Cancer Biomarkers
Tools for Cancer Biomarker Discovery and Detection Research
Cancer biomarkers are critical indicators present in tumor cells, tissues, or body fluids that are used to detect cancer, track tumor progression, and assess treatment efficacy. These biomarkers encompass a wide range of molecules, including metabolites, DNA repair proteins, immune checkpoint molecules, cell surface receptors, and metastasis-associated enzymes, among others, that serve as targets for cancer research and diagnostics, and the advancement of precision medicine.
The demand for effective cancer biomarkers is growing rapidly because of their potential to reduce cancer mortality rates. For cancer patients, these biomarkers facilitate:
- Early cancer detection: Biomarkers help detect cancers at the earliest, most treatable stages, enabling timely and effective intervention.
- Tracking disease progression: Continuous monitoring of tumor cells and biomarker expression provides critical insights into cancer metastasis and tumor behavior.
- Customizing therapy: Biomarker testing supports personalized cancer treatment strategies, including targeted therapies and immunotherapies, improving treatment outcomes for each patient.
- Evaluating treatment response: Clinicians use cancer biomarkers to monitor how patients respond to chemotherapy, immunotherapy, and other targeted inhibitors, helping to adjust treatments in real time.
Recent advances in cancer biology and technology have led to the discovery of multiple new tumor markers. Technologies such as tissue microarrays, antibody arrays, mass spectrometry, and high-throughput sequencing have been instrumental in identifying diverse types of biomarkers, including genetic, proteomic, and metabolic markers. As new biomarkers are identified, rigorous validation through clinical trials is essential to confirm their efficacy and reliability in cancer care. With biomarker testing becoming increasingly refined and expanding across more cancer types, it continues to improve cancer diagnosis accuracy and advance precision medicine.
Featured Products for Cancer Biomarker Research
R&D Systems products support cancer researchers at every stage of the cancer biomarker development process. Our suite of research-use-only reagents, including rigorously validated proteins, antibodies, ELISAs, and multiplex immunoassays, is designed to provide consistent, reliable results on your path to discovering and validating new biomarkers.
Human Tumor Biomarker Luminex® Panel
Enhance early cancer detection, characterize the tumor microenvironment, and assess disease progression with the Human Tumor Biomarker Luminex Performance Assay. This innovative panel simultaneously detects 27 tumor biomarkers in 50 µL of sample.
Primary IHC Antibodies for Cancer Biomarkers
Get the format you need for your IHC workflow for cancer biomarker detection. Unconjugated, enzyme- or fluorophore-conjugated, biotinylated, and BSA/azide-free options support both direct and indirect detection.
ELISA Kits for Measuring Cancer Biomarkers
Quantitatively measure disease markers with the most cited ELISA Kits on market. From our ready-to-use Quantikine™ Kits and 90-minute QuicKits™ to our flexible DuoSet™ ELISAs, our immunoassays consistently deliver accurate, reproducible results every time.
Select Cancer Biomarkers in Clinical Practice
Select Tumor Suppressors
Select Oncogenic Drivers
Biomarkers for Different Types of Cancer
From bladder and breast cancers to pancreatic and colon cancers, each cancer type exhibits unique biological characteristics that are reflected in specific biomarkers. Our high-quality products reliably detect biomarkers in relevant sample types, supporting all stages of cancer biomarker analysis.
Bladder Cancer | Blood Cancers | Breast Cancer | Colorectal Cancer | Gastric Cancer | Glioma |
Head & Neck Cancer | Liver Cancer | Lung Cancer | Medulloblastoma | Melanoma | Osteosarcoma |
Ovarian Cancer | Pancreatic Cancer | Prostate Cancer
Bladder Cancer
Bladder cancer spans a diverse spectrum of tumor types, ranging from non-invasive urothelial lesions to highly aggressive, muscle-invasive forms. Biomarker research is crucial for advancing early detection and tracking disease progression. Key markers such as FGFR3 mutations, p53, and Ki67 offer prognostic insights into tumor behavior. Additionally, angiogenesis-related proteins and proteolytic enzymes including VEGF and MMP-2/MMP-9 provide perspectives on vascular changes, invasion, and metastasis during tumor growth. Immune markers including PD-L1 (CD274) and diagnostic markers like Uroplakin help refine strategies for personalized therapies. Explore the biomarker table below to see how these and other markers are being used for bladder cancer research.
Biomarker | Relevance to Bladder Cancer Research |
|---|---|
| Apolipoprotein A-I (ApoA1) | Altered expression linked to bladder cancer progression; Urinary biomarker detectable via ELISA; Used for non-invasive diagnosis and surveillance of tumor recurrence. |
| EpCAM / TROP1 | Cell adhesion molecule overexpressed in bladder cancer cells; Associated with tumor aggressiveness. |
| FGF R3 | Mutation/overexpression linked to low-grade, non-invasive bladder cancers; Aids in tumor classification and prognosis; Involved in cell proliferation signaling. |
| IL-8 (CXCL8) | Pro-inflammatory cytokine elevated in bladder cancer microenvironment; Promotes angiogenesis and tumor growth; Used as a non-invasive biomarker reflecting tumor activity. |
| Ki67 | Cell proliferation marker; High expression correlates with aggressive bladder cancer and poor prognosis. |
| MMP-2 | Matrix metalloproteinases involved in extracellular matrix degradation, facilitating invasion and metastasis; Indicates tumor invasiveness and progression. |
| MMP-9 | |
| p53 | Tumor suppressor gene frequently mutated in bladder cancer, especially high-grade tumors; Affects cell cycle control; Prognostic indicator and predictor of treatment response. |
| PD-L1 (CD274) | Immune checkpoint protein expressed on tumor cells; Contributes to immunosuppression in tumor microenvironment; Guides immunotherapy eligibility. |
| Survivin | Inhibitor of apoptosis overexpressed in bladder cancer; Associated with resistance to therapy; Serves as prognostic marker and potential therapeutic target. |
| Uroplakin II | Bladder-specific differentiation marker; Expressed in urothelial cells and carcinomas; Used in diagnosis and confirmation of urothelial origin of tumors. |
| VEGF | Key angiogenic factor promoting blood vessel formation in tumors; Elevated in bladder cancer microenvironment; Indicative of tumor angiogenesis and potential target for anti-angiogenic therapy. |
Blood Cancers
Blood cancers encompass a heterogeneous group of hematologic malignancies, including leukemias, lymphomas, and myelomas, each with distinct molecular and cellular profiles. Critical lineage markers (CD5, CD10, CD19, CD20, CD23, CD30, CD33, CD34, CD38, CD56, CD117, and CD138) serve as essential diagnostic tools for identifying and classifying malignant cell populations, while apoptosis regulators (Bcl-2) and transcription factors (Bcl-6) provide prognostic insight into disease progression in hematologic cancers. Clinically relevant mutations in signaling and regulatory proteins including Flt-3, NPM1, Jak2 (V617F), and p53, alongside emerging markers such as MyD88 (L265P), ALK, MYC, EZH2, and Cereblon (CRBN) pathway components, further refine prognosis and guide targeted therapeutic strategies. Together, these biomarkers enable precise disease classification and support the development of targeted therapies for the treatment of hematologic cancers. Learn more about blood cancer biomarkers in the table below.
Biomarker | Relevance to Blood Cancer Research |
|---|---|
| ALK/CD246 | Receptor tyrosine kinase; Defines subtypes of anaplastic large cell lymphoma; Targetable oncogene. |
| Bcl-2 | Anti-apoptotic protein promoting cell survival; Overexpressed in follicular lymphoma and other B-cell malignancies; Prognostic and therapeutic target. |
| Bcl-6 | Transcriptional repressor involved in germinal center B-cell development; Expressed in diffuse large B-cell lymphoma (DLBCL); Prognostic marker. |
| CD5 | Marker expressed on T cells and some B cells; Differentiates CLL (CD5+) from other B cell lymphomas like mantle cell lymphoma; Prognostic implications in CLL. |
| CD10/Neprilysin | Common lymphoid progenitor marker; Expressed in precursor B-cell ALL and certain lymphomas; Diagnostic utility. |
| CD19 | B cell surface antigen involved in B cell development and activation; Widely used in diagnosing B cell leukemias and lymphomas; Target for CAR-T cell therapies. |
| CD20 | Surface protein expressed on mature B cells; Diagnostic marker for B-cell non-Hodgkin lymphomas. |
| CD23 | Low-affinity IgE receptor; Used to distinguish CLL (CD23+) from mantle cell lymphoma (CD23-). |
| CD25/IL-2 R alpha | IL-2 receptor alpha chain; Involved in cell proliferation; Diagnostic marker for hairy cell leukemia. |
| CD30 | Activation marker expressed in activated lymphocytes; Diagnostic marker in Hodgkin lymphoma and anaplastic large cell lymphoma; Therapeutic target. |
| CD33 | Myeloid cell surface antigen; Marker for myeloid leukemias. |
| CD34 | Stem/progenitor cell marker; Used to identify hematopoietic stem and progenitor cells; Important in leukemia diagnosis and stem cell transplantation. |
| CD38 | Surface protein involved in cell adhesion and signaling; Highly expressed in multiple myeloma and some leukemias. |
| CD56/NCAM-1 | Neural cell adhesion molecule expressed on NK cells and some leukemias; Marker for NK cell malignancies and a subset of acute leukemias; Prognostic significance. |
| CD117/c-kit | Receptor tyrosine kinase important for hematopoietic stem cell survival; Expressed in AML and mast cell disorders; Used for diagnosis and prognostic stratification. |
| CD138 | Plasma cell marker, involved in cell adhesion and signaling; Critical plasma cell marker for multiple myeloma diagnosis and minimal residual disease detection. |
| Cereblon | Substrate receptor of E3 ubiquitin ligase complex; Predicts response to immunomodulatory drugs in multiple myeloma. |
| EZH2 | Histone methyltransferase promoting gene silencing and proliferation; Mutations found in follicular lymphoma and other lymphomas; Emerging target for therapy. |
| Flt-3/Flk-2 | Receptor tyrosine kinase involved in proliferation; Mutations common in AML associated with poor prognosis; Guide use of Flt-3 inhibitors. |
| IRF4 | Transcription factor involved in plasma cell differentiation; Marker for activated B-cell lymphomas and multiple myeloma. |
| Jak2 (V617F) | Mutant kinase causing constitutive activation of JAK-STAT pathway; Found in myeloproliferative neoplasms (MPNs) such as polycythemia vera; Diagnostic and therapeutic target. |
| Ki67 | Cell proliferation marker; Used to assess tumor proliferation rate; Higher levels predict aggressive disease. |
| MPO | Enzyme expressed in myeloid cells; Marker of myeloid lineage; Used in distinguishing AML from ALL. |
| MYC (c-Myc) | Transcription factor regulating cell growth and proliferation; Myc rearrangement/fusion associated with aggressive lymphomas such as Burkitt lymphoma and double-hit lymphomas. |
| NPM1 | Involved in ribosome biogenesis and DNA repair; Mutated in AML; Favorable prognostic marker and used in minimal residual disease detection. |
| p53 | Tumor suppressor gene regulating DNA repair and apoptosis; Loss or mutations linked to poor prognosis and therapy resistance across blood cancers. |
| TDT | Nuclear enzyme indicative of lymphoid precursor cells; Used in diagnosis of acute lymphoblastic leukemia (ALL). |
Breast Cancer
Breast cancer remains one of the most prevalent malignancies worldwide, with heterogeneity that necessitates precise molecular characterization for optimal treatment. Well-established markers such as Estrogen Receptor, Progesterone Receptor, and Her2 form the basis for understanding breast cancer subtypes and determining precision treatments. BRCA1/2 mutations are genetic and used to assess risk, while markers like Ki67 and p53 are used to determine the likelihood of tumor progression. Potential therapeutic targets, including immune checkpoint molecules such as PD-L1, and metastasis-associated factors like uPA, are being investigated. Learn more about these and other breast cancer biomarkers in the table below.
Biomarker | Relevance to Breast Cancer Research |
|---|---|
| AGR2 | Overexpressed in breast tumors; Involved in tumor growth, metastasis, and drug resistance; Potential prognostic biomarker. |
| ATAD2 | Oncogene involved in chromatin remodeling and transcription regulation; Overexpressed in aggressive tumors; Potential prognostic marker and therapeutic target. |
| BRCA1 | Tumor suppressor gene involved in DNA repair; Germline mutations confer high risk for hereditary breast cancer. |
| BRCA2 | |
| CA 15-3 (MUC1) | Circulating mucin antigen elevated in advanced breast cancer; Used for monitoring disease progression. |
| Caveolin-1 | Scaffold protein involved in cell signaling; Linked to tumor progression and stromal interactions, potential prognostic marker indicator. |
| EGFR | Receptor tyrosine kinase overexpressed in triple negative/basal-like tumors; Drives proliferation; Therapeutic target and prognostic marker. |
| Estrogen Receptor (ER) alpha | Nuclear hormone receptor; Major driver of luminal breast cancer; Predictive of hormone therapy response. |
| Her2/ErbB2 | Receptor tyrosine kinase amplified/overexpressed in ~20% of breast cancers; Associated with aggressive disease. |
| HIN-1 | Tumor suppressor gene often silenced in breast cancer; Involved in cell differentiation; Potential early detection and prognostic marker. |
| Ki67 | Proliferation marker; Higher levels correlate with aggressive tumors and poor prognosis. |
| Nestin | Intermediate filament protein; Marker of basal-like and stem-like breast cancer cells; Identifies aggressive subtypes, useful in classification. |
| p53 | Tumor suppressor commonly mutated in breast cancer, especially triple-negative subtype; Prognostic marker and predictor of therapy response. |
| PARP | Enzyme involved in DNA repair; especially relevant in BRCA-mutated cancers. Target for PARP inhibitors; Expression and mutation status guide therapy. |
| PD-L1 (CD274) | Immune checkpoint ligand; Expression linked to immune evasion in tumor microenvironment; Companion diagnostic for immunotherapy eligibility. |
| Progesterone Receptor (PR) | Nuclear hormone receptor; Usually co-expressed with ER; Predicts response to hormone therapy. |
| RHAMM (CD168) | Receptor involved in cell motility and metastasis; Overexpression linked to aggressiveness and poor prognosis. |
| Serpin E1 (PAI-1) | Inhibitor of plasminogen activators; High levels linked to poor prognosis due to effects on tumor invasion. Prognostic biomarker influencing treatment decisions. |
| u-Plasminogen Activator (uPA) | Serine protease promoting ECM degradation, invasion, and metastasis of breast cancer cells. Elevated levels serve as a prognostic biomarker and when measured alongside PAI-1, help tailor adjuvant chemotherapy. |
| WISP3 | Tumor suppressor frequently lost or mutated in aggressive breast cancers; Involved in cell adhesion and invasiveness control. |
| YAP1 | Transcriptional coactivator in the Hippo pathway; Regulates proliferation; Dysregulated activity linked to tumor growth and therapy resistance. |
Colorectal Cancer
Colorectal cancer (CRC) remains a leading cause of cancer morbidity and mortality worldwide, driven by complex genetic and molecular heterogeneity that challenges effective diagnosis and treatment. CRC biomarkers play a pivotal role in advancing early detection, prognosis, and treatment strategies. Core markers include mismatch repair proteins (MLH1, MSH2, MSH6, PMS2), essential for microsatellite instability (MSI) testing, and KRAS and BRAF mutations. Widely used serum markers like carcinoembryonic antigen (CEA) and angiogenesis regulators such as VEGF further support disease monitoring and therapy selection. Novel markers, including 14-3-3 beta, Dkk-1, and Reg4, provide further insights into cancer biology and progression. Learn more about these and other colorectal cancer biomarkers in the table below.
Biomarker | Relevance to Colon Cancer Research |
|---|---|
| 14-3-3 beta | Involved in cell cycle control and apoptosis; Deregulated expression linked to tumor progression and metastasis. |
| 14-3-3 zeta | Regulates signaling pathways influencing proliferation and apoptosis; Overexpressed in CRC; Involved in EMT, metastasis, and chemotherapy resistance. |
| A33 | Cell surface glycoprotein expressed in >95% of colorectal cancers, not in surrounding normal tissue; Target for monoclonal antibody therapies and diagnostic imaging. |
| ATAD2 | Co-activator of oncogenic transcription factors; Overexpressed in CRC; Promotes proliferation and invasion. |
| Carcinoembryonic Antigen (CEA) | Classic serum tumor marker; Elevated levels widely used to monitor colon cancer presence and recurrence. |
| CEACAM-5 | Classical serum tumor marker used for CRC diagnosis, monitoring treatment response, and detecting recurrence. |
| B-Raf (V600E) | Activating mutation in the MAPK signaling pathway; Common in some CRC subtypes associated with poor prognosis and aggressive disease; Molecular testing for mutation guides targeted therapy and prognosis stratification. |
| Dkk-1 | Wnt pathway antagonist; Dysregulated in CRC; Potential prognostic marker and therapeutic target. |
| GPRC5A | Orphan GPCR altered in CRC; Involved in proliferation and Wnt signaling; Emerging biomarker for tumor progression. |
| KRAS | Small GTPase involved in proliferation signaling; Frequently mutated in CRC, locking it in an active state; Driver of oncogenesis via activation of MAPK/ERK signaling. |
| MLH1 | Mismatch repair genes; Loss causes microsatellite instability (MSI), a hallmark of Lynch syndrome-associated and sporadic CRC. |
| MSH2 | |
| p53 | Tumor suppressor gene frequently mutated in CRC; Mutation correlates with genomic instability, aggressive cancer features, and treatment resistance. |
| Progranulin | Growth factor involved in cell proliferation, migration, and inflammation; Elevated expression in CRC linked to poor prognosis. |
| Reg4 | Secreted protein highly expressed in CRC; Involved in proliferation and chemoresistance; Studied as a diagnostic and prognostic marker. |
| Serum Amyloid A | Acute phase protein elevated in CRC; Associated with inflammation, tumor progression, and poor prognosis. |
| TMEM25 | Proposed tumor suppressor gene with reduced expression in CRC; May correlate with disease progression. |
| UVRAG | Tumor suppressor involved in autophagy and DNA repair; Mutations/downregulation linked to CRC progression and genomic instability. |
| VEGF | Frequently overexpressed in CRC; Crucial for angiogenesis and tumor vascularization; Used as a target in anti-angiogenic therapies. |
Gastric Cancer
As one of the leading contributors to global cancer mortality, gastric cancer remains a focus of biomarker discovery to improve diagnosis and treatment. Commonly studied markers like CEA, CA 19-9, and CA 72-4 remain central to disease detection and monitoring, while Her2 and p53 provide insights into tumor aggression and response to targeted therapy. Research into immune regulators such as PD-L1, angiogenic factors like VEGF, and cancer cell-specific biomarkers such as Claudin-18 is uncovering promising therapeutic targets. Learn more about these and other gastric cancer biomarkers in the table below.
Biomarker | Relevance to Gastric Cancer Research |
|---|---|
| ARID1A | Chromatin remodeling and tumor suppressor gene frequently mutated in gastric cancer; ARID1A mutations are associated with DNA damage repair defects/microsatellite instability (MSI) and tumor progression. |
| ATAD2 | Oncogene overexpressed in gastric cancer; Linked with tumor progression, poor prognosis, and enhanced cell proliferation activities. |
| Carcinoembryonic Antigen (CEA) | Cell adhesion molecule elevated in gastric cancer patients, especially advanced stages; Used for diagnosis and monitoring recurrence. |
| CA19-9 | Glycolipid antigen; Elevated serum levels linked with advanced and metastatic disease. |
| Claudin-18 | Plays a crucial role in cell-cell adhesion as a tight junction protein; Overexpressed in certain types of gastric cancer; Selective and relatively restricted expression of isoform 2 in gastric cancer cells, makes it an attractive, highly-specific therapeutic target. |
| COX-2 | Enzyme involved in inflammation and tumorigenesis; Overexpressed in gastric cancer; Promotes angiogenesis and tumor growth; Potential therapeutic target. |
| DMBT1 | Tumor suppressor gene downregulated in gastric cancer; Involved in mucosal protection and immune response; Possible biomarker for early detection. |
| E-Cadherin | Tumor suppressor crucial for cell adhesion; Mutations or loss commonly found in diffuse-type gastric cancer, linked to tumor invasion. |
| EGFR | Receptor tyrosine kinase overexpressed or amplified in some gastric cancers; Contributes to proliferation and metastasis; Investigated as a therapeutic target. |
| Gastrokine 1 | Gastrointestinal mucosal protective factor frequently lost in gastric cancer. |
| Her2/ErbB2 | Receptor tyrosine kinase overexpressed or amplified in some gastric cancers; Contributes to proliferation and metastasis; Investigated as a therapeutic target. |
| MMP-2 | Matrix metalloproteinases involved in extracellular matrix degradation, facilitating invasion and metastasis; Indicates tumor invasiveness and progression. |
| MMP-9 | |
| p53 | Tumor suppressor gene frequently mutated in gastric cancer; Associated with poor prognosis. Diagnostic and prognosis marker. |
| PD-L1 (CD274) | Immune checkpoint ligand; Expression linked to immune evasion in tumor microenvironment; Predictive biomarker for immunotherapy response in gastric cancer. |
| SOX17 | Transcription factor with tumor-suppressing functions; Down-regulated by promoter methylation in gastric cancer. |
| TFF2 | Gastric-specific mucosal protein; Reduced expression linked to gastric cancer; Possible marker for early gastric mucosal changes. |
| TLR2 | Pattern recognition receptor involved in tumor-promoting inflammation; Altered expression implicated in gastric cancer. |
| VEGF | Angiogenic factor; Overexpression correlates with tumor vascularization and poor prognosis; Target for anti-angiogenic therapy. |
| VSIG1 | Epithelial marker specific to gastric tissue; Diagnostic marker distinguishing gastric cancer subtypes. |
Glioma
Gliomas constitute a heterogeneous group of primary brain tumors originating from glial cells, including astrocytes, oligodendrocytes, or ependymal cells. They are distinguished from other brain tumors by core diagnostic markers such as GFAP and neuron-specific enolase (NSE). Additional biomarkers, including Ki67 and p53, are used as markers of proliferation and genetic instability, respectively. Molecular biomarkers critical for glioma subclassification include IDH1/2 mutations, ATRX loss, and 1p/19q co-deletion. Among the glioma subclasses, glioblastoma (GBM) is the most aggressive and prevalent malignant glioma, arising from astrocytic lineage cells. GBM is classified as a WHO grade IV astrocytoma, reflecting its high malignancy and poor prognosis. Tumor pathogenesis and proliferation in GBM is frequently driven by EGFR amplification or mutation and TERT promoter mutations, while VEGF and PDGF receptor alpha play key roles in tumor angiogenesis and vascular proliferation. More details on glioma biomarkers are provided in the table below.
Biomarker | Relevance to Glioma Research |
|---|---|
| ASCL1/Mash1 | Transcription factor regulating neural progenitor differentiation; Used to identify neural stem-like tumor cells; Linked to glioblastoma progression and chemoresistance. |
| ATRX | Chromatin remodeling protein, involved in telomere maintenance; Frequently mutated in astrocytomas and IDH-mutant gliomas; Loss linked to alternative lengthening of telomeres; Used for molecular classification. |
| Cathepsin D | Lysosomal protease involved in protein degradation; Overexpressed in gliomas; Linked with invasive behavior and tumor progression. |
| CD31/PECAM-1 | Endothelial cell adhesion molecule; Marker of microvascular density in gliomas; Increased in glioblastoma due to angiogenesis; Used to assess tumor vascularization and angiogenesis. |
| CD44 | Cell surface adhesion molecule; Overexpresed in glioma stem-like cells; Marker for stemness and invasiveness; Linked to invasion, migration, and therapy resistance. |
| Chitinase 3-like 1/YKL40 | Glycoprotein involved in inflammation and tissue remodeling; Overexpressed in high-grade gliomas; Serum and tissue biomarker for tumor progression and prognosis; Correlates with poor prognosis, invasion, and glioma recurrence. |
| DLL3 | Notch pathway ligand; Overexpressed in some gliomas; Potential therapeutic target. |
| EGFR | Receptor tyrosine kinase driving cell proliferation and survival; Frequently amplified/mutated in glioblastoma; Therapeutic target for EGFR inhibitors and a marker of aggressive disease. |
| ELTD1 | GPCR involved in angiogenesis regulation; Overexpressed in glioblastoma vasculature, contributing to tumor blood vessel formation; Potential therapeutic target. |
| FABP5 | Fatty acid-binding protein involved in lipid metabolism and signaling; Elevated in glioblastoma; Linked to tumor proliferation, metabolic adaptation, and therapeutic resistance. |
| FoxM1 | Transcription factor regulating cell cycle progression and mitosis; Overexpressed in glioblastoma; Promotes tumor proliferation and survival; Associated with aggressive phenotype. |
| GFAP | An intermediate filament protein specific to astrocytes; Routinely used as an immunohistochemical marker to confirm the astrocytic lineage of gliomas, aiding in the diagnosis of astrocytomas in clinical pathology. |
| IDH1 | Metabolic enzymes; IDH1 mutations are common in lower-grade gliomas and impact treatment decisions; IDH2 mutations are less common in glioma but similarly confer prognostic significance; Used to define distinct molecular subtypes. |
| IDH2 | |
| Ki67 | Proliferation marker; High index correlates with tumor grade and aggressiveness; Prognostic marker; Used in grading. |
| MMP-9 | Matrix metalloproteinase; Facilitates tumor invasion through extracellular matrix degradation; Marker of invasiveness. |
| Nestin | Intermediate filament protein expressed in neural stem/progenitor cells; Marker of undifferentiated glioma stem-like cells; Associated with aggressive tumor behavior. |
| Neuron-specific Enolase | Glycolytic enzyme specific to neurons; May be elevated in some gliomas; Supplementary diagnostic marker. |
| p53 | Tumor suppressor involved in DNA repair; Mutated in variable subsets of gliomas, associated with genomic instability and poor prognosis. |
| PDGF R alpha | Receptor tyrosine kinase for PDGF; Overexpressed/amplified in gliomas; Promotes proliferation; Potential therapeutic target and diagnostic marker. |
| VEGF | Promotes angiogenesis, enabling tumor growth and metastasis; Target for anti-angiogenic therapy. |
Head and Neck Cancer
Head and neck cancers (HNC) encompass a wide range of malignancies arising from the oral cavity, pharynx, and larynx, with diverse biomarker profiles that influence their study and treatment. Key protein biomarkers such as p16INK4a and EGFR are used to identify HPV-driven and EGFR-amplified tumors, respectively. Proliferative markers, including Cyclin D1 and p53, and angiogenic regulators such as VEGF are being investigated to understand how these tumors grow and respond to therapy. IL-6, IL-8, and MMPs are additional biomarkers being used to investigate the inflammatory and proteolytic pathways associated with tumor progression. Learn more about head and neck cancer biomarkers in the table below.
Biomarker | Relevance to Head and Neck Cancer Research |
|---|---|
| 14-3-3 sigma | Acts as a tumor suppressor by inducing cell cycle arrest; Down-regulation is associated with tumor progression and poor prognosis in certain types of head and neck cancers; Used as a biomarker for detecting malignancy and predicting outcomes. |
| AMFR (gp78) | Receptor involved in endoplasmic reticulum-associated degradation; Overexpressed in some head and neck cancers and associated with increased tumor invasion and metastasis; Considered a biomarker for aggressive tumor phenotypes and poor clinical outcomes. |
| BMI-1 | Polycomb group protein involved in stem cell renewal and oncogenesis; Overexpression linked to poor prognosis, tumor growth, and therapy resistance; Used as a marker for cancer stem cells and tumor aggressiveness. |
| Cathepsin D | Lysosomal protease involved in protein degradation and apoptosis; Elevated in some head and neck cancers; Contributes to tumor invasion and metastasis; Used as a marker of tumor aggressiveness. |
| CRYAB | Molecular chaperone; Protects cells from stress and apoptosis; Overexpressed in head and neck cancers and linked to treatment resistance; Potential marker for poor response to treatment. |
| Cyclin D1 (CCND1) | Regulates G1 to S phase transition in the cell cycle; Frequently amplified/overexpressed in head and neck cancers; Used as a diagnostic marker and indicator of dysregulated proliferation. |
| Cytokeratin 19 | Intermediate filament in epithelial cells; Fragments found in serum (CYFRA 21-1) serve as tumor markers for HNC; Used for detection and monitoring disease progression. |
| EGFR | Receptor tyrosine kinase stimulating proliferation and survival; Commonly overexpressed/mutated in head and neck cancers; Diagnostic marker and therapeutic target. |
| FGF R3 | Receptor tyrosine kinase in cell differentiation and proliferation; FGF R3 mutations or overexpression is associated with a subset of head and neck cancers; Potential diagnostic and therapeutic marker. |
| Galectin-3BP | Involved in cell-cell and cell-matrix interactions; Elevated levels linked to tumor progression and metastasis; Considered a biomarker for diagnosis and prognosis in head and neck cancers. |
| HGF R | Receptor tyrosine kinase that controls cell growth, motility, and morphogenesis; Overexpressed in head and neck cancers; Drives tumor growth, invasion, and metastasis; Used as a prognostic marker and a therapeutic target. |
| IL-6 | Pro-inflammatory cytokine; Elevated serum levels in head and neck cancers; Associated with poor prognosis and tumor progression. |
| IL-8 | Pro-inflammatory chemokine promoting angiogenesis and neutrophil recruitment; Elevated in HNC; Correlates with angiogenesis, invasion and metastasis; Used as a prognostic marker. |
| MMP-1 | Matrix metalloproteinases involved in degradation of extracellular matrix components; Many MMPs are overexpressed in aggressive head and neck cancers; Regulators of tumor invasion and metastasis; Used to assess tumor aggressiveness. |
| MMP-2 | |
| MMP-3 | |
| MMP-9 | |
| MMP-10 | |
| Notch | Transmembrane receptor and transcription factor that regulates cell differentiation, proliferation, and apoptosis; Tumor suppressor in HPV-negative head and neck cancers; Tumor promoter in HPV-positive/metastatic head and neck cancers; Used for molecular classification and prognosis. |
| p16INK4a | Tumor suppressor; Inhibits CDK4/CDK6 to control the cell cycle; Overexpression indicative of HPV-driven oropharyngeal cancers; Used as a diagnostic and prognostic biomarker. |
| p53 | Tumor suppressor that regulates DNA repair and apoptosis; Frequently mutated in head and neck cancers; Loss of function leads to genomic instability; Used as a biomarker for malignancy and prognosis. |
| S100A9 | Calcium-binding protein involved in inflammation and immune regulation; Frequently overexpressed in head and neck cancers; Supports tumor growth and immune evasion. |
| Serum Amyloid A | Acute-phase protein involved in inflammation; Elevated in aggressive head and neck cancers and linked to poor prognosis; Potential serum biomarker reflecting tumor-associated inflammation. |
| SPARC | Modulates cell-matrix interactions and tissue remodeling; Overexpressed in some head and neck cancers; Associated with invasion and poor prognosis; Potential diagnostic marker. |
| VEGF | Promotes angiogenesis; Highly expressed in head and neck cancers; Correlates with tumor vascularization, aggressiveness, and poor prognosis; Targeted in anti-angiogenic therapies. |
Liver Cancer
Liver cancer, predominantly hepatocellular carcinoma (HCC), is one of the leading causes of cancer-related deaths worldwide, driven by its aggressive nature and frequent late stage diagnoses. Markers like Alpha-Fetoprotein (AFP) and Glypican-3 serve as critical diagnostic tools, while Des-gamma-carboxy prothrombin (DCP) and CYFRA 21-1 are used for disease monitoring and predicting risk of recurrence. Additionally, VEGF is used as a marker of vascular changes that drive tumor growth, and MMPs (MMP-2, MMP9) provide insights into tumor invasion and metastasis. Find more details on liver cancer biomarkers in the table below.
Biomarker | Relevance to Liver Cancer Research |
|---|---|
| Alpha-Fetoprotein (AFP) | Tumor-associated antigen; Widely used serum biomarker of HCC; Used for diagnosis and monitoring treatment efficacy. |
| ATAD2 | Oncogene involved in chromatin remodeling and transcription regulation; Overexpressed in HCC and linked to tumor proliferation and poor prognosis; Emerging biomarker and potential therapeutic target. |
| CTNNB1 (beta-Catenin) | Wnt pathway transcriptional co-activator; Frequently mutated/activated in HCC; Aberrant Wnt/beta-Catenin signaling promotes tumor growth and progression. |
| Dkk-1 | Wnt pathway antagonist; Elevated in HCC and used to distinguish HCC from benign liver diseases; Used to complement AFP testing. |
| Glypican-3 | Overexpressed in HCC but not in normal liver tissue; Plays a role in tumor growth; Therapeutic target for HCC. |
| Cytokeratin 19 fragment (CYFRA 21-1) | Intermediate filament protein fragment released during cell turnover; Elevated in a subset of HCC with cholangiocarcinoma features; Reflects tumor burden and aggressiveness. |
| Lamin B1 | Nuclear envelope structural protein; Overexpressed in HCC and correlates with tumor cell proliferation and aggressiveness. |
| MMP-2 | Matrix metalloproteinases involved in degradation of extracellular matrix components; Increased expression in HCC; Associated with angiogenesis, metastasis, and poor prognosis. |
| MMP-9 | |
| PD-L1 (CD274) | Immune checkpoint ligand; Overexpressed in some HCCs; Levels correlate with immune evasion; Predictive biomarker for immune checkpoint inhibitor therapy. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Overexpressed in HCC; Serum levels used as prognostic marker; Target of anti-angiogenic therapies. |
Lung Cancer
Lung cancer is the leading cause of cancer deaths worldwide, driven by its two main subtypes, non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC), each requiring distinct diagnostic and therapeutic strategies. To address this complexity, biomarker profiling is crucial for early diagnosis, guiding targeted therapies, and tracking response to treatment. Key markers like EGFR, ALK, and PD-L1 are critical for identifying actionable targets in NSCLC, while p53 and Neuron-specific Enolase (NSE) are commonly associated with SCLC and play important prognostic roles. Additional diagnostic biomarkers, including Carcinoembryonic Antigen (CEA), CYFRA 21-1, and Napsin A, support tumor classification and disease monitoring, while angiogenic indicators such as VEGF shed light on tumor vascularization and metastasis. Find more details on lung cancer biomarkers in the table below.
Biomarker | Relevance to Lung Cancer Research |
|---|---|
| ALK/CD246 | Receptor tyrosine kinase involved in cell growth and differentiation; Fusion mutations define a subset of non-small cell lung cancers (NSCLC); Biomarker for diagnosis and targeted therapy with ALK inhibitors. |
| ATAD2 | Oncogene involved in chromatin remodeling and transcription regulation; Overexpressed in lung cancer, particularly NSCLC; Associated with tumor growth, metastasis, and poor prognosis. |
| Carcinoembryonic Antigen (CEA) | Cell adhesion protein; Elevated in many lung adenocarcinomas and other cancers; Commonly used serum tumor marker for diagnosis, monitoring treatment response, and detecting recurrence. |
| CEACAM-5 | Member of CEA family, involved in cell adhesion and signaling; Widely used tumor marker; Overexpressed on lung adenocarcinoma cells; Closely linked to CEA for detection and monitoring. |
| Cytokeratin 19 fragment (CYFRA 21-1) | Intermediate filament protein fragment released during epithelial cell turnover; Commonly elevated in NSCLC; Widely used serum biomarker for diagnosis, monitoring treatment response, and detecting recurrence. |
| EGFR | Receptor tyrosine kinase that regulates cell proliferation and survival; Frequently mutated/overexpressed in NSCLC, particularly adenocarcinoma; Predictive biomarker for EGFR inhibitor therapies. |
| KRAS | Small GTPase involved in proliferation signaling; Frequently mutated in NSCLC, locking it in an active state; Driver of oncogenesis via activation of MAPK/ERK signaling. |
| Lactate dehydrogenase A | Enzyme in glycolysis that converts pyruvate to lactate; Frequently upregulated in lung cancer cells due to metabolic reprogramming; Elevated serum levels correlate with aggressive tumors and poor prognosis. |
| Napsin A | Aspartic protease involved in protein processing, highly expressed in lung alveolar cells and in lung adenocarcinoma; Aids in distinguishing lung adenocarcinoma from other lung cancer subtypes. |
| Neuron-specific Enolase | Glycolytic enzyme isoform expressed in neurons and neuroendocrine cells; Elevated in small cell lung cancer (SCLC); Serum biomarker used for SCLC diagnosis, monitoring treatment response, and detecting recurrence. |
| p53 | Tumor suppressor regulating DNA repair and apoptosis; Mutated in many lung cancers leading to loss of cell cycle control and genomic instability; Used as a prognostic marker. |
| PD-L1 (CD274) | Immune checkpoint ligand that inhibits T cell activation; Expressed on tumor cells and cells in the tumor microenvironment; Predictive biomarker for response to immune checkpoint inhibitors. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Frequently overexpressed in lung tumors; Promotes neovascularization facilitating tumor growth; Serum levels used as prognostic marker; Target of anti-angiogenic therapies. |
Medulloblastoma
Medulloblastoma is a complex and aggressive pediatric brain tumor, categorized into distinct molecular subgroups (WNT, SHH, Group 3, and Group 4), each with varying prognoses and responses to therapy. SHH pathway proteins (SHH, GLI-1, PTCH1) and beta-Catenin (CTNNB1) are markers used to differentiate molecular subtypes, while c-Myc and p53 are critical for understanding tumor aggressiveness and prognosis. Ki67 and VEGF are markers that provide insights into tumor growth and vascularization, respectively, while additional targets, including MMP-2, Otx2, and MMP-9, reveal pathways driving invasion and migration. Details on these and other medulloblastoma biomarkers can be found in the table below.
Biomarker | Relevance to Medulloblastoma Research |
|---|---|
| Apolipoprotein E (ApoE) | Involved in lipid metabolism; Implicated in medulloblastoma progression via tumor metabolic alterations. |
| Clusterin | Chaperone protein involved in apoptosis regulation and cell survival; Overexpressed in medulloblastoma; Contributes to tumor progression and chemoresistance. |
| CTNNB1 (beta-Catenin) | Wnt pathway transcriptional co-activator; Mutations/accumulation indicate Wnt subtype; Used in molecular subgroup classification. |
| FGF-basic | Growth factor that promotes cell proliferation and angiogenesis; Potential marker of aggressive tumors. |
| GLI-1 | Regulator of the SHH pathway promoting cell proliferation; Mutations or altered expression identify SHH-driven tumors. Used for subtype classification. |
| IGFBP-2 | Modulates IGF signaling, enhances cell proliferation; Frequently overexpressed in medulloblastoma; Elevated levels correlate with tumor growth. |
| IGFBP-3 | Regulates IGF bioavailability, can promote apoptosis; Altered expression has been observed in medulloblastoma and levels can reflect tumor behavior; Used in prognostic assessment. |
| Ki67 | Marker of cellular proliferation; Widely used to assess tumor proliferative index; Higher expression indicates aggressive tumors. |
| Lipocalin-2 (NGAL) | Involved in iron transport and immune response; Elevated levels associated with tumor invasion and poor prognosis. |
| MMP-2 | Matrix metalloproteinases involved in degradation of extracellular matrix components; Markers of invasiveness; Higher expression indicates aggressive phenotype. |
| MMP-9 | |
| MYC (c-Myc) | Transcription factor regulating cell growth and proliferation; Amplification associated with high-risk medulloblastoma subtype. |
| Netrin-1 | Guidance cue protein involved in axon pathfinding; Affects cell migration and survival; Overexpressed in certain medulloblastoma subtypes correlates with tumor progression and invasiveness. |
| Osteopontin | Glycoprotein involved in cell adhesion and migration; Highly expressed in medulloblastoma and associated with tumor aggressiveness. |
| Otx2 | Transcription factor overexpressed in some medulloblastomas subgroups (excluding Wnt subgroup); Useful marker for molecular classification. |
| p53 | Tumor suppressor regulating DNA repair and apoptosis; Mutations linked to poor prognosis. |
| PTCH1 | Regulator of the SHH pathway promoting cell proliferation; Mutations or altered expression identify SHH-driven tumors. Used for subtype classification. |
| SHH | Regulator of the SHH pathway promoting cell proliferation; Mutations or altered expression identify SHH-driven tumors. Used for subtype classification. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Overexpression correlates with increased vascularity and tumor growth; Target of anti-angiogenic therapies. |
| YAP1 | Effector of Hippo pathway, promotes cell proliferation and survival; Key oncogenic driver in WNT subgroup and some SHH subtype medulloblastomas; YAP1 expression or nuclear localization serves as both a diagnostic and prognostic marker. |
Melanoma
Melanoma is an aggressive and often therapy-resistant form of skin cancer, marked by significant molecular diversity that impacts its progression and response to treatment. Genetic alterations such as B-Raf (V600E) mutations and loss or mutation of p16INK4a are vital for classifying melanoma subtypes, while Ki67 and p53 provide insights into tumor proliferation and genetic instability, respectively. Additionally, Melan-A (MART-1) and S100B, are important markers of the melanocyte lineage that are both widely used to confirm the presence of melanoma. Immune-focused biomarkers, including PD-L1 (CD274) and CD8, are aimed at investigating tumor-immune cell interactions in the tumor microenvironment (TME). Assessment of these markers helps guide immunotherapy development and predict response. Find more details on melanoma biomarkers in the table below.
Biomarker | Relevance to Melanoma Research |
|---|---|
| AIM2 | Component of the inflammasome; Regulates immune response and tumor suppression; Down-regulation associated with melanoma progression and immune evasion. |
| Akt | Key kinase in PI3K/Akt pathway promoting survival, proliferation, and metabolism; Frequently upregulated in melanoma, promoting tumor growth and therapy resistance. |
| B-Raf (V600E) | Mutated serine/threonine kinase driving MAPK pathway activation; Most common oncogenic driver mutation in melanoma; Diagnostic and therapeutic target. |
| CD8 | Marker for cytotoxic T cells; Used to evaluate tumor-infiltrating lymphocytes; Predictor of immune response and prognosis. |
| CD44 | Cell surface glycoprotein mediating adhesion and migration; Associated with melanoma stem cell-like properties and metastatic potential. |
| CEACAM-1/CD66a | Cell adhesion molecule involved in immune evasion and metastasis; Correlates with immune suppression and melanoma progression; Potential prognostic marker. |
| CTNNB1 (beta-Catenin) | Wnt pathway transcriptional co-activator; Nuclear β-catenin is linked with melanoma progression; Involved in immune evasion. |
| CXCR4 | Chemokine receptor promoting metastasis by regulating migration and homing to the lymph nodes and lungs; Target for anti-metastatic therapies. |
| E-Cadherin | Cell-cell adhesion molecule and tumor suppressor; Loss of E-Cadherin is a hallmark of EMT and promotes melanoma invasion and metastasis. |
| EZH2 | Histone methyltransferase promoting gene silencing and proliferation; Frequently overexpressed in advanced melanoma; Therapeutic target and prognostic indicator. |
| FGF-basic | Growth factor promoting cell proliferation and angiogenesis; Elevated levels support tumor growth and neovascularization. |
| FXYD5/Dysadherin | Modulates cell adhesion and enhances metastatic behavior; Linked to melanoma invasiveness and poor prognosis. |
| HIF-2 alpha | Hypoxia-inducible factor promoting angiogenesis and metabolic adaptation; Marker of hypoxic tumor regions and aggressiveness. |
| HLA-DR | MHC class II molecule involved in antigen presentation; Expression induced on melanoma cells by IFNs; Associated with immune evasion. |
| HSP90 | Molecular chaperone involved in stabilizing numerous oncogenic proteins; Overexpressed in melanoma; Target for HSP90 inhibitors. |
| ID1 | Transcriptional regulator involved in proliferation and inhibition of differentiation; Frequently overexpressed in melanoma; Associated with tumor aggressiveness and therapy resistance. |
| Integrin beta 1 | Cell surface receptor mediating adhesion, migration, and signaling; Promotes invasion and metastasis. |
| Integrin beta 3 | Cell adhesion receptor involved in angiogenesis and metastasis; Marker of melanoma progression. |
| JNK | Stress-activated protein kinase with dual role in apoptosis and survival; Elevated activity linked to apoptosis resistance and melanoma progression. |
| Ki67 | Cell proliferation marker; Widely used to assess proliferation rate; High levels correlate with aggressive melanoma |
| Lactate dehydrogenase A | Enzyme in glycolysis that converts pyruvate to lactate; High serum LDH levels are a strong prognostic marker in metastatic melanoma, with elevated levels correlating with increased tumor burden, aggressive disease, and reduced survival. |
| MCAM/CD146 | Cell adhesion molecule promoting angiogenesis and metastasis; Highly expressed in melanoma and correlated with poor prognosis and metastatic potential. |
| Melan-A (MART-1) | Melanocyte differentiation antigen; Diagnostic marker for melanoma cells. |
| MITF | Master regulator of melanocyte differentiation and melanoma survival; Critical marker for melanoma lineage; Regulates proliferation and pigmentation. |
| MMP-2 | Matrix metalloproteinase involved in degradation of extracellular matrix components; Marker of invasiveness; Higher expression indicates aggressive phenotype. |
| MYC (c-Myc) | Oncogenic transcription factor that promotes proliferation; Overexpression drives melanoma growth; Prognostic indicator. |
| NM23-H1/NH23-H2 | Metastasis suppressor proteins; Reduced expression correlates with increased metastatic potential. |
| p16INK4a | Tumor suppressor; Inhibits CDK4/CDK6 to control the cell cycle; Frequently mutated or lost in melanoma; Loss predicts poor prognosis. |
| p21 | Tumor suppressor; Dysregulation alters cell cycle control and promotes melanoma progression. |
| p53 | Tumor suppressor regulating DNA repair and apoptosis; Rarely mutated in melanoma but frequently inactivated; Role in prognosis. |
| PD-L1 (CD274) | Immune checkpoint ligand inhibiting T cell activity; Expression predicts response to immune checkpoint inhibitors. |
| PLK1 | Mitotic kinase promoting cell cycle progression; Overexpressed in melanoma; Potential therapeutic target. |
| S100A4 | Calcium-binding protein promoting cell motility and metastasis; Overexpressed in melanoma; Associated with melanoma progression and metastasis. |
| S100B | Calcium-binding protein that is involved in regulating cell proliferation and survival; Frequently overexpressed in melanoma; Used clinically as a serum biomarker for monitoring melanoma progression, detecting recurrence, and providing prognostic information, especially in advanced stages. |
| Skp2 | E3 ubiquitin ligase targeting cell cycle inhibitors for degradation; Overexpression linked to melanoma aggressiveness. |
| SOX10 | Neural crest transcription factor essential for melanocyte development; Highly sensitive diagnostic marker for melanoma cells. |
Osteosarcoma
Osteosarcoma is a rare but aggressive bone cancer that primarily affects children and young adults. Critical biomarkers include Alkaline Phosphatase (ALP) and Osteocalcin (BGLAP), which are used to evaluate bone turnover and tumor metabolism, while mutations in p53 and Runx2 expression levels shed light on genetic predisposition and tumor differentiation, respectively. Angiogenic factors, including VEGF, and invasion-related markers such as MMP-2, and MMP-9, play a vital role in understanding tumor progression and metastatic potential. Nestin and CD44 are emerging biomarkers that suggest the involvement of cancer stem-like cells in osteosarcoma aggressiveness. Refer to the table below for additional details on osteosarcoma biomarkers.
Biomarker | Relevance to Osteosarcoma Research |
|---|---|
| ABCG2 | ATP-binding cassette transporter involved in drug efflux and stem cell phenotype; Marker of cancer stem-like cells; Associated with chemoresistance and poor prognosis. |
| Alkaline Phosphatase (ALPL) | Enzyme involved in bone mineralization; Commonly elevated in osteosarcoma patients; Used as a diagnostic and prognostic serum biomarker. |
| CD44 | Cell surface glycoprotein involved in adhesion, migration, and interaction with the extracellular matrix; Marker of tumor aggressiveness and metastasis; Linked to chemoresistance. |
| Cyr61/CCN1 | Matricellular protein promoting angiogenesis, proliferation, and migration; Frequently overexpressed in osteosarcoma; Associated with poor prognosis. |
| Endoglin | Co-receptor for TGF-β involved in angiogenesis; Marker of tumor neovascularization; Potential prognostic indicator. |
| ERK5/BMK1 | MAP kinase regulating proliferation, survival, and migration; Promotes osteosarcoma growth and metastasis; Linked to chemoresistance in preclinical studies. |
| MKK4 | MAP kinase kinase involved in stress signaling and JNK/p38 activation; Modulates osteosarcoma cell apoptosis and metastasis; Expression levels correlate with tumor behavior. |
| MMP-2 | Matrix metalloproteinases involved in degradation of extracellular matrix components; Markers of invasiveness; Higher expression indicates aggressive phenotype. |
| MMP-9 | |
| Nestin | Intermediate filament protein; Marker of neural and mesenchymal stem cells; Linked with tumor stemness, invasion, metastasis, and chemoresistance. |
| Osteocalcin (BGLAP) | Bone-specific protein produced by osteoblasts; Used to assess osteoblastic differentiation; Lower levels may indicate less differentiated tumor. |
| p53 | Tumor suppressor regulating DNA repair and apoptosis; Frequently mutated in osteosarcoma; Loss correlates with poor prognosis and therapy resistance. |
| Runx2 | Master transcription factor critical for osteoblast differentiation; Overexpressed in osteosarcoma; Promotes tumor proliferation and bone matrix deposition. |
| STRO-1 | Marker for mesenchymal stem cells; Used to identify osteosarcoma stem-like cells contributing to tumor initiation and maintenance. |
| TSPAN31 | Member of tetraspanin family implicated in cell motility and proliferation; Frequently amplified in osteosarcoma; Associated with aggressive tumor behavior. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Overexpression correlates with increased vascularity and tumor growth; Target of anti-angiogenic therapies. |
Ovarian Cancer
Ovarian cancer is one of the deadliest gynecologic malignancies, often diagnosed at advanced stages due to its asymptomatic nature and molecular complexity. Established markers like CA125 (MUC16) and HE4 are used for disease monitoring, while mutations in BRCA1/2 and p53 help determine genetic susceptibility and aggressive tumor behavior. VEGF provides insight into tumor angiogenesis, and Mesothelin is used to assess active tumor-microenvironment engagement, tumor burden, and metastatic potential. Additional targets, including EpCAM, Her2/ErbB2, and HIF-1 alpha, contribute to understanding tumor biology and advancing therapeutic development. Learn more about ovarian cancer biomarkers in the table below.
Biomarker | Relevance to Ovarian Cancer Research |
|---|---|
| ATAD2 | Oncogene involved in chromatin remodeling and transcription regulation; Overexpressed in aggressive ovarian cancers; Linked to tumor progression, metastasis, and poor prognosis. |
| BRCA1 | DNA repair proteins involved in homologous recombination; Germline/somatic mutations in these genes increase cancer risk and are associated with hereditary ovarian cancer. Used for genetic testing and therapy selection. |
| BRCA2 | |
| CA 15-3 (MUC1) | Mucin glycoprotein often elevated in adenocarcinomas; Occasionally used adjunctively but less specific for ovarian cancer. |
| CA125/MUC16 | High molecular weight mucin; Most widely used serum biomarker for diagnosis, assessing treatment response, and monitoring recurrence. |
| EGFR | Receptor tyrosine kinase activating MAPK and PI3K pathways; Prognostic biomarker for epithelial ovarian cancer, overexpressed in 40%-60% of cases and often associated with a poor prognosis; Potential therapeutic target. |
| EpCAM / TROP1 | Cell adhesion molecule overexpressed on epithelial tumors; Frequently found on circulating tumor cells in advanced/metastatic ovarian cancers; Contributes to tumor cell adhesion, invasion, and immune evasion. |
| FOLR1 | Folate receptor alpha facilitates folate uptake critical for tumor proliferation; Highly overexpressed in ovarian cancers; Therapeutic target. |
| HE4 | Secreted protease inhibitor with expression in ovarian tumors; Serum biomarker complementary to CA125 for diagnosis and prognosis. |
| Her2/ErbB2 | Receptor tyrosine kinase; Overexpressed/amplified in a subset of ovarian cancers; Targeted by anti-Her2 therapies. |
| HIF-1 alpha | Transcription factors activated under hypoxic conditions in ovarian tumors, driving angiogenesis, metabolic adaptation, and therapy resistance. |
| HIF-2 alpha | |
| IL-6 | Pro-inflammatory cytokine promoting tumor growth and immune modulation; Elevated serum levels correlate with poor prognosis, therapy resistance, and cancer-related cachexia. |
| Mesothelin | Cell surface glycoprotein overexpressed in ovarian cancer; Promotes tumor adhesion, invasion, and metastasis; Diagnostic and therapeutic target; Detected in serum and tumor tissue. |
| MSH2 | A key DNA mismatch repair (MMR) protein; Deficiency/mutation associated with Lynch syndrome and increased ovarian cancer risk. |
| p53 | Tumor suppressor controlling DNA repair, and apoptosis; Frequently mutated in ovarian cancer; Mutation status guides prognosis and subtype classification, especially in high-grade serous tumors. |
| PTEN | Tumor suppressor regulating the PI3K/Akt pathway; Frequently dysregulated in ovarian cancer; Loss of function mutations contribute to tumorigenesis and progression. |
| Ras | Small GTPase involved in proliferation signaling; Mutations less frequent but present in some ovarian cancers. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Overexpression correlates with increased vascularity and tumor growth; Target of anti-angiogenic therapies. |
Pancreatic Cancer
Pancreatic cancer, primarily pancreatic ductal adenocarcinoma (PDAC), is one of the deadliest malignancies due to its late-stage diagnosis, rapid progression, and resistance to treatment. Widely recognized biomarkers such as CA19-9 and Carcinoembryonic Antigen (CEA) are essential for monitoring disease progression and recurrence, while mutational markers like Ras (KRAS), p53, and p16INK4a are critical for identifying genetic drivers of disease. Studies of Mesothelin, VEGF, and Glypican-1 are uncovering the microenvironmental and metastatic mechanisms in pancreatic cancer, offering hope for novel therapeutic interventions. Find more details on pancreatic cancer biomarkers in the table below.
Biomarker | Relevance to Pancreatic Cancer Research |
|---|---|
| CA 15-3 (MUC1) | Transmembrane mucin involved in cell adhesion and signaling; MUC1 overexpression is linked to tumor growth and immune evasion in pancreatic ductal adenocarcinoma (PDAC); Being investigated as a diagnostic and therapeutic marker. |
| CA19-9 | A sialylated Lewis blood group antigen and carbohydrate tumor-associated antigen; Most widely used serum biomarker in the clinical management of pancreatic ductal adenocarcinoma (PDAC); Widely accepted and used clinically as a serum biomarker in pancreatic cancer for prognosis and therapy monitoring. |
| Carcinoembryonic Antigen (CEA) | Cell adhesion molecule elevated in several adenocarcinomas; Used alongside CA19-9 to improve diagnostic and monitoring accuracy; More useful in advanced disease for monitoring disease progression or recurrence, rather than early detection. |
| CD44 | Cell surface adhesion receptor; Associated with cancer stem cell properties; Marker of tumor aggressiveness, metastatic potential, and chemoresistance. |
| CEACAM-1/CD66a | Cell adhesion molecule involved in immune evasion and tumor progression; Overexpressed in pancreatic cancer and associated with immune suppression and metastatic potential. |
| EGFR | Receptor tyrosine kinase activating proliferation pathways; Frequently overexpressed in pancreatic cancer; Drives cell proliferation and angiogenesis; Target for anti-EGFR therapies. |
| Glypican-1 | Cell surface heparan sulfate proteoglycan regulating growth factor signaling; Overexpressed in pancreatic cancer; Candidate for liquid biopsy and diagnostic marker. |
| Her2/ErbB2 | Receptor tyrosine kinase involved in cell proliferation; Amplified in a subset of pancreatic cancers; Potential therapeutic target. |
| HIF-1 alpha | Transcription factor activated under hypoxic conditions that controls genes involved in angiogenesis, metabolism, and survival; High expression correlates with aggressive phenotype and poor prognosis. |
| KRAS | Oncogenic GTPase; Frequently mutated in PDAC, locking it in an active state; Drives constitutive MAPK/ERK and PI3K/AKT signaling; Central to tumor initiation and progression; A critical therapeutic target. |
| LKB1/STK11 | Tumor suppressor kinase that regulates cell metabolism, polarity, and growth; Lost in sporadic cases of pancreatic cancer; Associated with aggressive disease and therapeutic resistance. |
| Mesothelin | Cell surface glycoprotein involved in cell adhesion and signaling; Overexpression in pancreatic cancer facilitates tumor spread and immune evasion; Diagnostic biomarker and immunotherapy target. |
| MSPR/Ron | Receptor tyrosine kinase involved in promoting proliferation and metastatic spread; Overexpression associated with tumor aggression and poor prognosis. |
| MUC4 | Transmembrane mucin linked to cell signaling and immune evasion; Overexpressed supports immune evasion and chemoresistance. |
| MUC5AC | Secreted mucin that modulates cell adhesion and signaling; Overexpression in pancreatic cancer promotes tumor progression and immune evasion, making it relevant as a diagnostic/prognostic marker. |
| p16INK4a | Tumor suppressor; Inhibits CDK4/CDK6 to control the cell cycle; Frequently inactivated in PDAC, resulting in cell cycle dysregulation and tumor growth. |
| p53 | Tumor suppressor controlling DNA repair and apoptosis; Frequently mutated in PDAC and linked to poor prognosis; Key prognostic and biological marker. |
| PSCA | Cell surface antigen implicated in cell proliferation and adhesion; Overexpressed in pancreatic cancer; Promotes tumor growth and invasion. |
| Ras | Small GTPase involved in proliferation signaling; Frequently mutated (KRAS) in PDAC; Driver of oncogenesis via activation of MAPK/ERK signaling. |
| Smad4 | Mediator of TGF-β signaling, tumor suppressor involved in growth inhibition; Frequently lost in PDAC; Loss/inactivation is associated with distant metastasis and poor prognosis. |
| VEGF | Growth factor that acts as a major driver of angiogenesis; Overexpression correlates with increased vascularity and tumor growth; Target of anti-angiogenic therapies. |
Prostate Cancer
Prostate cancer ranks among the most prevalent malignancies in men worldwide, characterized by diverse molecular profiles and variable clinical outcomes. Established biomarkers, including Prostate-Specific Antigen (PSA) and Prostate-Specific Membrane Antigen (PSMA), play a central role in diagnostics and disease monitoring, while the Androgen Receptor (AR) and its variants offer critical insights into resistance to hormone therapy. Emerging biomarkers such as EZH2 and AMACR (alpha-Methylacyl-CoA Racemase), alongside proliferation indicators like Ki67, are gaining recognition for their potential to predict aggressive disease. Additionally, tumor suppressors, including PTEN and DNA mismatch repair proteins MLH1 and MSH2, aid in identifying genetic alterations that can inform personalized treatment strategies. Further details on prostate cancer biomarkers are provided in the table below.
Biomarker | Relevance to Prostate Cancer Research |
|---|---|
| AKR1C3 (Aldo-keto-reductase 1C3) | Enzyme involved in androgen biosynthesis and steroid metabolism; Linked to castration-resistant prostate cancer progression and therapy resistance. |
| AMACR (alpha-Methylacyl-CoA Racemase) | Enzyme involved in fatty acid metabolism that is overexpressed in prostate cancer tissue; Standard diagnostic marker in pathology. |
| Androgen Receptor (AR) | Nuclear receptor regulating androgen-responsive gene expression; Central driver of prostate cancer development and progression; Target of androgen deprivation therapy (ADT). |
| Androgen Receptor V7 | Splice variant of AR lacking ligand-binding domain; Linked to resistance to androgen-deprivation therapies (ADT) and poor prognosis in metastatic prostate cancer. |
| Aurora A Kinase | Mitotic kinase regulating cell cycle and chromosomal stability; Overexpressed in aggressive tumors; Potential prognostic marker and therapeutic target. |
| Chromogranin A (CgA) | Secretory protein marker of neuroendocrine differentiation; Elevated in neuroendocrine prostate cancer; Associated with poor prognosis and resistance to therapy. |
| EZH2 | Histone methyltransferase involved in epigenetic silencing; Overexpressed in aggressive and metastatic prostate cancers; Important for prognosis and targeted therapies. |
| Hepsin | Transmembrane serine protease involved in extracellular matrix degradation; Overexpressed in aggressive and metastatic prostate cancer; Potential diagnostic and therapeutic relevance. |
| IL-6 | Pro-inflammatory cytokine driving tumor progression and androgen receptor activation; Elevated levels correlate with aggressive disease and therapy resistance. |
| Kallikrein 2 | Serine protease related to PSA; Biomarker complementary to PSA for early detection and risk stratification. |
| Ki67 | Cell proliferation marker; Used to assess tumor proliferation rate; Higher levels predict aggressive disease. |
| MLH1 | DNA mismatch repair proteins; Deficiency associated with microsatellite instability; Frequently investigated in advanced or metastatic cases of prostate cancer, often looking at loss of MSH2 and MSH6 proteins, which are associated with aggressive tumors and reduced survival. |
| MSH2 | |
| MSH6 | |
| PMS2 | |
| Prostate-Specific Antigen (PSA)/Kallikrein 3 | Serine protease produced by prostate epithelial cells; The most widely used serum biomarker for prostate cancer screening, diagnosis, and monitoring. |
| PSCA | Cell surface antigen involved in cell proliferation and adhesion; Overexpressed in prostate cancer; Potential diagnostic and therapeutic target. |
| PSMA | Cell surface protein with enzymatic activity that is highly expressed in prostate cancer cells, especially in advanced or metastatic cases; A key target for diagnostic imaging and therapies. |
| PSP94/MSMB | Secreted protein with tumor suppressive properties; Decreases in expression as prostate cancer progresses or becomes more aggressive; Being studied as a biomarker. |
| PTEN | Tumor suppressor gene regulating the PI3K/AKT pathway; Frequently lost or mutated in advanced prostate cancer; Associated with poor prognosis. |
| STEAP1 | Cell surface metalloreductase involved in metal ion transport and proliferation; Overexpressed in prostate cancer; Being investigated as a potential therapeutic target. |
| uPA | Protease involved in degradation of the extracellular matrix, promoting invasion and metastasis; Elevated in metastatic prostate cancer; Being investigated as a prognostic factor and therapeutic target. |
Immune Cell Therapy Workflow Wall Poster
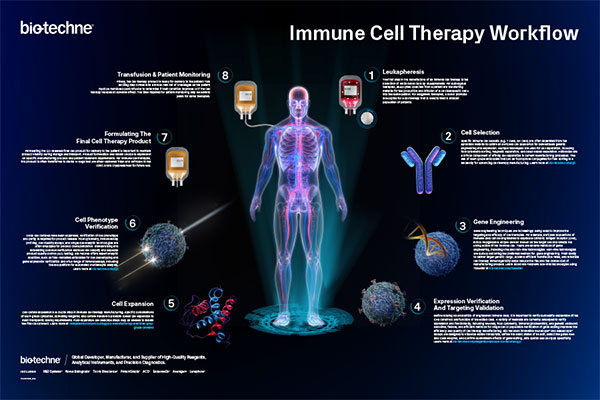
Explore this poster showing the workflow for producing immune cells at scale for therapeutic use. It will serve as a great reference tool for your lab.
Immune Checkpoint Targets eBook
Download this eBook to learn about the latest research on some of the most promising immune checkpoint targets for cancer immunotherapy.
Cell & Gene Therapy Manufacturing Solutions
Discover solutions to successfully scale cell therapies from preclinical development to commercialization.
Related Information
- Immunotherapy Research
- A Look Inside a Tumor Wall Poster
- Cancer Research
- Immune Cell Therapy Research
- Immune Checkpoint Research
- Immune Checkpoint Targets Wall Poster
- Multiplex Measurement of Biomarkers in Human Breast Cancer Cell Culture Supernates
- Tools: Human Tissue Kallikrein Products
- Technical Note: R&D Systems Cell Selection Kits for Cancer-associated Cells
- Axl Overexpression Promotes TKI Resistance in Non-small Cell Lung Cancer
- Custom Services - Tailor-Made Reagents
