SARS-CoV-2 Spike RBD LlaMABodyTM Bivalent VHH HuIgG2 Fusion Antibody
R&D Systems | Catalog # LMAB10731

Key Product Details
Species Reactivity
Applications
Label
Antibody Source
Product Specifications
Immunogen
Arg319-Phe541
Accession # YP_009724390.1
Specificity
Clonality
Host
Isotype
Endotoxin Level
Scientific Data Images
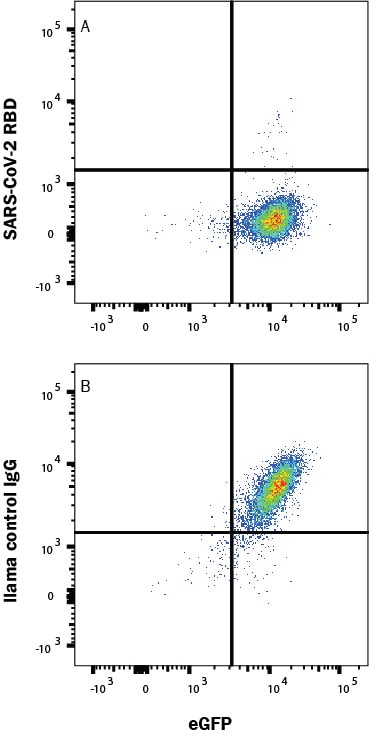
SARS-Cov-2 Spike 1 protein binding to ACE-2-transfected Human Cell Line is Blocked by SARS-Cov-2 Spike RBD Antibody.
In a functional flow cytometry test, Recombinant SARS-Cov-2 Spike 1 His-tagged protein (10522-CV) binds to HEK293 human embryonic kidney cell line transfected with recombinant human ACE-2 and eGFP. (A) Binding is completely blocked by 50 µg/mL of Llama Anti-SARS-Cov-2 Spike RBD Monoclonal Antibody (Catalog # LMAB10731) but not by (B) Llama IgG1 Control. Protein binding was detected with Mouse Anti-His APC-conjugated Monoclonal Antibody (IC050A). Staining was performed using our Staining Membrane-Associated Proteins protocol.Applications
Blockade of Receptor-ligand Interaction
Sample: In a functional flow cytometry test, 50 μg/mL of Llama Anti-SARS-Cov-2 Spike RBD Antibody(Catalog # LMAB10731) will block the binding of Recombinant SARS-Cov-2 Spike 1 His-tagged protein (Catalog # 10522-CV) to HEK293 human embryonic kidney cell line transfected with recombinant human ACE-2.
Formulation, Preparation, and Storage
Purification
Reconstitution
Reconstitute at 0.5 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Formulation
Shipping
Stability & Storage
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: Spike RBD
References
- Wu, F. et al. (2020) Nature 579:265.
- Tortorici, M.A. and D. Veesler (2019). Adv. Virus Res. 105:93.
- Bosch, B.J. et al. (2003). J. Virol. 77:8801.
- Belouzard, S. et al. (2009) Proc. Natl. Acad. Sci. 106:5871.
- Millet, J.K. and G. R. Whittaker (2015) Virus Res. 202:120.
- Yuan, Y. et al. (2017) Nat. Commun. 8:15092.
- Walls, A.C. et al. (2010) Cell 180:281.
- Jiang, S. et al. (2020) Trends. Immunol. https://doi.org/10.1016/j.it.2020.03.007.
- Ortega, J.T. et al. (2020) EXCLI J. 19:410.
- Wrapp, D. et al. (2020) Science 367:1260.
- Tai, W. et al. (2020) Cell. Mol. Immunol. https://doi.org/10.1016/j.it.2020.03.007.
- Okba, N. M. A. et al. (2020). Emerg. Infect. Dis. https://doi.org/10.3201/eid2607.200841.
- Wang, X. et al. (2020) https://doi.org/10.1038/s41423-020-0424-9.
- Wang, K. et al. (2020) bioRxiv https://www.biorxiv.org/content/10.1101/2020.03.14.988345v1.
Long Name
Gene Symbol
UniProt
Additional Spike RBD Products
Product Documents
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices
For research use only
Related Research Areas
Customer Reviews
There are currently no reviews for this product. Be the first to review SARS-CoV-2 Spike RBD LlaMABodyTM Bivalent VHH HuIgG2 Fusion Antibody and earn rewards!
Have you used SARS-CoV-2 Spike RBD LlaMABodyTM Bivalent VHH HuIgG2 Fusion Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
FAQs
-
Q: What are Camelid Antibodies?
A: Camelid antibodies are antibodies from the Camelidae family of mammals that include llamas, camels, and alpacas. These animals produce 2 main types of antibodies. One type of antibody camelids produce is the conventional antibody that is made up of 2 heavy chains and 2 light chains. They also produce another type of antibody that is made up of only 2 heavy chains. This is known as heavy chain IgG (hcIgG). While these antibodies do not contain the CH1 region, they retain an antigen binding domain called the VHH region. VHH antibodies, also known as single domain antibodies or Nanobodies®, contain only the VHH region from the camelid antibody.
