XBP1 Antibody - BSA Free
Novus Biologicals | Catalog # NBP1-77681

![Western Blot: XBP1 AntibodyBSA Free [NBP1-77681] Western Blot: XBP1 AntibodyBSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Western-Blot-NBP1-77681-img0006.jpg)
Key Product Details
Species Reactivity
Validated:
Cited:
Applications
Validated:
Cited:
Label
Antibody Source
Format
Product Specifications
Immunogen
Localization
Specificity
Clonality
Host
Isotype
Scientific Data Images for XBP1 Antibody - BSA Free
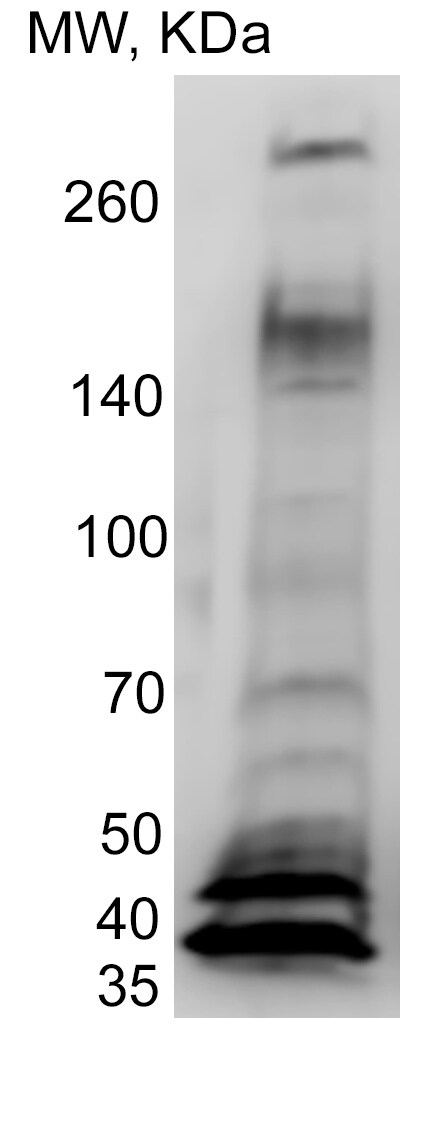
Western Blot: XBP1 AntibodyBSA Free [NBP1-77681]
Western Blot: XBP1 Antibody [NBP1-77681] - Total protein from HeLa cells was separated on a 12% gel by SDS-PAGE, transferred to PVDF membrane and blocked in 5% non-fat milk in TBST. The membrane was probed with 1.0 ug/ml anti-XBP1 in block buffer and detected with an anti-rabbit HRP secondary antibody using chemiluminescence. Arrow delineates XBP1s and arrowhead XBP1u.Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681]
Immunocytochemistry/Immunofluorescence: XBP1 Antibody [NBP1-77681] - Mouse MS1 cells were fixed in 4% paraformaldehyde for 10 minutes and permeabilized in 0.05% Triton X-100 in PBS for 5 minutes. The cells were incubated with XBP1 Antibody (NBP1-77681) at 1 ug/ml overnight at 4C and detected with an anti-rabbit DyLight 488 (Green) at a 1:1000 dilution for 60 minutes. Nuclei were counterstained with DAPI (Blue). Cells were imaged using a 40X objective.Immunohistochemistry: XBP1 Antibody - BSA Free [NBP1-77681]
Immunohistochemistry: XBP1 Antibody [NBP1-77681] - Staining of XBP1 in mouse bladder using DAB with hematoxylin counterstain.Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681]
Flow Cytometry: XBP1 Antibody [NBP1-77681] - An intracellular stain was performed on NIH3T3 cells with XBP1 Antibody NBP1-77681 (blue) and a matched isotype control NBP2-24891 (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 1.0 ug/mL for 30 minutes at room temperature, followed by Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Dylight 550 (SA5-10033, Thermo Fisher).Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681]
Immunocytochemistry/Immunofluorescence: XBP1 Antibody [NBP1-77681] - HeLa cells were fixed in 4% paraformaldehyde for 10 minutes and permeabilized in 0.05% Triton X-100 in PBS for 5 minutes. The cells were incubated with XBP1 Antibody (NBP1-77681) at 1 ug/ml overnight at 4C and detected with an anti-rabbit DyLight 488 (Green) at a 1:1000 dilution for 60 minutes. Nuclei were counterstained with DAPI (Blue). Cells were imaged using a 40X objective.Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681]
Flow Cytometry: XBP1 Antibody [NBP1-77681] - An intracellular stain was performed on HepG2 cells with XBP1 Antibody NBP1-77681 (blue) and a matched isotype control NBP2-24891 (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 1.0 ug/mL for 30 minutes at room temperature, followed by Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Dylight 550 (SA5-10033, Thermo Fisher).Western Blot: XBP1 Antibody [NBP1-77681] -
Western Blot: XBP1 Antibody [NBP1-77681] - Western Blot of PC-3 cell lysate. Image from verified customer review.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
ER stress inhibition impairs Zn (II)-curc-induced autophagy and mutp53 degradation. (a) Western blot analysis of BiP, LC3I/II, and p53 protein levels in U373 cells untreated or treated with Zn(II)-curc (100 ug/mL) for 24 h, with or without 1 h pre-treatment with 4-BPA (2.5 mM). (b) Densitometric analysis was performed using Image J software to calculate the ratio of the protein levels, as detected in (a), vs. beta -actin. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (c) Total mRNA was extracted from U373 cells untreated or treated as in (a). Spliced (s) and unspliced (u) Xbp1 gene expression were assayed by the PCR of reverse-transcribed cDNA. Densitometric analysis was performed using Image J software to calculate the Xbp1s/28S ratio. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32138264), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
Zn (II)-curc induces endoplasmic reticulum (ER) stress in mutant p53H273-carrying cells. (a) Representative photomicrographs of ER-Red Fluorescence staining in U373 cells untreated (Mock) or treated with Zn (II)-curc (100 ug/mL) for 16 h (Original magnification: 40×). (b) Quantization of ER content in U373 cells from ER-Red Fluorescence-stained cells. Mean fluorescence intensity (MFI) of each individual cell was normalized to cell size and expressed as fold-change compared with untreated cells at the same time point. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (c) Western blot analysis of p53, BiP, total (tot) IRE1 alpha, phosphorylated (p) IRE1 alpha, and XBP1 spliced (s) protein levels evaluated in U373 and HT29 cells untreated or treated with Zn (II)-curc (100 ug/mL) for 24 h. (d) Densitometric analysis was performed using Image J software to calculate the ratio of the protein levels, as detected in (c), vs. beta -actin. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (e) Total mRNA was extracted from U373 and HT29 cells untreated or treated with Zn (II)-curc (100 ug/mL) for 24 h. Spliced (s) Xbp1 gene expression was assayed by the polymerase chain reaction (PCR) of reverse-transcribed cDNA. Densitometric analysis was performed using Image J software to calculate the Xbp1s/28S ratio. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (f) p53 gene expression was assayed by PCR as in (e). The p53/28S ratio is indicated. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32138264), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
Mutp53 knockdown abrogates the Zn (II)-curc-induced ER stress. (a) U373 cells were transfected with control pSuper (si-ctr) and pSuper-p53 (si-p53) vectors for p53 knockdown and 36 h after transfection, cells were treated with Zn (II)-curc (100 ug/mL) for 16 h, before undergoing ER-Red Fluorescence staining. Quantization of ER content in U373 cells from ER-Red Fluorescence-stained cells as evaluated by the mean fluorescence intensity (MFI) of each individual cell normalized to cell size and expressed as fold-change compared with untreated cells at the same time point. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (b) Western blot analysis of BiP and p53 protein levels in U373 and HT29 cells transfected for 36 h with si-ctr and si-p53, and then treated with Zn (II)-curc (100 ug/mL) for 24 h. Densitometric analysis was performed using Image J software to calculate the ratio of BiP and p53 protein levels vs. beta -actin, as indicated ns: not specific signal. (c) Total mRNA was extracted from U373 cells treated as in (b), and spliced (s) and unspliced (u) Xbp1 gene expression were assayed by the PCR of reverse-transcribed cDNA. (upper panel) Densitometric analysis was performed using Image J software to calculate the Xbp1s/28S ratio. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32138264), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
Selective utilization of SHMT2 promoter 2 by HIF1 alpha and TFE3 complex.A, B Western blot analysis of SHMT2, SHMT2 alpha, XBP1, TFE3, HIF1 alpha protein expression upon XBP1, TFE3, HIF1 alpha knockdown respectively in cisplatin-sensitive (parental) and cisplatin-resistant (DDP) SKOV3 or A2780 cells under different cell culture. C Luciferase assay different promoter activity upon XBP1, TFE3, HIF1 alpha knockdown respectively cisplatin-resistant (DDP) SKOV3 or A2780 cells under low-glucose and hypoxic environment. D Co-IP experiment testing the interaction between TFE3, HIF1 alpha in cisplatin-resistant (DDP) SKOV3 or A2780 cells under low-glucose and hypoxic environment. E Binding pose of TFE3 with HIF1 alpha in 3D. F, G PLA analyze showing the distribution of TFE3 with HIF1 alpha in cisplatin-resistant (DDP) SKOV3 or A2780 cells under different cell culture. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/40097394), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
KSHV activates UPR and up-regulates PD-L1 on KSHV- infected macrophages.a Ire1 alpha and XBP1s expression in mock- and KSHV-infected macrophages was evaluated by western blot analysis; b ATF4, CHOP and BIP expression in mock- and KSHV-infected macrophages was evaluated by western blot analysis. beta -actin ( beta -ACT) was used as loading control. A representative experiment out of three is shown. Histograms represent the mean plus S.D. of the densitometric analysis of the ratio of each protein/ beta -ACT. *p-value < 0.05. c PD-L1 expression on mock- and KSHV-infected macrophages was evaluated by FACS analysis. A representative experiment is shown, and the mean of fluorescence intensity is indicated. Grey peaks represent the isotype controls. d Histograms representing the mean plus SD of PD-L1 MFI (Mean fluorescence Intensity) are also reported. *p-value < 0.05. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32418990), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] -
IRE1 alpha inhibition impairs Zn (II)-curc-induced autophagy and mutp53 degradation. (a) Total mRNA was extracted from U373 cells untreated or treated with Zn(II)-curc (100 ug/mL) for 24 h, with or without inhibitor of XBP1 cleavage STF-083010 (STF) (60 μM). Spliced (s) and unspliced (u) Xbp1 gene expression was assayed by the PCR of reverse-transcribed cDNA. (lower panel) Densitometric analysis was performed using Image J software to calculate the Xbp1s/28S ratio. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. (b) Western blot analysis of p53, BiP, and LC3I/II protein levels in U373 cells untreated or treated, as in (a). (c) Densitometric analysis was performed using Image J software to calculate the ratio of the protein levels, as detected in (b), vs. beta -actin. Histograms represent the mean +/- SD of three independent experiments. * p ≤ 0.05. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/32138264), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for XBP1 Antibody - BSA Free
Flow Cytometry
Immunocytochemistry/ Immunofluorescence
Immunohistochemistry
Immunohistochemistry-Paraffin
Knockdown Validated
Western Blot
Reviewed Applications
Read 1 review rated 5 using NBP1-77681 in the following applications:
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Formulation
Format
Preservative
Concentration
Shipping
Stability & Storage
Background: XBP1
Long Name
Alternate Names
Gene Symbol
UniProt
Additional XBP1 Products
Product Documents for XBP1 Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for XBP1 Antibody - BSA Free
Manufactured by Genomic Antibody Technology™. GAT FAQs
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Related Research Areas
Citations for XBP1 Antibody - BSA Free
Customer Reviews for XBP1 Antibody - BSA Free (1)
Have you used XBP1 Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Application: Western BlotSample Tested: PC-3 CellsSpecies: HumanVerified Customer | Posted 06/27/2023W-B of PC-3 cell lysate
There are no reviews that match your criteria.
Protocols
View specific protocols for XBP1 Antibody - BSA Free (NBP1-77681):
Immunocytochemistry Protocol
Culture cells to appropriate density in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and add 10% formalin to the dish. Fix at room temperature for 30 minutes.
2. Remove the formalin and add ice cold methanol. Incubate for 5-10 minutes.
3. Remove methanol and add washing solution (i.e. PBS). Be sure to not let the specimen dry out. Wash three times for 10 minutes.
4. To block nonspecific antibody binding incubate in 10% normal goat serum from 1 hour to overnight at room temperature.
5. Add primary antibody at appropriate dilution and incubate at room temperature from 2 hours to overnight at room temperature.
6. Remove primary antibody and replace with washing solution. Wash three times for 10 minutes.
7. Add secondary antibody at appropriate dilution. Incubate for 1 hour at room temperature.
8. Remove antibody and replace with wash solution, then wash for 10 minutes. Add Hoechst 33258 to wash solution at 1:25,0000 and incubate for 10 minutes. Wash a third time for 10 minutes.
9. Cells can be viewed directly after washing. The plates can also be stored in PBS containing Azide covered in Parafilm (TM). Cells can also be cover-slipped using Fluoromount, with appropriate sealing.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow safe laboratory procedures.
Immunohistochemistry-Paraffin Embedded Sections
Antigen Unmasking:
Bring slides to a boil in 10 mM sodium citrate buffer (pH 6.0) then maintain at a sub-boiling temperature for 10 minutes. Cool slides on bench-top for 30 minutes.
Staining:
1. Wash sections in deionized water three times for 5 minutes each.
2. Wash sections in wash buffer for 5 minutes.
3. Block each section with 100-400 ul blocking solution for 1 hour at room temperature.
4. Remove blocking solution and add 100-400 ul diluted primary antibody. Incubate overnight at 4C.
5. Remove antibody solution and wash sections in wash buffer three times for 5 minutes each.
6. Add 100-400 ul biotinylated diluted secondary antibody. Incubate 30 minutes at room temperature.
7. Remove secondary antibody solution and wash sections three times with wash buffer for 5 minutes each.
8. Add 100-400 ul Streptavidin-HRP reagent to each section and incubate for 30 minutes at room temperature.
9. Wash sections three times in wash buffer for 5 minutes each.
10. Add 100-400 ul DAB substrate to each section and monitor staining closely.
11. As soon as the sections develop, immerse slides in deionized water.
12. Counterstain sections in hematoxylin.
13. Wash sections in deionized water two times for 5 minutes each.
14. Dehydrate sections.
15. Mount coverslips.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow safe laboratory procedures.
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Liperfluo
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars

![Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681] Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Immunocytochemistry-Immunofluorescence-NBP1-77681-img0010.jpg)
![Immunohistochemistry: XBP1 Antibody - BSA Free [NBP1-77681] Immunohistochemistry: XBP1 Antibody - BSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Immunohistochemistry-NBP1-77681-img0004.jpg)
![Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681] Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Flow-Cytometry-NBP1-77681-img0008.jpg)
![Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681] Immunocytochemistry/ Immunofluorescence: XBP1 Antibody - BSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Immunocytochemistry-Immunofluorescence-NBP1-77681-img0009.jpg)
![Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681] Flow Cytometry: XBP1 Antibody - BSA Free [NBP1-77681]](https://resources.rndsystems.com/images/products/XBP1-Antibody-Flow-Cytometry-NBP1-77681-img0007.jpg)
![Western Blot: XBP1 Antibody [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/antibody/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-672023151929..jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-13220261534221.jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-132202615335815.jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-13220261530266.jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-13220261756120.jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-13220261739386.jpg)
![Western Blot: XBP1 Antibody - BSA Free [NBP1-77681] - XBP1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-77681_rabbit-polyclonal-xbp1-antibody-western-blot-13220261812225.jpg)