The embryonic form of NCAM, PSA-NCAM, is highly polysialylated and is mainly expressed in the developing nervous system.74 PSA-NCAM may be related to synaptic rearrangement and plasticity.75 In the adult, PSA-NCAM expression is restricted to regions that retain plasticity.76A neuronal-restricted precursor identified by its high expression of PSA-NCAM can undergo self-renewal and differentiate into multiple neuronal phenotypes.77PSA-NCAM+ neonatal brain precursors are restricted to a glial fate and thyroid hormone can modulate them into an oligodendrocyte fate.78, 79, 80 Polysialic acid modification significantly decreases NCAM adhesiveness and therefore, it was originally suggested PSA-NCAM works as a purely anti-adhesive factor that modulates cell-cell interactions in promoting brain plasticity. Increasing evidence indicates that PSA-NCAM may interact with secreted signaling molecules to perform an instructive role in development.81, 82

Figure 3. The structure of NGF with a model of the p75 Neurotrophin Receptor. The extracellular domain of the receptor is taken from the tumor necrosis factor receptor structure and the intracellular portion contains a death domain.
p75 Neurotrophin R (NTR): p75 NTR, also named low affinity nerve growth factor (NGF) receptor, is a type I transmembrane protein that belongs to the tumor necrosis factor receptor superfamily (Figure 3).83 It binds to NGF, BDNF, NT-3 and NT-4 equally (with low affinity). p75NTR, when activated in the presence of Trk, enhances responses to neurotrophin (see reference 84 for a review). TrkC receptors working together with p75 NTR have been suggested to serve critical functions during the development of the nervous system.85 Neural crest stem cells (NCSCs) have been isolated based on their surface expression of p75NTR.86, 87 Freshly isolated p75NTR+ NCSCs from peripheral nerve tissues can self-renew and generate neurons and glia both in vitro and in vivo. In addition, neuroepithelial-derived p75NTR+ cells are also able to differentiate into neurons, smooth muscle and Schwann cells in culture.88 Recently, p75 NTR has been used as a marker to identify mesenchymal precursors as well as hepatic stellate cells.89, 90
References
- Scholer, H.R. et al. (1990) Nature 344:435.
- Scholer, H.R. et al. (1989) EMBO J. 8:2543.
- Rosner, M.H. et al. (1990) Nature 345:686.
- Niwa, H. et al. (2000) Nat. Genet. 24:372.
- Pesce, M. et al. (2001) Stem Cells 19:271.
- Bruce, A. et al. (1990) BioEssays 12:173.
- Solter, D. et al. (1978) Proc. Natl. Acad. Sci. USA 75:5565.
- Knowles, B.B. et al. (1980) Nature 288:615.
- Fox, N. et al. (1981) Dev. Biol. 83:391.
- Shevinsky, L.H. et al. (1982) Cell 30:697.
- Kannagi, R. et al. (1983) EMBO J. 2:2355.
- Thomson, J.A. et al. (1998) Science 282:1145.
- Thomson, J.A. et al. (1998) Curr. Top. Dev. Biol. 38:133.
- Civin, C.I. et al. (1984) J. Immunol. 133:157.
- Sutherland, H.J. et al. (1989) Blood 74:1563.
- Bhatis, M. et al. (1997) Proc. Natl. Acad. Sci. USA 94:5320.
- Berenson, R.J. et al. (1988) J. Clin. Invest. 81:951.
- Civin, C.I. et al. (1996) J. Clin. Oncol. 14:2224.
- Link, H. et al. (1996) Blood 87:4903.
- Shpall, E.J. et al. (1997) Blood 90:4313.
- Yabe, H. et al. (1996) Bone Marrow Transplant. 17:985.
- Sutherland, D.R. et al. (1992) J. Hematother. 1:115.
- Andrew, R. et al. (1989) J. Exp. Med. 169:1721.
- Osawa, M. et al. (1996) Science 273:242.
- Bhatis, M. et al. (1998) Nat. Med. 4:1038.
- Zanjani, E.D. et al. (1998) Exp. Hematol. 26:353.
- Nakamura, Y. et al. (1999) Blood 94:4053.
- Sato, T. et al. (1999) Blood 94:2548.
- Yin, A.H. et al. (1997) Blood 90:5002.
- Miraglia, S. et al. (1997) Blood 90:5013.
- Yu, Y. et al. (2002) J. Biol. Chem. 277:20711.
- Kobari, L. et al. (2001) J. Hematother. Stem Cell Res. 10:273.
- Gehling, U.M. et al. (2000) Blood 95:3106.
- Uchida, N. et al. (2000) Proc. Natl. Acad. Sci. USA 97:14720.
- Zhou, S. et al. (2001) Nat. Med. 7:1028.
- Kim, M. et al. (2002) Clin. Cancer Res. 8:22.
- Doyle, L.A. et al. (1998) Proc. Natl. Acad. Sci. USA 95:15665.
- Rocchi, E. et al. (2000) Biochem. Biophys. Res. Commun.271:42.
- Scharenberg, C.W. et al. (2002) Blood 99:507.
- Jackson, K.A. et al. (2001) J. Clin. Invest. 107:1395.
- Gussoni, E. et al. (1999) Nature 401:390.
- Bunting, K.D. (2002) Stem Cells 20:11.
- Van de Rijn, M. et al. (1989) Proc. Natl. Acad. Sci. USA 86:4634.
- Spangrude, G.I. et al. (1988) Science 241:58.
- Spangrude, G.I. et al. (1993) Blood 82:3327.
- Morrison, S.J. et al. (1995) Proc. Natl. Acad. Sci. USA 92:10302.
- Kawamoto, H. et al. (1997) Int. Immunol. 9:1011.
- Yamamoto, Y. et al. (1996) Blood 88:445.
- Morrison, S.J. et al. (1997) Proc. Natl. Acad. Sci. USA 94:1908.
- Welm, B.E. et al. (2002) Dev. Biol. 245:42.
- Codias, E.K. et al. (1990) J. Immunol. 144:2197.
- Malek, T.R. et al. (1986) J. Exp. Med. 164:709.
- Codias, E.K. et al. (1990) J. Immunol. 145:1407.
- Flood, P.M. et al. (1990) J. Exp. Med. 172:115.
- Simmons, P.J. et al. (1991) Blood 78:55.
- Gronthos, S. et al. (1994) Blood 84:4164.
- Encina, N.R. et al. (1999) Lab. Invest. 79:449.
- Oyajobi, B.O. et al. (1999) J. Bone. Miner. Res. 14:351.
- Dennis, J.E. et al. (2002) Cells Tissues Organs 170:73.
- Hockfield, S. et al. (1985) J. Neurosci. 5:3310.
- Lendahl, U. et al. (1990) Cell 60:585.
- Frederiksen, K. et al. (1988) J. Neurosci. 8:1144.
- Tohyama, T. et al. (1992) Lab. Invest. 66:303.
- Frederiksen, K. et al. (1988) Neuron 1:439.
- Cattaneo, C. et al. (1990) Nature 347:762.
- Reynolds, B.A. et al. (1992) Science 255:1707.
- Rietze, R.L. et al. (2001) Nature 412:736.
- Carpenter, M.K. et al. (2001) Exp. Neurol. 172:383.
- Steinert, P.M. et al. (1999) J. Biol. Chem. 274:9881.
- Zulewski, H. et al. (2001) Diabetes 50:521.
- Lumelsky, N. et al. (2001) Science 292:1389.
- Lechner, A. et al. (2002) Biochem. Biophys. Res. Commun. 293:670.
- Shih, C.C. et al. (2001) Blood 98:2412.
- Kiss, J.Z. et al. (2001) Rev. Neurosci. 12:297.
- Muller, D. et al. (1996) Neuron 17:413.
- Theodosis, D.T. et al. (1994) Psychoneuroendocrinology 19:455.
- Mayer-Proschel, M. et al. (1997) Neuron 19:773.
- Ben-Hur, T. et al. (1998) J. Neurosci. 18:5777.
- Theodosis, D.T. et al. (1999) J. Neurosci. 19:10228.
- Keirstead, H.S. et al. (1999) J. Neurosci. 19:7529.
- Muller, D. et al. (2000) Proc. Natl. Acad. Sci. USA 97:4315.
- Kiss, J.Z. et al. (2001) Brain Res. Brain Res. Rev. 36:175.
- Barker, P.A. et al. (1992) Mol. Cell. Biochem. 110:1.
- Kaplan, D.R. et al. (1997) Curr. Opin. Cell. Biol. 9:213.
- Hapner, S.J. et al. (1998) Dev. Biol. 201:90.
- Stemple, D.L. et al. (1992) Cell 71:973.
- Morrison, S.J. et al. (1999) Cell 96:737.
- Mujtaba, T. et al. (1998) Dev. Biol. 200:1.
- Campagnolo, L. et al. (2001) Biol. Reprod. 64:464.
- Cassiman, D. et al. (2001) Hepatology 33:148.
