Histone H3 [ac Lys4] Antibody - BSA Free
Novus Biologicals | Catalog # NB21-1024

Key Product Details
Species Reactivity
Validated:
Human, Mouse, Rat, C. elegans
Cited:
Rat
Predicted:
Chicken (100%), Drosophila (100%), Plant (100%), Xenopus (100%). Backed by our 100% Guarantee.
Applications
Validated:
Western Blot, Immunocytochemistry/ Immunofluorescence, Chromatin Immunoprecipitation (ChIP), Dot Blot
Cited:
Western Blot
Label
Unconjugated
Antibody Source
Polyclonal Rabbit IgG
Format
BSA Free
Loading...
Product Specifications
Immunogen
This Histone H3 [ac Lys4] antibody was raised against synthetic acetylated peptide surrounding Lysine 4 of human Histone H3.2 [Swiss Prot Q71DI3].
Modification
ac Lys4
Localization
Nucleus; Chromosome
Clonality
Polyclonal
Host
Rabbit
Isotype
IgG
Theoretical MW
15 kDa.
Disclaimer note: The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Disclaimer note: The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Scientific Data Images for Histone H3 [ac Lys4] Antibody - BSA Free
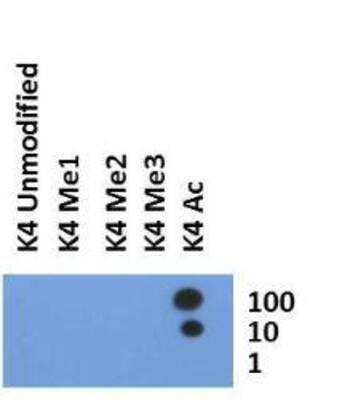
Dot Blot: Histone H3 [ac Lys4] Antibody [NB21-1024] - Analysis of Histone H3 K4ac antibody in picomoles of peptide.
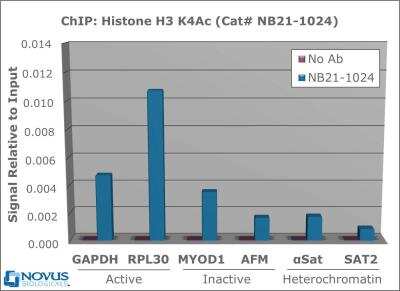
Chromatin Immunoprecipitation: Histone H3 [ac Lys4] Antibody [NB21-1024] - Histone H3 [ac Lys4] Antibody [NB21-1024] - Chromatin from one million formaldehyde cross-linked Hela cells was used with 2ug of NB21-1024 and 20ul of magnetic IgG beads per immunoprecipitation. A no antibody (No Ab) control was also used. Immunoprecipitated DNA was quantified using quantitative real-time PCR and SYBR green dye, then normalized to the non-precipitated input chromatin, which is equal to one.
Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024]
Western Blot: Histone H3 [ac Lys4] Antibody [NB21-1024] - Analysis of Histone H3 K4ac in C. elegans embryo lysate. Observed molecular weight is ~17 kDa.Immunocytochemistry/ Immunofluorescence: Histone H3 [ac Lys4] Antibody - BSA Free [NB21-1024]
Immunocytochemistry/Immunofluorescence: Histone H3 [ac Lys4] Antibody [NB21-1024] - Histone H3 K4ac1 antibody was tested in HeLa cells with DyLight 488 (green). Nuclei and alpha-tubulin were counterstained with DAPI (blue) and DyLight 550 (red).Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024]
Western Blot: Histone H3 [ac Lys4] Antibody [NB21-1024] - Analysis of Histone H3 K4ac in (1) HeLa histone prep and (2) NIH 3T3 histone prep lysates. Observed molecular weight is ~16 kDa.Applications for Histone H3 [ac Lys4] Antibody - BSA Free
Application
Recommended Usage
Chromatin Immunoprecipitation (ChIP)
2-5 ug per million cells
Dot Blot
1:1000
Immunocytochemistry/ Immunofluorescence
1:1000
Western Blot
1:500
Application Notes
In Western Blot a band is seen at ~15-16 kDa in HeLa histone prep, NIH 3T3 histone prep, and C. elegans embryo lysate. The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Reviewed Applications
Read 1 review rated 5 using NB21-1024 in the following applications:
Formulation, Preparation, and Storage
Purification
Immunogen affinity purified
Formulation
PBS and 30% Glycerol
Format
BSA Free
Preservative
0.05% Sodium Azide
Concentration
1.15 mg/ml
Shipping
The product is shipped with polar packs. Upon receipt, store it immediately at the temperature recommended below.
Stability & Storage
Store at 4C short term. Aliquot and store at -20C long term. Avoid freeze-thaw cycles.
Background: Histone H3
Histones are nuclear proteins responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Changes in chromatin structure play a large role in the regulation of transcription. The chromatin fibers are compacted through the interaction of a linker histone, H1, with the DNA between the nucleosomes to form higher order chromatin structures.
Common histone modifications include methylation of lysine and arginine, acetylation of lysine, phosphorylation of threonine and serine, and sumoylation, biotinylation, and ubiquitylation of lysine. Posttranslational modifications such as acetylation of core histones regulates gene expression, thus altering protein function and regulation (1). Histone H3 is primarily acetylated at lysines 9, 14, 18, and 23 and have a theoretical molecular weight of 15 kDa. Acetylation at lysine 9 and 14 appears to control histone deposition, chromatin assembly and active transcription. Methylation of arginine residues within histone H3 has also been linked to transcription regulation. Histone H3 has been linked to various types of cancer as a biomarker through the aberrant expression of histone deacetylase (HDAC) enzymes and changes to chromatins (2-4).
References
1. Zhang, Y. X., Akumuo, R. C., Espana, R. A., Yan, C. X., Gao, W. J., & Li, Y. C. (2018). The histone demethylase KDM6B in the medial prefrontal cortex epigenetically regulates cocaine reward memory. Neuropharmacology, 141, 113-125. doi:10.1016/j.neuropharm.2018.08.030
2. Nandakumar, V., Hansen, N., Glenn, H. L., Han, J. H., Helland, S., Hernandez, K,...Meldrum, D. R. (2016). Vorinostat differentially alters 3D nuclear structure of cancer and non-cancerous esophageal cells. Sci Rep, 6, 30593. doi:10.1038/srep30593
3. Zhou, M., Li, Y., Lin, S., Chen, Y., Qian, Y., Zhao, Z., & Fan, H. (2019). H3K9me3, H3K36me3, and H4K20me3 Expression Correlates with Patient Outcome in Esophageal Squamous Cell Carcinoma as Epigenetic Markers. Dig Dis Sci, 64(8), 2147-2157. doi:10.1007/s10620-019-05529-2
4. Li, Y., Guo, D., Sun, R., Chen, P., Qian, Q., & Fan, H. (2019). Methylation Patterns of Lys9 and Lys27 on Histone H3 Correlate with Patient Outcome in Gastric Cancer. Dig Dis Sci, 64(2), 439-446. doi:10.1007/s10620-018-5341-8
Alternate Names
H3F3A, H3K4ac
Gene Symbol
H3C14
UniProt
Additional Histone H3 Products
Product Documents for Histone H3 [ac Lys4] Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for Histone H3 [ac Lys4] Antibody - BSA Free
Epi-Plus antibody production in collaboration with Rockland Immunochemicals Inc.
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Related Research Areas
Citations for Histone H3 [ac Lys4] Antibody - BSA Free
Customer Reviews for Histone H3 [ac Lys4] Antibody - BSA Free (1)
5 out of 5
1 Customer Rating
Have you used Histone H3 [ac Lys4] Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
1 of
1 review
Showing All
Filter By:
-
Application: Western BlotSample Tested: Human breast cell lysate (MDA-MB-231)Species: HumanVerified Customer | Posted 09/26/2013
There are no reviews that match your criteria.
Protocols
View specific protocols for Histone H3 [ac Lys4] Antibody - BSA Free (NB21-1024):
Histone H3 [ac Lys4] Antibody:
Chromatin Immunoprecipitation Protocol
Cell Fixation and Preparation
1. Begin with a cell culture that has reached 80% confluency.
2. Add formaldehyde to a final concentration of 1% in growth media and incubate for 10 minutes at room temperature.
3. Add glycine to reach a final concentration of 125 mM in the media. Incubate for 5 minutes at room temperature.
4. Remove all media and wash twice with 20 mL of ice cold PBS.
5. Add 2 mL of ice cold PBS with protease inhibitors*. Scrape cells into microcentrifuge tube.
6. Spin cells at 4 C for 5 minutes at 800 x g.
7. Remove supernatant and resuspend cells in 750 ul of RIPA lysis buffer containing protease inhibitors* per 10,000,000 cells (enough for 10 IPs). Incubate at 4 C for 15 minutes.
8. Spin cells at 4 C for 5 minutes at 800 x g.
DNA Shearing by Sonication
1. Sonicate crosslinked DNA to fragments sizes of 200-1000 base pairs. Keep samples ice cold to prevent denaturing of chromatin. Conditions for fragmenting must be empirically derived, and vary depending on equipment, cell type, cell density, and cross-linking efficiency.
2. Centrifuge samples to remove debris at 4 C for 10 minutes at 12,500 x g. Remove supernatant and transfer to a new tube. Discard pellet. Set aside 75 ul of sample for input fraction, which will not go through the subsequent IP steps. The remaining sample can be moved into 75 ul aliquots, each of which is sufficient for a single IP. Although it is preferable to proceed directly to the following steps, sheared chromatin can now be frozen at -80 C for up to 1 month.
a. Optional: Test the efficiency of the shearing by running 5-10 ul of sample on a 2% agarose gel after reversal of crosslinking, RNase treatment (0.5 mg/ml) and proteinase K treatment (0.1 ug/ul) as described below.
Chromatin Immunoprecipitation
Recommended controls include: No antibody negative control OR normal IgG negative control, positive control antibody.
1. Dilute each IP sample 1:10 by adding 75 ul sheared chromatin to 675 ul dilution buffer, along with appropriate protease inhibitors*. Save undiluted input fraction for step 10.
2. Add 25 ul of fully suspended protein A/G magnetic bead slurry. Do not allow the beads to dry.
3. Add antibody of interest to the diluted sample. For best results, incubate tubes with rotation at 4 C overnight. Alternatively, incubate at room temperature for 1-2 hours.
4. Pellet magnetic beads with a magnetic separator and remove the supernatant.
5. Add 750 ul cold low salt buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
6. Add 750 ul cold high salt buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
7. Add 750 ul cold LiCl buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
8. Add 750 ul cold TE buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
9. Elute complex by adding 200 ul elution buffer and agitate at RT for 15 minutes. Pellet beads with separator and discard beads. Keep the supernatant.
10. Add 8 ul of 5M NaCl and 2 ul of proteinase K (10 ug/ul) to each sample to reverse cross-linking. For the input fraction, add 125 ul of elution buffer along with NaCl and proteinase K. Incubate at 62 C for 4 hours or overnight. Incubate at 95 C for 10 minutes to deactivate proteinase K.
a. Optional: To decrease time, this step may also be performed at 95 C for 20 minutes without proteinase K treatment.
DNA Purification and Amplification
1. DNA can now be purified with either a commercially available column kit or by phenol/chloroform extraction.
2. Perform real-time PCR with 2 ul of purified DNA and primers (catalog numbers NBP1-71650, NBP1-71651, NBP1-71652, NBP1-71653, NBP1-71654 and NBP1-71655) per reaction. Dilute input fraction to 1% before PCR. Normalize all IPs and no antibody control IP to adjusted input fraction.
a. Ex. Raw input CT=30, adjusted input: 30 - 6.44 = 23.4
b. Antibody CT=28
c. Normalized signal relative to input: 2 ^ (23.4-28) = 0.04
Buffers
Dilution Buffer: 0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCL (pH8.1), 167 mM NaCl
Low Salt Wash Buffer: 0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 150mM NaCl.
High Salt Wash Buffer: 0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 500mM NaCl.
LiCl Wash Buffer: 250mM LiCl, 1% NP-40, 1% Na-deoxycholate, 1mM EDTA, 10mM Tris, pH 8.1.
TE Buffer: 10mM Tris-HCL pH 8.1, 1 mM EDTA
Elution buffer: 1% SDS, 100mM NaHCO3
* Please note that protease inhibitors have variable half-lives and should be freshly prepared as applicable.
Western Blot Protocol
1. Perform SDS-PAGE on samples to be analyzed, loading 10 ug of histone preps per lane.
2. Transfer proteins to membrane according to the instructions provided by the manufacturer of the membrane and transfer apparatus.
3. Stain according to standard Ponceau S procedure (or similar product) to assess transfer success, and mark molecular weight standards where appropriate.
4. Rinse the blot.
5. Block the membrane using standard blocking buffer for at least 1 hour.
6. Wash the membrane in wash buffer three times for 10 minutes each.
7. Dilute primary antibody in blocking buffer and incubate 1 hour at room temperature.
8. Wash the membrane in wash buffer three times for 10 minutes each.
9. Apply the diluted HRP conjugated secondary antibody in blocking buffer (as per manufacturers instructions) and incubate 1 hour at room temperature.
10. Wash the blot in wash buffer three times for 10 minutes each (this step can be repeated as required to reduce background).
11. Apply the detection reagent of choice in accordance with the manufacturers instructions.
Note: Tween-20 can be added to the blocking or antibody dilution buffer at a final concentration of 0.05-0.2%.
Immunocytochemistry Protocol
Culture cells to appropriate density in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and add 10% formalin to the dish. Fix at room temperature for 30 minutes.
2. Remove the formalin and add ice cold methanol. Incubate for 5-10 minutes.
3. Remove methanol and add washing solution (i.e. PBS). Be sure to not let the specimen dry out. Wash three times for 10 minutes.
4. To block nonspecific antibody binding incubate in 10% normal goat serum from 1 hour to overnight at room temperature.
5. Add primary antibody at appropriate dilution and incubate at room temperature from 2 hours to overnight at room temperature.
6. Remove primary antibody and replace with washing solution. Wash three times for 10 minutes.
7. Add secondary antibody at appropriate dilution. Incubate for 1 hour at room temperature.
8. Remove antibody and replace with wash solution, then wash for 10 minutes. Add Hoechst 33258 to wash solution at 1:25,0000 and incubate for 10 minutes. Wash a third time for 10 minutes.
9. Cells can be viewed directly after washing. The plates can also be stored in PBS containing Azide covered in Parafilm (TM). Cells can also be cover-slipped using Fluoromount, with appropriate sealing.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow safe laboratory procedures.
Chromatin Immunoprecipitation Protocol
Cell Fixation and Preparation
1. Begin with a cell culture that has reached 80% confluency.
2. Add formaldehyde to a final concentration of 1% in growth media and incubate for 10 minutes at room temperature.
3. Add glycine to reach a final concentration of 125 mM in the media. Incubate for 5 minutes at room temperature.
4. Remove all media and wash twice with 20 mL of ice cold PBS.
5. Add 2 mL of ice cold PBS with protease inhibitors*. Scrape cells into microcentrifuge tube.
6. Spin cells at 4 C for 5 minutes at 800 x g.
7. Remove supernatant and resuspend cells in 750 ul of RIPA lysis buffer containing protease inhibitors* per 10,000,000 cells (enough for 10 IPs). Incubate at 4 C for 15 minutes.
8. Spin cells at 4 C for 5 minutes at 800 x g.
DNA Shearing by Sonication
1. Sonicate crosslinked DNA to fragments sizes of 200-1000 base pairs. Keep samples ice cold to prevent denaturing of chromatin. Conditions for fragmenting must be empirically derived, and vary depending on equipment, cell type, cell density, and cross-linking efficiency.
2. Centrifuge samples to remove debris at 4 C for 10 minutes at 12,500 x g. Remove supernatant and transfer to a new tube. Discard pellet. Set aside 75 ul of sample for input fraction, which will not go through the subsequent IP steps. The remaining sample can be moved into 75 ul aliquots, each of which is sufficient for a single IP. Although it is preferable to proceed directly to the following steps, sheared chromatin can now be frozen at -80 C for up to 1 month.
a. Optional: Test the efficiency of the shearing by running 5-10 ul of sample on a 2% agarose gel after reversal of crosslinking, RNase treatment (0.5 mg/ml) and proteinase K treatment (0.1 ug/ul) as described below.
Chromatin Immunoprecipitation
Recommended controls include: No antibody negative control OR normal IgG negative control, positive control antibody.
1. Dilute each IP sample 1:10 by adding 75 ul sheared chromatin to 675 ul dilution buffer, along with appropriate protease inhibitors*. Save undiluted input fraction for step 10.
2. Add 25 ul of fully suspended protein A/G magnetic bead slurry. Do not allow the beads to dry.
3. Add antibody of interest to the diluted sample. For best results, incubate tubes with rotation at 4 C overnight. Alternatively, incubate at room temperature for 1-2 hours.
4. Pellet magnetic beads with a magnetic separator and remove the supernatant.
5. Add 750 ul cold low salt buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
6. Add 750 ul cold high salt buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
7. Add 750 ul cold LiCl buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
8. Add 750 ul cold TE buffer and wash for 5 minutes with rotation. Pellet beads with separator and discard supernatant.
9. Elute complex by adding 200 ul elution buffer and agitate at RT for 15 minutes. Pellet beads with separator and discard beads. Keep the supernatant.
10. Add 8 ul of 5M NaCl and 2 ul of proteinase K (10 ug/ul) to each sample to reverse cross-linking. For the input fraction, add 125 ul of elution buffer along with NaCl and proteinase K. Incubate at 62 C for 4 hours or overnight. Incubate at 95 C for 10 minutes to deactivate proteinase K.
a. Optional: To decrease time, this step may also be performed at 95 C for 20 minutes without proteinase K treatment.
DNA Purification and Amplification
1. DNA can now be purified with either a commercially available column kit or by phenol/chloroform extraction.
2. Perform real-time PCR with 2 ul of purified DNA and primers (catalog numbers NBP1-71650, NBP1-71651, NBP1-71652, NBP1-71653, NBP1-71654 and NBP1-71655) per reaction. Dilute input fraction to 1% before PCR. Normalize all IPs and no antibody control IP to adjusted input fraction.
a. Ex. Raw input CT=30, adjusted input: 30 - 6.44 = 23.4
b. Antibody CT=28
c. Normalized signal relative to input: 2 ^ (23.4-28) = 0.04
Buffers
Dilution Buffer: 0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCL (pH8.1), 167 mM NaCl
Low Salt Wash Buffer: 0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 150mM NaCl.
High Salt Wash Buffer: 0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 500mM NaCl.
LiCl Wash Buffer: 250mM LiCl, 1% NP-40, 1% Na-deoxycholate, 1mM EDTA, 10mM Tris, pH 8.1.
TE Buffer: 10mM Tris-HCL pH 8.1, 1 mM EDTA
Elution buffer: 1% SDS, 100mM NaHCO3
* Please note that protease inhibitors have variable half-lives and should be freshly prepared as applicable.
Western Blot Protocol
1. Perform SDS-PAGE on samples to be analyzed, loading 10 ug of histone preps per lane.
2. Transfer proteins to membrane according to the instructions provided by the manufacturer of the membrane and transfer apparatus.
3. Stain according to standard Ponceau S procedure (or similar product) to assess transfer success, and mark molecular weight standards where appropriate.
4. Rinse the blot.
5. Block the membrane using standard blocking buffer for at least 1 hour.
6. Wash the membrane in wash buffer three times for 10 minutes each.
7. Dilute primary antibody in blocking buffer and incubate 1 hour at room temperature.
8. Wash the membrane in wash buffer three times for 10 minutes each.
9. Apply the diluted HRP conjugated secondary antibody in blocking buffer (as per manufacturers instructions) and incubate 1 hour at room temperature.
10. Wash the blot in wash buffer three times for 10 minutes each (this step can be repeated as required to reduce background).
11. Apply the detection reagent of choice in accordance with the manufacturers instructions.
Note: Tween-20 can be added to the blocking or antibody dilution buffer at a final concentration of 0.05-0.2%.
Immunocytochemistry Protocol
Culture cells to appropriate density in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and add 10% formalin to the dish. Fix at room temperature for 30 minutes.
2. Remove the formalin and add ice cold methanol. Incubate for 5-10 minutes.
3. Remove methanol and add washing solution (i.e. PBS). Be sure to not let the specimen dry out. Wash three times for 10 minutes.
4. To block nonspecific antibody binding incubate in 10% normal goat serum from 1 hour to overnight at room temperature.
5. Add primary antibody at appropriate dilution and incubate at room temperature from 2 hours to overnight at room temperature.
6. Remove primary antibody and replace with washing solution. Wash three times for 10 minutes.
7. Add secondary antibody at appropriate dilution. Incubate for 1 hour at room temperature.
8. Remove antibody and replace with wash solution, then wash for 10 minutes. Add Hoechst 33258 to wash solution at 1:25,0000 and incubate for 10 minutes. Wash a third time for 10 minutes.
9. Cells can be viewed directly after washing. The plates can also be stored in PBS containing Azide covered in Parafilm (TM). Cells can also be cover-slipped using Fluoromount, with appropriate sealing.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow safe laboratory procedures.
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ChIP Protocol Video
- Chromatin Immunoprecipitation (ChIP) Protocol
- Chromatin Immunoprecipitation Protocol
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...

![Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024] Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024]](https://resources.rndsystems.com/images/products/Histone-H3-[ac-Lys4]-Antibody-Western-Blot-NB21-1024-img0017.jpg)
![Immunocytochemistry/ Immunofluorescence: Histone H3 [ac Lys4] Antibody - BSA Free [NB21-1024] Immunocytochemistry/ Immunofluorescence: Histone H3 [ac Lys4] Antibody - BSA Free [NB21-1024]](https://resources.rndsystems.com/images/products/Histone-H3-[ac-Lys4]-Antibody-Chromatin-Immunoprecipitation-NB21-1024-img0021.jpg)
![Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024] Western Blot: Histone H3 [ac Lys4] AntibodyBSA Free [NB21-1024]](https://resources.rndsystems.com/images/products/Histone-H3-[ac-Lys4]-Antibody-Western-Blot-NB21-1024-img0016.jpg)
![Histone H3 [ac Lys4] Antibody - BSA Free NB21-1024](https://resources.rndsystems.com/images/reviews/review_image_14181_5291_1380215317.JPG)