EGLN1/PHD2 Antibody - BSA Free
Novus Biologicals | Catalog # NB100-137

![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Knockout Validated: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Knockout-Validated-NB100-137-img0019.jpg)
Key Product Details
Validated by
Species Reactivity
Validated:
Cited:
Predicted:
Applications
Validated:
Cited:
Label
Antibody Source
Format
Product Specifications
Immunogen
Reactivity Notes
Localization
Clonality
Host
Isotype
Theoretical MW
Disclaimer note: The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Scientific Data Images for EGLN1/PHD2 Antibody - BSA Free
Simple Western: EGLN1/PHD2 Antibody [NB100-137]
Simple Western: EGLN1/PHD2 Antibody [NB100-137] - Simple Western lane view shows a specific band for PHD2/HIF Prolyl Hydroxylase 2 in 0.5 mg/mL of hypoxic HeLa cell lysate. This experiment was performed under reducing conditions using the 12-230 kDa separation system.Western Blot: EGLN1/PHD2 Antibody [NB100-137]
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - Analysis of PHD2 expression in human prostate cancer cell lines. Western blot image submitted by a verified customer review.Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137]
Immunocytochemistry/Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137] - Staining of endogenous PHD2 in U2OS cells. ICC/IF image submitted by a verified customer review.Western Blot: EGLN1/PHD2 Antibody [NB100-137]
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - Detection of human PHD2 by Western blot. Recombinant epitope-tagged PHD1, PHD2 or PHD3 (10 ng/lane) or whole cell lysate from HeLa cells. EGLN1/PHD2 antibody used at 1 ug/mL. Detection by chemiluminescence.Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137]
EGLN1-PHD2-Antibody-Immunohistochemistry-NB100-137-img0018.jpgWestern Blot: EGLN1/PHD2 Antibody [NB100-137]
EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0017.jpgFlow Cytometry: EGLN1/PHD2 Antibody [NB100-137]
Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137] - An intracellular stain was performed on Jurkat cells with NB100-137AF647 (blue) and a matched isotype control (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 5 ug/mL for 30 minutes at room temperature. Both antibodies were conjugated to Alexa Fluor 647.Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137]
Immunocytochemistry/Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137] - Staining of PHD2 in HeLa cells with DyLight 488 (green). Nuclei and alpha-tubulin were counterstained with DAPI (blue) and DyLight 550 (red).Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137]
Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137] - Staining of lung vascular endothelium with EGLN1/PHD2 antibody. Image at 40X.Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137]
Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137] - Detection of PHD2 in Jurkat cells. One million Jurkat cells were fixed, permeabilized, and stained with 3.0 ug/mL anti-EGLN1/PHD2 antibody in a 150 uL reaction. Isotype control (black), anti-MLL1 (red).Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137] -
PHD2 and EGFR expression levels positively correlate in breast cancerProcessed tissue microarrays of breast cancer biopsies from 313 patients were stained with PHD2 and EGFR antibodies (cf Materials and methods). Four representative immunohistochemistries of human breast cancer with low and high expression of PHD2 and EGFR are shown. Magnification 10x.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Initial characterization of the OS-9 protein.(A) OS-9 expression in various human cell lines. Equal protein amounts of total cell lysates were used for SDS-PAGE & subsequent Western blotting. For each cell line, two independent samples are shown. Endogenous OS-9 was detected with a polyclonal antibody raised against a peptide corresponding to amino acids 600–667 of isoform 1 of OS-9. (B) Protein stability assay of endogenous OS-9. U2OS cells were treated with the translational inhibitor cycloheximide (100 µM). At indicated time points, whole cell lysates were analysed by immunoblotting. (C) Effect of hypoxia on OS-9 expression. For hypoxia, UT-7 cells were exposed to 1% O2 for 24 h prior to Western blot analysis. To determine any influence of HIF-1 alpha on OS-9 expression under normoxia, cells were incubated with the prolyl hydroxylase inhibitor DMOG (0.5 mM) for 24 h. (D) Protein interaction between OS-9 & PHD2 in vitro. For co-immunoprecipitation, U2OS cells were transiently co-transfected with the plasmids pOS-9-V5 & pPHD2-His, lysed in NP40 buffer, & subjected to immunoisolation with anti-V5 antibody recognizing OS-9 by its V5-tag. OS-9 & its associated proteins were separated by SDS-PAGE & analyzed by Western blot (lane 2). As controls, samples of untransfected (lane 1) cells or cells transfected with a single plasmid (lanes 3–4) were loaded. Representative Western blots are shown for each subfigure. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/21559462), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Cellular localization of OS-9 & PHD2.(A) A lectin gel-shift assay was conducted to test for glycosylated proteins. Total cell lysates of U2OS cells were incubated in the presence or absence of the endoglycosidases EndoH & PNGaseF for 6 h at 37°C. Digest products were separated on a reducing 10% SDS-PAGE gel which contained concanavalin A co-polymerized in the top layer of the separating gel to retard mobility of glycosylated proteins [56]. Glycosylated OS-9 is indicated as ‘g’, deglycosylated OS-9 as ‘d’. (B) Detection of OS-9 & PHD2 in the nuclear fraction. HEK293 cells with & without transfection of the plasmid pcDNA3-OS-9 were separated into nuclear fraction (N) & postnuclear supernatant (PS), the latter containing cytoplasm & organelles. Western blot analysis included BiP, GAPDH & lamin A as typical marker proteins for the ER, the cytoplasm & the nucleus, respectively. (C) Detection of OS-9 & PHD2 in the cytoplasm. HEK293 cells were co-transfected with pcDNA3-OS-9 & pPHD2-V5. For hypoxia, cells were exposed to 3% O2 for 4 h. Cells were treated with 50 µg/ml digitonin & centrifuged to obtain a cytoplasmic (C) & an organelle fraction (O) & subjected to immunoblotting. (D) Isolation of cellular endomembranes. HEK293 cells were lysed mechanically by several passages through a 30½G needle. The postnuclear supernatant was processed further by ultra-centrifugation to separate the organelles (O) from the cytosol (C). High salt treatment (1 M KCl) of the organelle fraction produced a wash fraction (W) that contained dissociated peripheral membrane proteins. For immunoblot analysis of subcellular fractionations, cell aliquots were normalized for cell number prior to loading (B–D). Representative Western blots are shown for each subfigure. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/21559462), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - Representative picture of western blot in histopathologically unchanged tissue (N) & primary cancerous tissue (C) from patients with CRC. Immunodetection of bands was performed with Rp anti- PHD1, - PHD2, - PHD3 & - FIH Ab, followed by incubation with goat anti-rabbit HRP-conjugated Ab. The membrane was stripped & incubated with Rp anti-GAPDH Ab, followed by incubation with goat anti-rabbit HRP-conjugated Ab. Bands were revealed using SuperSignal West Femto Chemiluminescent Substrate, Thermo Fisher Scientific (Rockford, IL) & Biospectrum® Imaging System 500, UVP Ltd. (Upland, CA). Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/24195777), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - OS-9 shows no effect on regulation of HIF-1 alpha.Total cell lysates were used for SDS-PAGE & subsequent Western blotting. To generate nearly anoxic conditions, cells were exposed to an oxygen consuming chemical system or to 1% or 3% O2 for 4 h to generate hypoxia. (A) U2OS, HeLa & Hep3B cells were transiently transfected with the plasmid pOS-9-V5 48 h prior to the experiment. Lamin A & actin were used as loading controls. (B) U2OS cells were subjected to ER stress by incubation either with tunicamycin (1 µg/ml) or thapsigargin (0.5 µg/ml) for 20 h. To detect HIF-1 alpha under normoxia, cells were treated with DMOG (1 mM) for 4 h. A sample of DMSO-only treated cells was loaded to exclude unspecific side effects of the solvent. (C) U2OS cells were transduced with lentiviral construct pLKO.1-shRNA-OS-9 (shOS-9) mediating a stable knockdown of OS-9 expression. Control cells (c) were transduced with plasmid pLKO.1-puro. Representative Western blots are shown for each subfigure. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/21559462), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - Oxygen chemotaxis is independent of the PHD/HIF pathway. a–dHIF1A, HIF2A, HIF1A+HIF2A & PHD2 CRISPR/Cas9 KO clones characterisation, respectively, regarding O2-directed migration. Left panels: immunoblot validation of HIF1A, HIF2A, HIF1A+HIF2A & PHD2 KO clones. To blot HIFs factors (a, b, c), cells were first pre-treated for 5 h with CoCl2 300 µM before protein extraction, a condition that promotes HIF factors accumulation (cf. Supplementary Fig. 8f). Middle panels: cell trajectories under confinement. Red dashed lines indicate the border of the cell cluster at 0 h. Right panels: relative distribution of MCF10A KO clones versus wt cells at the edge of the cluster at 48 h. These experiments demonstrate that HIF factors deletion does not prevent aerotaxis. e, f Tracks & redistribution of wt & PHD2 KO clone silenced for PHD3 (siPHD3) or not (siCTR). g Tracks & redistribution of MCF10A cells treated with DMOG (50 µM) or CoCl2 (50 µM) to inhibit PHDs, or with vehicle only (DMSO). These experiments demonstrate that PHDs do not participate in O2-sensing during aerotaxis. Confinement was applied for 48 h (a–g). Scale bars, 500 µm Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30382089), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - PHD3 expression correlates with a mesenchymal-like morphology in pancreatic ductal adenocarcinoma cell lines.NHF-1 (Fibroblast) MiaPaca2, Panc1, CAPAN1 & BxPC3 cells were harvested for RNA & protein following 24 hours exposure to normoxia (21% O2) or hypoxia (1% O2). (A) Phase-contrast images at 10× magnification were taken of MiaPaca2 (mesenchymal-like) morphology & BxPC3 cells (differentiated, epithelial morphology) under normoxic conditions. (B) PHD3 mRNA expression was determined by qRT-PCR & graphed relative to BxPC3 in normoxia. All samples were normalized to 18S rRNA & graphed as expression relative to BxPC3-Vec Normoxia (lane 1). n = 3, error bars = 1 S.D. (C) Whole cell lysate was resolved by SDS-PAGE & blotted for beta -tubulin PHD3 & PHD2. N = normoxia, H = hypoxia (1% O2). Image collected & cropped by CiteAb from the following publication (https://dx.plos.org/10.1371/journal.pone.0083021), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - Iron supplementation restores HIF-1 alpha levels to normal following ATP6V0E1 inhibition in HeLa cells. (A) Schematic diagram of the multimeric V-ATPase complex. (B) Chemical inhibition of V-ATPase by 10 nM BafA treatment for 24 h, increased HIF-1 alpha levels in HIF alpha -GFPODD reporter cells. Treatment with 100 μM Fe (III) citrate significantly reduced the elevated HIF-1 alpha levels associated with loss of ATP6V0E1 (1 × 106 cells per sample harvested & analyzed; N = 2). (C) Knock-down of ATP6V0E1 subunit with three different CRISPR-Cas9 guides resulted in significant upregulation of HIF-1 alpha levels in HIF alpha -GFPODD reporter cells. Co-treating cells with 100 μM Fe (III) citrate led to a reduction in HIF-1 alpha levels across the three ATP6V0E1 depleted cells. phd2 was knocked down as a control & treatment with 100 μM Fe (III) citrate did not result in reduction of HIF-1 alpha levels (1 × 106 cells per sample harvested & analyzed; N = 2). FACs plot shown is a representative image of two biological repeats performed. (D) Immunoblot analysis for HIF-1 alpha & PHD2 levels in HIF alpha -GFPODD reporter cells with either ATP6V0E1 or PHD2 depleted or treated with 10 nM BafA. The cells were treated with 100 μM Fe (III) citrate for 24 h. beta actin was used as a control. Results validated findings observed by flow cytometry, whereby HIF-1 alpha levels were upregulated following ATP6V0E1 knock-down or inhibition & levels were re-normalized upon Fe (III) citrate treatment. Treatment of Fe (III) citrate in PHD2 depleted cells did not alter HIF-1 alpha levels. All experiments were performed in biological duplicate. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/32984302), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: EGLN1/PHD2 Antibody [NB100-137] -
Western Blot: EGLN1/PHD2 Antibody [NB100-137] - PHD3 depletion stabilizes hypoxic p27 expression by increasing p27 half-life. a Cell cycle arrest at G0 & subsequent release shows an increase of p27 expression in siPHD3 exposed cells. b Quantification for p27 expression under PHD3 depletion at indicated time points after cell cycle release in HeLa & 786-O cells. Asterisk indicates significant difference (p < 0,05; n = 3). c Cell cycle arrest at G0 & inhibition of protein synthesis with cycloheximide indicate increased p27 stability in PHD3 depleted HeLa cells. d Quantification of p27 expression using siPHD3 or control at indicated time points. Four independent experiments (± SEM) are shown (p < 0,05; n = 4). e Analysis of p27 stability in 786–0 cells by cycloheximide chase during reoxygenation after 24 h hypoxia demonstrates markedly increased half-life of p27 upon PHD3 depletion Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/26223520), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for EGLN1/PHD2 Antibody - BSA Free
Flow Cytometry
Immunocytochemistry/ Immunofluorescence
Immunohistochemistry
Immunohistochemistry-Paraffin
Simple Western
Western Blot
In Simple Western only 10 - 15 uL of the recommended dilution is used per data point.
See Simple Western Antibody Database for Simple Western validation: Tested in Hypoxic HeLa lysate 0.5 mg/mL, separated by Size, antibody dilution of 1:500, apparent MW was 57 kDa. Separated by Size-Wes, Sally Sue/Peggy Sue.
The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Reviewed Applications
Read 12 reviews rated 4.8 using NB100-137 in the following applications:
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Formulation
Format
Preservative
Concentration
Shipping
Stability & Storage
Background: EGLN1/PHD2
EGLN1/PHD2 has been implicated in several critical processes including erythropoiesis, angiogenesis, and metabolism as well as various pathologies such as cancer (2, 5, 6). Studies in mice have found that somatic deletion of PHD2 resulted in higher vascular endothelial growth factor A (VEGF-A) levels, increased blood vessel formation, and more erythropoietin (EPO), leading to severe polycythemia or erythrocytosis (high red blood cell (RBC) volume) (6). Another study revealed that specific point mutations in EGLN1/PHD2 led to elevated EPO and RBC mass associated with hemorrhages and strokes (6). Accordingly, given the known role of PHD2 in inhibition of EPO production, PHD2 inhibitors are being studied as a potential therapeutic for anemia (6). Additionally, dysregulation in EGLN1, and specifically the PHD2-VHL-HIF-1alpha pathway, has been associated with the development of pheochromocytomas (PCC) and sympathetic paragangliomas (PGL), which are rare neuroendocrine tumors (2). Besides pathological features, EGLN1/PHD2 may also be important for high altitude adaptation as two coding sequence variants in PHD2 are prevalent in the Tibetan population but is very rare in people at lower altitudes (2).
Alternate names for EGLN1/PHD2 include HIF Prolyl Hydroxylase 2, PH2, Prolyl hydroxylase domain containing protein 2, HIF2PH2, HIF-Prolyl hydroxylase 2, egl nine homolog 1, and C1orf12.
References
1. Amorim-Pires, D., Peixoto, J., & Lima, J. (2016). Hypoxia Pathway Mutations in Pheochromocytomas and Paragangliomas. Cytogenetic and genome research. https://doi.org/10.1159/000457479
2. Gardie, B., Percy, M. J., Hoogewijs, D., Chowdhury, R., Bento, C., Arsenault, P. R., Richard, S., Almeida, H., Ewing, J., Lambert, F., McMullin, M. F., Schofield, C. J., & Lee, F. S. (2014). The role of PHD2 mutations in the pathogenesis of erythrocytosis. Hypoxia (Auckland, N.Z.). https://doi.org/10.2147/HP.S54455
3. Minervini, G., Quaglia, F., & Tosatto, S. C. (2015). Insights into the proline hydroxylase (PHD) family, molecular evolution and its impact on human health. Biochimie. https://doi.org/10.1016/j.biochi.2015.07.009
4. Semenza G. L. (2007). Hypoxia-inducible factor 1 (HIF-1) pathway. Science's STKE : signal transduction knowledge environment. https://doi.org/10.1126/stke.4072007cm8
5. Chan, D. A., & Giaccia, A. J. (2010). PHD2 in tumour angiogenesis. British journal of cancer. https://doi.org/10.1038/sj.bjc.6605682
6. Meneses, A. M., & Wielockx, B. (2016). PHD2: from hypoxia regulation to disease progression. Hypoxia (Auckland, N.Z.). https://doi.org/10.2147/HP.S53576
Long Name
Alternate Names
Gene Symbol
UniProt
Additional EGLN1/PHD2 Products
Product Documents for EGLN1/PHD2 Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for EGLN1/PHD2 Antibody - BSA Free
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Related Research Areas
Citations for EGLN1/PHD2 Antibody - BSA Free
Customer Reviews for EGLN1/PHD2 Antibody - BSA Free (12)
Have you used EGLN1/PHD2 Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
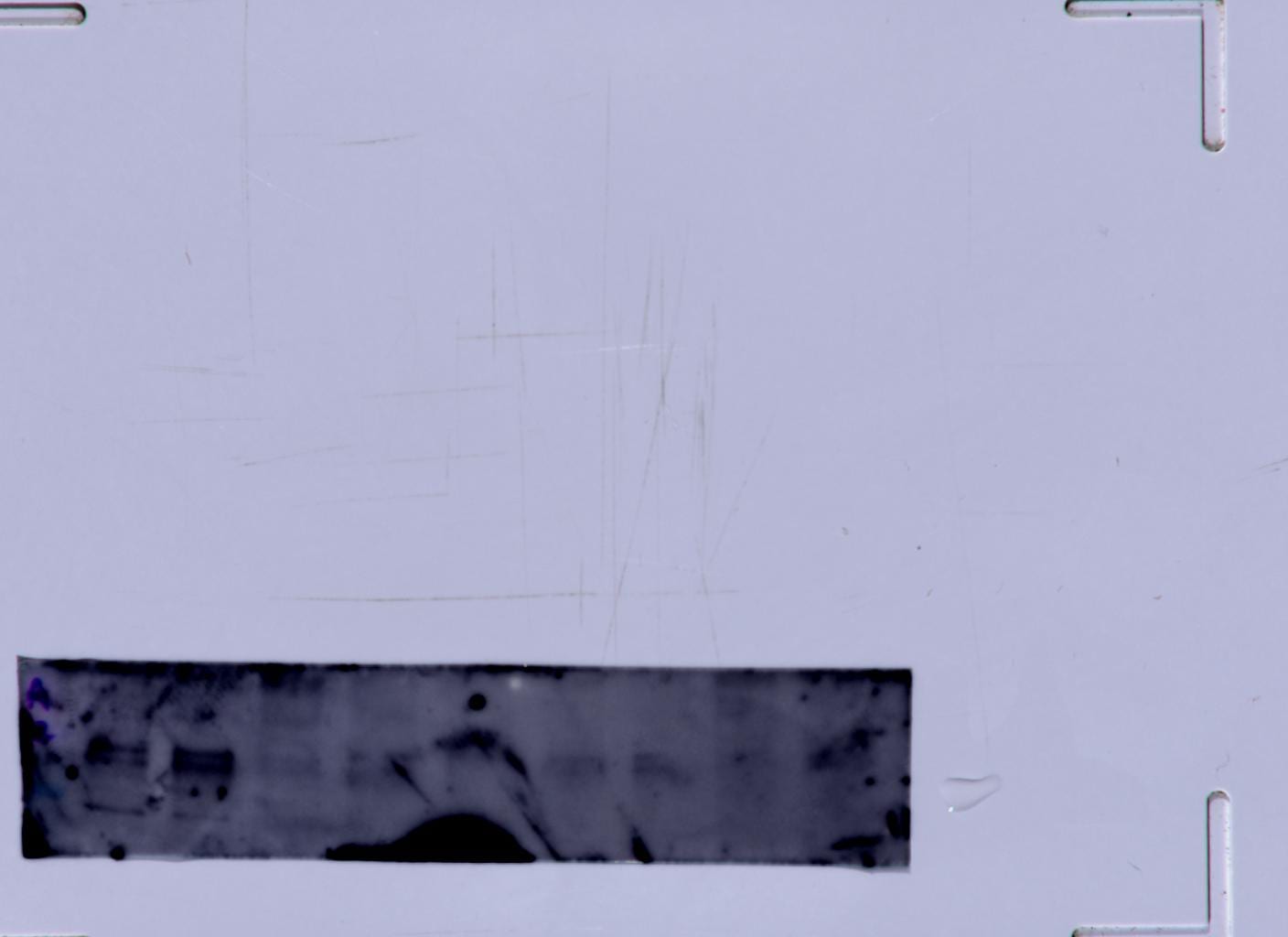
Application: Immunofluorescence and WBSample Tested: Protein and paraffin sectionsSpecies: Human and mouse liver cellsVerified Customer | Posted 01/19/2020PHD2 in human liver cells
-
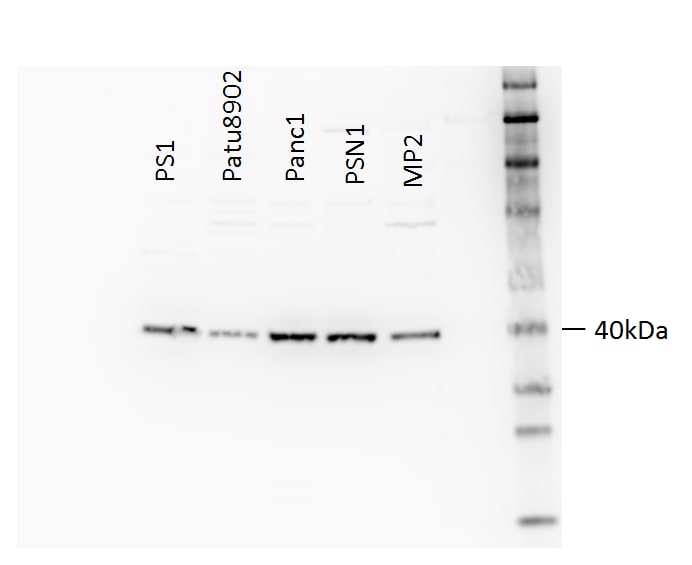
Application: Western BlotSample Tested: Human epithelial cell linesSpecies: HumanVerified Customer | Posted 10/23/201720 ug lysate. (Cell lines L-R): PS-1, PA-TU-8902, PANC-1, PSN-1, MIA PaCa2
-
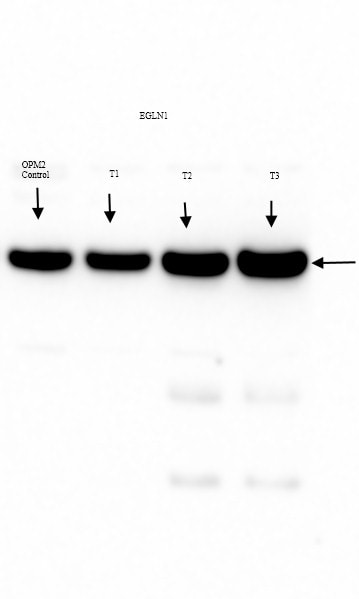
Application: Western BlotSample Tested: OPM2 cells whole cell lysateSpecies: HumanVerified Customer | Posted 10/08/2015EGLN1 expression in untreated control and drug treated OPM2 clones
-
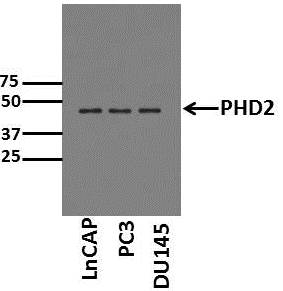
Application: Western BlotSample Tested: Cell lysates from prostate cancer cell linesSpecies: HumanVerified Customer | Posted 06/30/2015Western Blot for PHD2
-
Application: Western BlotSample Tested: See PMID 23225569Species: HumanVerified Customer | Posted 01/05/2015
-
Application: Western BlotSample Tested: See PMID 23328582Species: HumanVerified Customer | Posted 12/12/2014
-
Application: Western BlotSample Tested:Species: HumanVerified Customer | Posted 11/21/2014
-
Application: Western BlotSample Tested: Human SH-SY5Y or rat PC12 whole cell lysate.Species: OtherVerified Customer | Posted 09/22/2014
-
Application: Western BlotSample Tested: SK-N-BE(2) whole cell lysateSpecies: OtherVerified Customer | Posted 08/24/2014
-
Application: ImmunofluorescenceSample Tested: U2OS (human osteosarcoma cell line)Species: HumanVerified Customer | Posted 04/28/2014Immunofluorescence of endogenous PHD2 in U2OS cells
-
Application: Western BlotSample Tested: U87MG, U138MG, U343MG whole cell lysatesSpecies: HumanVerified Customer | Posted 04/28/2014Western blot detection of PHD2 human glioblastoma cells
-
Application: Western BlotSample Tested: HeLa cell lysates, Sample Amount: 20 ugSpecies: HumanVerified Customer | Posted 07/08/2010
There are no reviews that match your criteria.
Protocols
View specific protocols for EGLN1/PHD2 Antibody - BSA Free (NB100-137):
IHC-FFPE sections
I. Deparaffinization:
A. Treat slides with Xylene: 3 changes for 5 minutes each. Drain slides for 10 seconds between changes.
B. Treat slides with 100% Reagent Alcohol: 3 changes for 5 minutes each. Drain slides for 10 seconds between changes.
II. Quench Endogenous Peroxidase:
A. Place slides in peroxidase quenching solution: 15-30 minutes. To Prepare 200 ml of Quenching Solution: Add 3 ml of 30% Hydrogen Peroxide to 200 ml of Methanol.
Use within 4 hours of preparation
B. Place slides in distilled water: 2 changes for 2 minutes each.
III. Retrieve Epitopes:
A. Preheat Citrate Buffer. Place 200 ml of Citrate Buffer Working Solution into container, cover and place into steamer. Heat to 90-96 degrees Celsius.
B. Place rack of slides into hot Citrate Buffer for 20 minutes. Cover.
C. Carefully remove container with slides from steamer and cool on bench, uncovered, for 20 minutes.
D. Slowly add distilled water to further cool for 5 minutes.
E. Rinse slides with distilled water. 2 changes for 2 minutes each.
IV. Immunostaining Procedure:
A. Remove each slide from rack and circle tissue section with a hydrophobic barrier pen (e.g. Liquid Blocker-Super Pap Pen).
B. Flood slide with Wash Solution. Do not allow tissue sections to dry for the rest of the procedure.
C. Drain wash solution and apply 4 drops of Blocking Reagent to each slide and incubate for 15 minutes.
D. Drain Blocking Reagent (do not wash off the Blocking Reagent), apply 200 ul of Primary Antibody solution to each slide, and incubate for 1 hour.
E. Wash slides with Wash Solution: 3 changes for 5 minutes each.
F. Drain wash solution, apply 4 drops of Secondary antibody to each slide and incubate for 1 hour.
G. Wash slides with Wash Solution: 3 changes for 5 minutes each.
H. Drain wash solution, apply 4 drops of DAB Substrate to each slide and develop for 5-10 minutes. Check development with microscope.
I. Wash slides with Wash Solution: 3 changes for 5 minutes each.
J. Drain wash solution, apply 4 drops of Hematoxylin to each slide and stain for 1-3 minutes. Increase time if darker counterstaining is desired.
K. Wash slides with Wash Solution: 2-3 changes for 2 minutes each.
L. Drain wash solution and apply 4 drops of Bluing Solution to each slide for 1-2 minutes.
M. Rinse slides in distilled water.
N. Soak slides in 70% reagent alcohol: 3 minutes with intermittent agitation.
O. Soak slides in 95% reagent alcohol: 2 changes for 3 minutes each with intermittent agitation.
P. Soak slides in 100% reagent alcohol: 3 changes for 3 minutes each with intermittent agitation. Drain slides for 10 seconds between each change.
Q. Soak slides in Xylene: 3 changes for 3 minutes each with intermittent agitation. Drain slides for 10 seconds between each change.
R. Apply 2-3 drops of non-aqueous mounting media to each slide and mount coverslip.
S. Lay slides on a flat surface to dry prior to viewing under microscope.
NOTES:
-Use treated slides (e.g. HistoBond) to assure adherence of FFPE sections to slide.
-Prior to deparaffinization, heat slides overnight in a 60 degrees Celsius oven.
-All steps in which Xylene is used should be performed in a fume hood.
-For Epitope Retrieval, a microwave or pressure cooker may be substituted for the steamer method. Adjust times as necessary depending on conditions.
-For the initial IHC run with a new primary antibody, test tissues with and without Epitope Retrieval. In some instances, Epitope Retrieval may not be necessary.
-200 ul is the recommended maximum volume to apply to a slide for full coverage. Using more than 200 ul may allow solutions to wick off the slide and create drying artifacts. For small tissue sections less than 200 ul may be used.
-5 minutes of development with DAB Substrate should be sufficient. Do not develop for more than 10 minutes. If 5 minutes of development causes background staining, further dilution of the primary antibody may be necessary.
-Hematoxylin should produce a light nuclear counterstain so as not to obscure the DAB staining. Counterstain for 1-1.5 minutes for nuclear antigens. Counterstain for 2-3 minutes for cytoplasmic and membranous antigens. If darker counterstaining is desired increase time (up to 10 minutes).
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Immunoprecipitation Protocol
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Liperfluo
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
FAQs for EGLN1/PHD2 Antibody - BSA Free
-
Q: I would like to use this antibody but it has not been validated in my species of interest. Is there any way I can find out if it will work?
A: We offer risk-free testing of all of our primary antibodies. Please check out our Innovator's Reward Program and test this EGLN1-PHD2 antibody in any unvalidated species or application, without the financial risk of failure.
-
Q: What is the probability of this antibody cross-reacting with the bovine protein. Can you send me the alignment score of the immunogen with the bovine protein?
A:
The immunogen used to generate this antibody shares 100% homology with bovine. This antibody will likely cross react with bovine, however we have not yet tested it in that species. If you would be interested in testing this species, please take a look at our Innovators Reward Program.
-
Q: I would like to use this antibody but it has not been validated in my species of interest. Is there any way I can find out if it will work?
A: We offer risk-free testing of all of our primary antibodies. Please check out our Innovator's Reward Program and test this EGLN1-PHD2 antibody in any unvalidated species or application, without the financial risk of failure.
-
Q: What is the probability of this antibody cross-reacting with the bovine protein. Can you send me the alignment score of the immunogen with the bovine protein?
A:
The immunogen used to generate this antibody shares 100% homology with bovine. This antibody will likely cross react with bovine, however we have not yet tested it in that species. If you would be interested in testing this species, please take a look at our Innovators Reward Program.

![Simple Western: EGLN1/PHD2 Antibody [NB100-137] Simple Western: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Simple-Western-NB100-137-img0010.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Western Blot: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0011.jpg)
![Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137] Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Immunocytochemistry-Immunofluorescence-NB100-137-img0006.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Knockdown Validated: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0005.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Western Blot: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0013.jpg)
![Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137] Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Immunohistochemistry-NB100-137-img0018.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Western Blot: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0017.jpg)
![Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137] Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Flow-Cytometry-NB100-137-img0014.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] Western Blot: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Western-Blot-NB100-137-img0012.jpg)
![Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137] Immunocytochemistry/ Immunofluorescence: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Immunocytochemistry-Immunofluorescence-NB100-137-img0003.jpg)
![Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137] Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Immunohistochemistry-NB100-137-img0015.jpg)
![Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137] Flow Cytometry: EGLN1/PHD2 Antibody [NB100-137]](https://resources.rndsystems.com/images/products/EGLN1-PHD2-Antibody-Flow-Cytometry-NB100-137-img0007.jpg)
![Immunohistochemistry: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-27122023125334.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-210202423452425.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-210202423454817.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-310202415293310.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-31020241683931.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-31020241612618.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-310202415541942.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-310202415533838.jpg)
![Western Blot: EGLN1/PHD2 Antibody [NB100-137] - EGLN1/PHD2 Antibody](https://resources.rndsystems.com/images/products/nb100-137_rabbit-polyclonal-egln1-phd2-antibody-310202415533825.jpg)


-(01-ml)_NB100-137_7091.jpg)
-(01-ml)_NB100-137_7051.jpg)