Dcp1a Antibody (3G4) - Azide and BSA Free
Novus Biologicals | Catalog # H00055802-M06

![Western Blot: Dcp1a Antibody (3G4) [H00055802-M06] Western Blot: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Western-Blot-H00055802-M06-img0001.jpg)
Loading...
Key Product Details
Species Reactivity
Validated:
Human, Mouse, Rat, Insect
Cited:
Human, Mouse
Applications
Validated:
Immunohistochemistry, Western Blot, ELISA, Immunocytochemistry/ Immunofluorescence, Immunoprecipitation, Proximity Ligation Assay
Cited:
Western Blot, Immunocytochemistry/ Immunofluorescence, Immunoprecipitation
Label
Unconjugated
Antibody Source
Monoclonal Mouse IgG2a Kappa Clone # 3G4
Format
Azide and BSA Free
Loading...
Product Specifications
Immunogen
DCP1A (NP_060873, 186 a.a. ~ 285 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. STQLSNLGSTETLEEMPSGSQDKSAPSGHKHLTVEELFGTSLPKEQPAVVGLDSEEMERLPGDASQKEPNSFLPFPFEQLGGAPQSETLGVPSAAHHSVQ
Reactivity Notes
Mouse reactivity reported in scientific literature (PMID: 25530357).
Specificity
DCP1A - DCP1 decapping enzyme homolog A (S. cerevisiae)
Clonality
Monoclonal
Host
Mouse
Isotype
IgG2a Kappa
Scientific Data Images for Dcp1a Antibody (3G4) - Azide and BSA Free
Western Blot: Dcp1a Antibody (3G4) [H00055802-M06]
Western Blot: Dcp1a Antibody (3G4) [H00055802-M06] - Western Blot analysis of DCP1A expression in IMR-32 ( Cat # L008V1 ).Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0011.jpgImmunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Immunocytochemistry/Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] - Analysis of monoclonal antibody to DCP1A on HeLa cell. Antibody concentration 10 ug/mlImmunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0008.jpgImmunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0009.jpgImmunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0010.jpgImmunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]
Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0012.jpgELISA: Dcp1a Antibody (3G4) [H00055802-M06]
ELISA: Dcp1a Antibody (3G4) [H00055802-M06] - Detection limit for recombinant GST tagged DCP1A is approximately 0.1ng/ml as a capture antibody.Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] -
Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] - OTUD4 is recruited to stress granules. (A) Immunofluorescence of OTUD4 (shown in red) in SH-SY5Y cells that were either untreated, arsenite-treated (30 min, 0.5 mM) or heat-shocked (42°C, 1 h). Cells were co-stained for the stress granule marker protein TIA1 (green) & DAPI as a nuclear marker (blue). OTUD4 is redistributed to granular structures upon arsenite & heat-shock treatment. Granules also contain TIA1 & are considered as stress granules. Scale bar: 20 µm. The experiment was repeated two times. (B) Exogenously expressed FLAG–OTUD4 is recruited to stress granules in HeLa cells. Transfected cells were arsenite-treated (0.5 mM) for 40 min or left untreated & co-stained with anti-FLAG (red) & anti-G3BP1 antibodies (green). Nuclei are shown in blue (DAPI). Scale bar: 20 µm. The experiment was done at least three times. (C) OTUD4 granules do not colocalize with P-bodies. Shown is immunofluorescence of HeLa cells (untreated or 0.5 mM arsenite for 30 min) stained with anti-OTUD4 (red) & anti-DCP1a (green) antibodies. Scale bar: 20 µm. The experiment was performed two times. (D) OTUD4 granules contain mRNA. HeLa cells were treated with arsenite (0.5 mM) for 1 h. FISH was carried out with Cy3-labeled oligo(dT) (red), & cells were co-stained with anti-OTUD4 (shown in green). Scale bar: 20 µm. A representative image from four independent experiments is shown. (E) Scheme illustrating OTUD4 fragments used in F, numbers indicate amino acid borders of expression constructs. IDR, intrinsically disordered region. (F) HeLa cells were transfected with EGFP-tagged OTUD4 expression constructs as shown in E) & treated with arsenite (0.5 mM for 30 min) or left untreated to monitor intrinsic ability to form granules. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/31138677), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] -
Coronavirus infection does not alter steady state levels of most processing body proteins.A. HUVECs were transduced with human ACE2 (HUVECACE2), selected, and infected with SARS-CoV-2 TO-1 isolate (MOI = 3). Cells were lysed at 6 and 12 hours post infection and immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, SARS-CoV-2 N, and beta -actin specific antibodies. One representative experiment of two is shown. B-C. HUVECs were infected with OC43 (B, TCID50 = 2 x 104) or 229E (C, TCID50 = 2.4 x 103). Cells were lysed at 12 and 24 hours post infection (B, OC43) or 6 and 12 hours post infection (C, 229E). Immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, OC43 N protein (B only), and beta -actin specific antibodies. One representative experiment of three is shown. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/35998203), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] -
Processing body disassembly is not a common feature of all human coronavirus N proteins.A-D. HUVECs were transduced with recombinant lentiviruses ectopically expressing N protein from the Betacoronaviruses MERS-CoV and OC43 (A-B) or N protein from Alphacoronaviruses 229E and NL63 (C-D) or N protein from SARS-CoV-1 (E-F). A control lentiviral expressing an empty vector (EV) was used as a negative control and SARS-CoV-2 N protein expressing lentiviruses were used as a positive control in each experiment. Cells were selected, fixed and immunostained for DDX6 (PBs; white; Alexa555) and either authentic N protein or a FLAG tag (green; Alexa488). Nuclei were stained with Hoechst (blue). Scale bar = 20 μm. DDX6 puncta in EV or N-transduced cells were quantified using CellProfiler as in Fig 1. DDX6 puncta were quantified as in Fig 1. Representative images from one independent experiment of three are shown. These data represent three independent biological replicates (n = 3) with >30 cells measured per condition (EV and N) per replicate. Each EV and N replicate pair plotted independently; mean. Statistics were performed using Kruskal-Wallis H test with Dunn’s correction (*, p < 0.0332; **, p < 0.0021; ****, p < 0.0001; ns, nonsignificant). G-I. HUVECs were transduced as above, protein lysate was harvested and immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, N protein or FLAG, and beta -actin specific antibodies. One representative experiment of three is shown. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/35998203), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] -
Processing body disassembly is not a common feature of all human coronavirus N proteins.A-D. HUVECs were transduced with recombinant lentiviruses ectopically expressing N protein from the Betacoronaviruses MERS-CoV and OC43 (A-B) or N protein from Alphacoronaviruses 229E and NL63 (C-D) or N protein from SARS-CoV-1 (E-F). A control lentiviral expressing an empty vector (EV) was used as a negative control and SARS-CoV-2 N protein expressing lentiviruses were used as a positive control in each experiment. Cells were selected, fixed and immunostained for DDX6 (PBs; white; Alexa555) and either authentic N protein or a FLAG tag (green; Alexa488). Nuclei were stained with Hoechst (blue). Scale bar = 20 μm. DDX6 puncta in EV or N-transduced cells were quantified using CellProfiler as in Fig 1. DDX6 puncta were quantified as in Fig 1. Representative images from one independent experiment of three are shown. These data represent three independent biological replicates (n = 3) with >30 cells measured per condition (EV and N) per replicate. Each EV and N replicate pair plotted independently; mean. Statistics were performed using Kruskal-Wallis H test with Dunn’s correction (*, p < 0.0332; **, p < 0.0021; ****, p < 0.0001; ns, nonsignificant). G-I. HUVECs were transduced as above, protein lysate was harvested and immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, N protein or FLAG, and beta -actin specific antibodies. One representative experiment of three is shown. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/35998203), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] -
Coronavirus infection does not alter steady state levels of most processing body proteins.A. HUVECs were transduced with human ACE2 (HUVECACE2), selected, and infected with SARS-CoV-2 TO-1 isolate (MOI = 3). Cells were lysed at 6 and 12 hours post infection and immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, SARS-CoV-2 N, and beta -actin specific antibodies. One representative experiment of two is shown. B-C. HUVECs were infected with OC43 (B, TCID50 = 2 x 104) or 229E (C, TCID50 = 2.4 x 103). Cells were lysed at 12 and 24 hours post infection (B, OC43) or 6 and 12 hours post infection (C, 229E). Immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, OC43 N protein (B only), and beta -actin specific antibodies. One representative experiment of three is shown. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/35998203), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] -
Coronavirus infection does not alter steady state levels of most processing body proteins.A. HUVECs were transduced with human ACE2 (HUVECACE2), selected, and infected with SARS-CoV-2 TO-1 isolate (MOI = 3). Cells were lysed at 6 and 12 hours post infection and immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, SARS-CoV-2 N, and beta -actin specific antibodies. One representative experiment of two is shown. B-C. HUVECs were infected with OC43 (B, TCID50 = 2 x 104) or 229E (C, TCID50 = 2.4 x 103). Cells were lysed at 12 and 24 hours post infection (B, OC43) or 6 and 12 hours post infection (C, 229E). Immunoblotting was performed using XRN1, Hedls, DCP1A, DDX6, OC43 N protein (B only), and beta -actin specific antibodies. One representative experiment of three is shown. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/35998203), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for Dcp1a Antibody (3G4) - Azide and BSA Free
Application
Recommended Usage
ELISA
1:100-1:2000
Immunocytochemistry/ Immunofluorescence
1:10-1:2000
Western Blot
1:500
Application Notes
Antibody reactive against cell lysate and recombinant protein for Western Blot. Has also been used for immunofluoresence and ELISA. Use in immunprecipitation reported in scientific literature (PMID 21883093)
Reviewed Applications
Read 1 review rated 5 using H00055802-M06 in the following applications:
Formulation, Preparation, and Storage
Purification
IgG purified
Formulation
In 1x PBS, pH 7.4
Format
Azide and BSA Free
Preservative
No Preservative
Concentration
Concentrations vary lot to lot. See vial label for concentration. If unlisted please contact technical services.
Shipping
The product is shipped with polar packs. Upon receipt, store it immediately at the temperature recommended below.
Stability & Storage
Aliquot and store at -20C or -80C. Avoid freeze-thaw cycles.
Background: Dcp1a
Alternate Names
DCP1 decapping enzyme homolog A (S. cerevisiae), decapping enzyme hDcp1a, EC 3.-, FLJ21691, mRNA-decapping enzyme 1A, Nbla00360, putative protein product of Nbla00360, Smad4-interacting transcriptional co-activator, SMAD4IP1, SMIFHSA275986, Transcription factor SMIF
Entrez Gene IDs
55802 (Human)
Gene Symbol
DCP1A
OMIM
607010 (Human)
UniProt
Additional Dcp1a Products
Product Documents for Dcp1a Antibody (3G4) - Azide and BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for Dcp1a Antibody (3G4) - Azide and BSA Free
This product is produced by and distributed for Abnova, a company based in Taiwan.
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Citations for Dcp1a Antibody (3G4) - Azide and BSA Free
Customer Reviews for Dcp1a Antibody (3G4) - Azide and BSA Free (1)
5 out of 5
1 Customer Rating
Have you used Dcp1a Antibody (3G4) - Azide and BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
1 of
1 review
Showing All
Filter By:
-
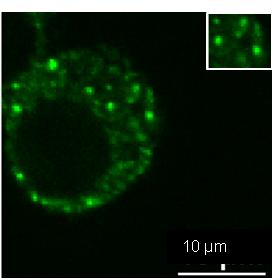
Application: ImmunocytochemistrySample Tested: RAW264.7 cellsSpecies: Mus MusculusVerified Customer | Posted 03/16/2018Dcp1a enriched in P bodies of RAW cell lines..IF analysis of Dcp1a protein with antibody at 1:100 dilution in RAW cell lines using a Nikon Widefield Fluorescence microscope
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ELISA Sample Preparation & Collection Guide
- ELISA Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- How to Run an R&D Systems DuoSet ELISA
- How to Run an R&D Systems Quantikine ELISA
- How to Run an R&D Systems Quantikine™ QuicKit™ ELISA
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Immunoprecipitation Protocol
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Quantikine HS ELISA Kit Assay Principle, Alkaline Phosphatase
- Quantikine HS ELISA Kit Principle, Streptavidin-HRP Polymer
- R&D Systems Quality Control Western Blot Protocol
- Sandwich ELISA (Colorimetric) – Biotin/Streptavidin Detection Protocol
- Sandwich ELISA (Colorimetric) – Direct Detection Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: ELISA
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...

![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0011.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0005.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0008.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0009.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0010.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-Immunocytochemistry-Immunofluorescence-H00055802-M06-img0012.jpg)
![ELISA: Dcp1a Antibody (3G4) [H00055802-M06] ELISA: Dcp1a Antibody (3G4) [H00055802-M06]](https://resources.rndsystems.com/images/products/Dcp1a-Antibody-3G4-ELISA-H00055802-M06-img0007.jpg)
![Immunocytochemistry/ Immunofluorescence: Dcp1a Antibody (3G4) [H00055802-M06] - Dcp1a Antibody (3G4)](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-310202415483610.jpg)
![Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] - Dcp1a Antibody (3G4) - Azide and BSA Free](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-western-blot-13220261328104.jpg)
![Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] - Dcp1a Antibody (3G4) - Azide and BSA Free](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-western-blot-13220261212518.jpg)
![Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] - Dcp1a Antibody (3G4) - Azide and BSA Free](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-western-blot-132202613165623.jpg)
![Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] - Dcp1a Antibody (3G4) - Azide and BSA Free](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-western-blot-13220261328105.jpg)
![Western Blot: Dcp1a Antibody (3G4) - Azide and BSA Free [H00055802-M06] - Dcp1a Antibody (3G4) - Azide and BSA Free](https://resources.rndsystems.com/images/products/h00055802-m06_mouse-monoclonal-dcp1a-antibody-3g4-western-blot-13220261755390.jpg)