CXCL1/GRO alpha/KC/CINC-1 is an ELR+ angiogenic chemokine that interacts with CXCR2/IL-8 RB to attract and activate neutrophils. It is produced at sites of microbial infection by macrophages, monocytes, Kupffer cells, hepatocytes, endothelial and epithelial cells, cardiac and vascular smooth muscle cells, fibroblasts, melanocytes, mast cells, and neurons. CXCL1 function is associated with the development and extent of atherosclerosis, Alzheimer’s disease, and tumor angiogenesis.
Mouse CXCL1/KC Quantikine ELISA Kit
R&D Systems | Catalog # MKC00B


Key Product Details
Assay Length
Sample Type & Volume Required Per Well
Sensitivity
Assay Range
Product Summary for Mouse CXCL1/KC Quantikine ELISA Kit
Product Specifications
Measurement
Detection Method
Conjugate
Species
Specificity
Cross-reactivity
Interference
Precision
Intra-Assay Precision (Precision within an assay) Three samples of known concentration were tested on one plate to assess intra-assay precision.
Inter-Assay Precision (Precision between assays) Three samples of known concentration were tested in separate assays to assess inter-assay precision.
Cell Culture Supernates, Serum
| Intra-Assay Precision | Inter-Assay Precision | |||||
|---|---|---|---|---|---|---|
| Sample | 1 | 2 | 3 | 1 | 2 | 3 |
| n | 20 | 20 | 20 | 20 | 20 | 20 |
| Mean (pg/mL) | 34.9 | 169 | 530 | 35.0 | 183 | 521 |
| Standard Deviation | 1.9 | 5.2 | 26.3 | 2.1 | 18.0 | 15.6 |
| CV% | 5.4 | 3.1 | 5.0 | 6.0 | 9.8 | 3.0 |
Recovery for Mouse CXCL1/KC Quantikine ELISA Kit
The recovery of mouse KC spiked to three levels throughout the range of the assay in various matrices was evaluated.
| Sample Type | Average % Recovery | Range % |
|---|---|---|
| Cell Culture Supernates (n=7) | 98 | 89-103 |
| Serum (n=9) | 95 | 83-104 |
Linearity
To assess the linearity of the assay, five or more samples containing and/or spiked with various concentrations of mouse KC in each matrix were diluted with Calibrator Diluent and then assayed.

Scientific Data Images for Mouse CXCL1/KC Quantikine ELISA Kit
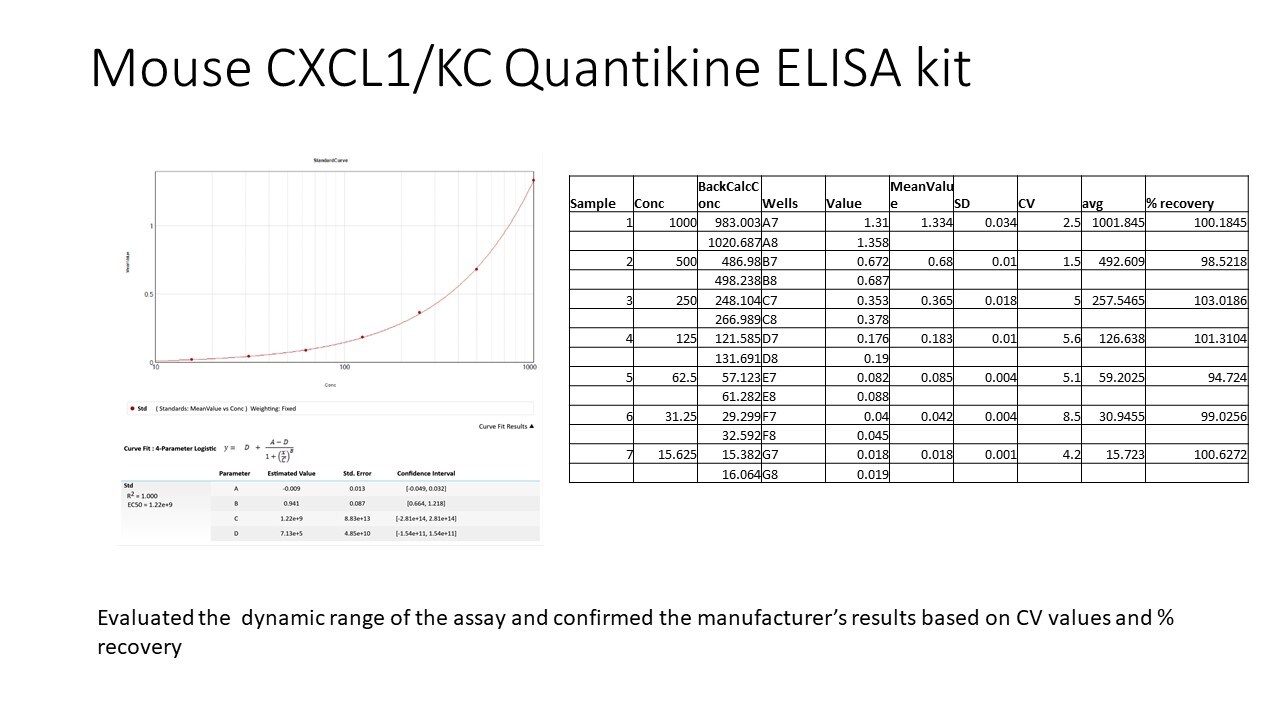
Mouse CXCL1/GRO alpha/KC/CINC-1 ELISA Standard Curve
Detection of Mouse CXCL1/GRO alpha/KC/CINC-1 by ELISA
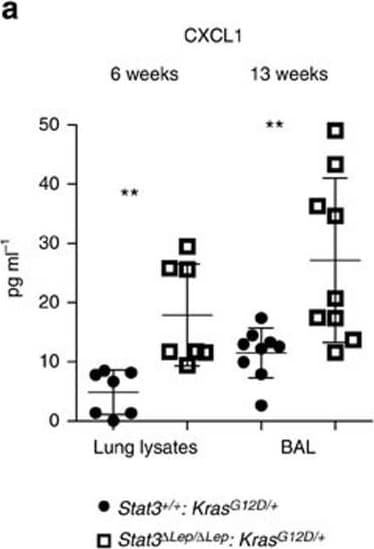
STAT3 regulates chemoattractive CXCL1 expression.(a) ELISA of CXCL1 levels in lung lysates and BAL at 6 and 13 weeks post AdCre. Data were analysed by Student’s t-test and shown as mean±s.d., n≥7 mice per genotype and time point. (b) Representative in situ hybridization with probes specific for murine Cxcl1 (AS) in lung tumours at 13 weeks post AdCre. Sense probes (S) were used as negative control and surfactant protein C (SP-C) as lung epithelial maker. Scale bar, 100 μm. (c) Kaplan–Meier plot showing overall survival of lung adenocarcinoma patient samples harbouring smoking history with high or low IL-8 mRNA expression (expression values of upper quartile, log-rank test) n≥35–104 per group. (d) Human A549 lung cancer cell line (KRASG12S mutated) was stimulated with designated cytokines for indicated time points. IL-8 or CXCL1 expression was analysed via qRT–PCR. Data were analysed by one-way ANOVA with Tukey’s multiple comparison test and shown as mean±s.d. (e) A549 cells transducted with lentiviral vectors expressing nonspecific scrambled small hairpin (sh)RNA or shRNA against STAT3 were stimulated with indicated cytokines for 60 min. Data were analysed by one-way ANOVA with Tukey’s multiple comparison test and shown as mean±s.d. (f) Primary pneumocytes were isolated and infected with AdCre. 120 h post infection cells were stimulated with indicated cytokines for 60 min. Cxcl1 expression was analysed by qRT–PCR. Data are shown as mean±s.e.m. (g) Murine 3T3 fibroblasts and STAT3 overexpressing 3T3 fibroblasts were stimulated as in f. Cxcl1 expression was analysed by qRT–PCR. Data were analysed by one-way ANOVA with Tukey’s multiple comparison test and shown as mean± s.d. Values in d–g are presented as fold change of relative mRNA expression compared with each unstimulated individual cell line. At least two independent experiments with three individual plates per stimulation were performed. For all graphs: *P<0.05; **P<0.01; ***P<0.001. Image collected and cropped by CiteAb from the following publication (https://www.nature.com/articles/ncomms7285), licensed under a CC-BY license. Not internally tested by R&D Systems.Preparation and Storage
Shipping
Stability & Storage
Background: CXCL1/GRO alpha/KC/CINC-1
Alternate Names
Gene Symbol
Additional CXCL1/GRO alpha/KC/CINC-1 Products
Product Documents for Mouse CXCL1/KC Quantikine ELISA Kit
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Mouse CXCL1/KC Quantikine ELISA Kit
For research use only
⚠ WARNING: This product can expose you to chemicals including N,N-Dimethylforamide, which is known to the State of California to cause cancer. For more information, go to www.P65Warnings.ca.gov.Related Research Areas
Citations for Mouse CXCL1/KC Quantikine ELISA Kit
Customer Reviews for Mouse CXCL1/KC Quantikine ELISA Kit (1)
Have you used Mouse CXCL1/KC Quantikine ELISA Kit?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Sample Tested: Recombinant proteinVerified Customer | Posted 10/26/2021
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- ELISA Sample Preparation & Collection Guide
- ELISA Troubleshooting Guide
- How to Run an R&D Systems DuoSet ELISA
- How to Run an R&D Systems Quantikine ELISA
- How to Run an R&D Systems Quantikine™ QuicKit™ ELISA
- Quantikine HS ELISA Kit Assay Principle, Alkaline Phosphatase
- Quantikine HS ELISA Kit Principle, Streptavidin-HRP Polymer
- Sandwich ELISA (Colorimetric) – Biotin/Streptavidin Detection Protocol
- Sandwich ELISA (Colorimetric) – Direct Detection Protocol
- Troubleshooting Guide: ELISA
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Associated Pathways