Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
R&D Systems | Catalog # 11060-CV

Key Product Details
- R&D Systems HEK293-derived Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein (11060-CV)
- Quality control testing to verify active proteins with lot specific assays by in-house scientists
- All R&D Systems proteins are covered with a 100% guarantee
Source
Accession Number
Applications
Product Specifications
Source
SARS-CoV-2 B.1.1.529 Spike (Val16 - Lys1211) (Ala67Val, His69del, Val70del, Thr95Ile, Gly142Asp, Val143del, Tyr144del, Tyr145del, Asn211del, Leu212Ile, ins214Glu-Pro-Glu, Gly339Asp, Ser371Leu, Ser373Pro, Ser375Phe, Lys417Asn, Asn440Lys, Gly446Ser, Ser477Asn, Thr478Lys, Glu484Ala, Gln493Arg, Gly496Ser, Gln498Arg, Asn501Tyr, Tyr505His, Thr547Lys, Asp614Gly, His655Tyr, Asn679Lys, Pro681His, Asn764Lys, Asp796Tyr, Asn856Lys, Gln954His, Asn969Lys, Leu981Phe) (Arg682Ser, Arg685Ser, Lys986Pro, Val987Pro) Accession # YP_009724390.1 |
6-His tag |
| N-terminus | C-terminus |
Purity
Endotoxin Level
N-terminal Sequence Analysis
Predicted Molecular Mass
SDS-PAGE
Activity
Scientific Data Images for Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
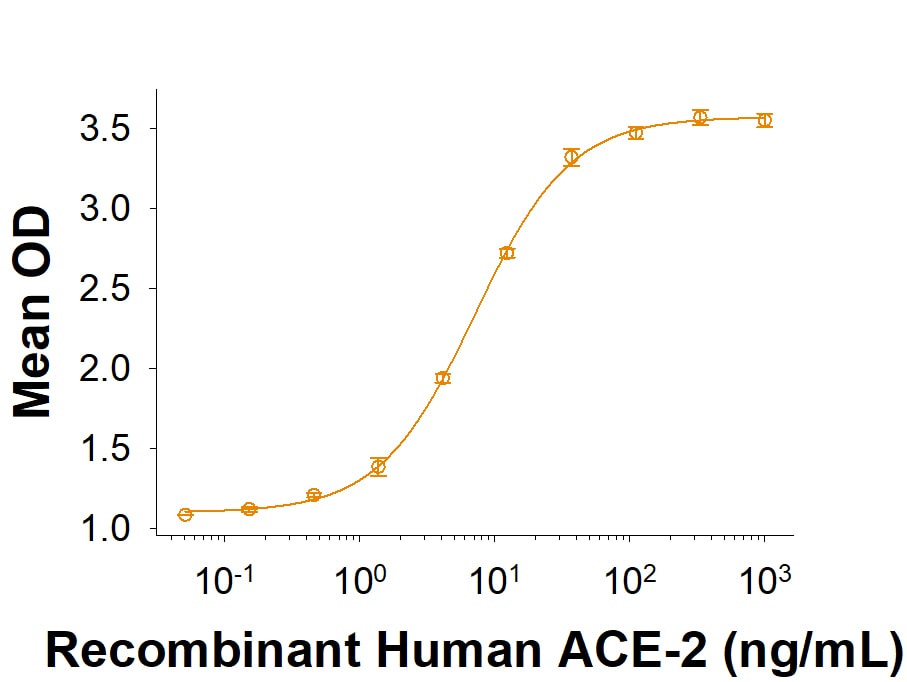
Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein Binding Activity.
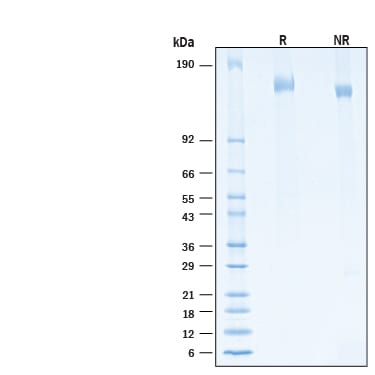
Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag (Catalog # 11060-CV) binds Recombinant Human ACE-2 His-tag (933-ZN) in a functional ELISA.Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein SDS-PAGE.
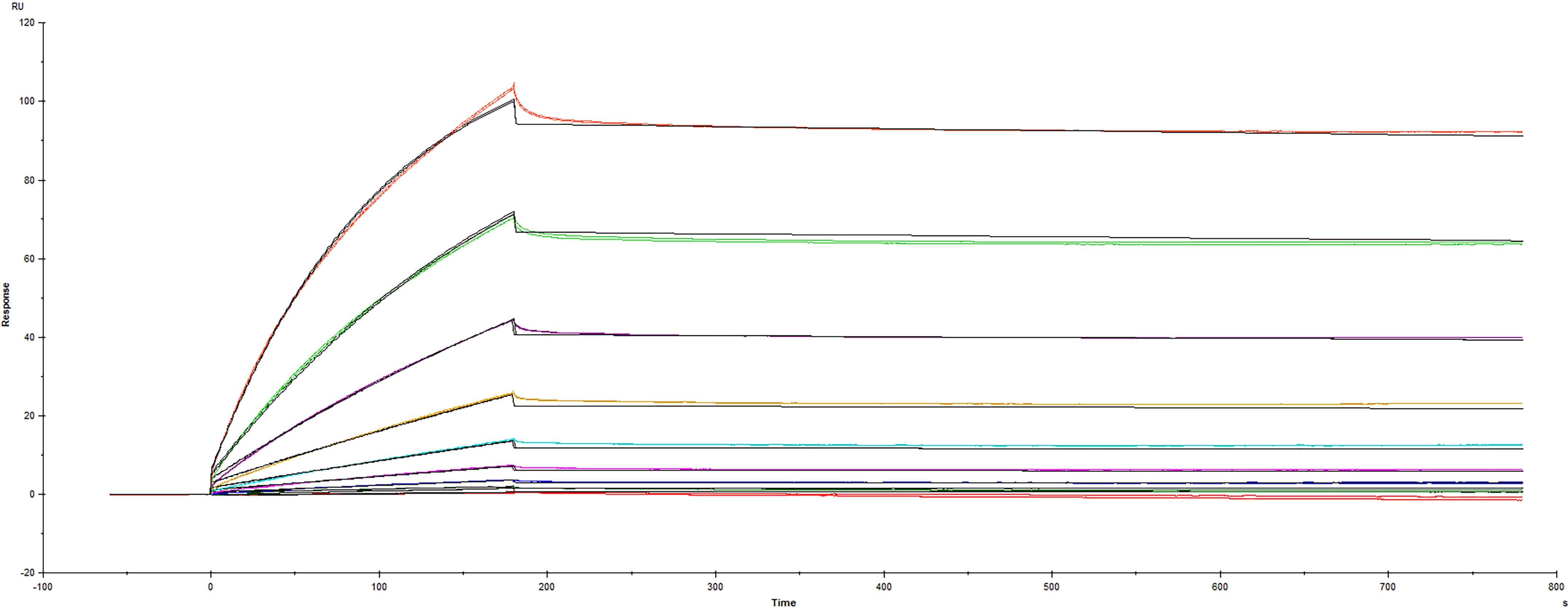
2 μg/lane of Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein (Catalog # 11060-CV) was resolved with SDS-PAGE under reducing (R) and non-reducing (NR) conditions and visualized by Coomassie® Blue staining, showing bands at 146 - 166 kDa.Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein binds to Recombinant Human ACE-2 by surface plasmon resonance
Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein (Catalog #11060-CV) was immobilized on a Biacore Sensor Chip CM5, and binding to recombinant human ACE-2 (933-ZN) was measured at a concentration range between 0.021 nM and 53.1 nM. The double-referenced sensorgram was fit to a 1:1 binding model to determine the binding kinetics and affinity, with an affinity constant of Kd=3.63 nM.Formulation, Preparation, and Storage
11060-CV
| Formulation | Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. |
| Reconstitution | Reconstitute at 500 μg/mL in PBS. |
| Shipping | The product is shipped at ambient temperature. Upon receipt, store it immediately at the temperature recommended below. |
| Stability & Storage | Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
|
Calculators
Background: Spike
References
- Wu, F. et al. (2020) Nature 579:265.
- Tortorici, M.A. and D. Veesler (2019) Adv. Virus Res. 105:93.
- Bosch, B.J. et al. (2003) J. Virol. 77:8801.
- Belouzard, S. et al. (2009) Proc. Natl. Acad. Sci. 106:5871.
- Millet, J.K. and G.R. Whittaker (2015) Virus Res. 202:120.
- Ortega, J.T. et al. (2020) EXCLI J. 19:410.
- Wang, K. et al. (2020) bioRxiv https://www.biorxiv.org/content/10.1101/2020.03.14.988345v1.
- Isabel, S. et al. (2020) Sci Rep. 10, 14031. https://doi.org/10.1038/s41598-020-70827-z.
- Tai, W. et al. (2020) Cell. Mol. Immunol. 17:613.
- Okba, N. M. A. et al. (2020). Emerg. Infect. Dis. https://doi.org/10.3201/eid2607.200841.
- Shah, M. and Woo, H.G. (2021) bioRxiv https://doi.org/10.1101/2021.12.04.471200.
- Lupala, C.S. et al. (2021) bioRxiv https://doi.org/10.1101/2021.12.10.472102.
- Zhang, L. et al. (2020) Nat Commun. 11:6013.
- Lasek-Nesselquist, E. et al. (2021) medRxiv https://doi.org/10.1101/2021.03.10.21253285.
- Scheepers, C. et al. (2021) medRxiv https://doi.org/10.1101/2021.08.20.21262342.
- Callaway, E. and Ledford, H. (2021) Nature 600:197.
Long Name
Alternate Names
Entrez Gene IDs
Gene Symbol
UniProt
Additional Spike Products
Product Documents for Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
For research use only
Related Research Areas
Citations for Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
Customer Reviews for Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF
There are currently no reviews for this product. Be the first to review Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF and earn rewards!
Have you used Recombinant SARS-CoV-2 B.1.1.529 Spike His-tag Protein, CF?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review