SP‑D (surfactant protein‑D; also PSP‑D) is a 43 kDa member of the collectin family of innate immune modulators (1 ‑ 5). It is constitutively secreted by alveolar lining cells and epithelium associated with tubular structures. SP‑D is found in serum, plasma, broncho‑alveolar lavage (BAL) fluid, and amniotic fluid (1, 2, 6). Lung injuries often increase release of SP‑D to the circulation (3, 6). Mouse SP‑D is synthesized as a 374 amino acid (aa) precursor. Mouse SP‑D cDNA encodes a 19 aa signal sequence and a 355 aa mature region with a 25 aa N‑terminal linking‑region, a 177 aa hydroxyproline and hydroxylysine collagen‑like domain, a 46 aa coiled‑coil segment, and a 106 aa, C‑terminal collectin‑like C‑type lectin domain (CRD) (5). Mature mouse SP‑D shares 72 ‑ 76% aa sequence identity with human, porcine, equine, canine and bovine SP‑D, and 92% with rat SP‑D. SP‑D is usually found as a glycosylated, disulfide‑linked 150 kDa alpha ‑helical coiled‑coil trimer with a “head” of three symmetrical CRDs (2‑4, 7). Each CRD recognizes the hydroxides of one monosaccharide, and trimerization allows for the discrimination of monosaccharide patterns specific to microbial pathogens (4, 7, 8). Typically, SP‑D forms a higher‑order 620 kDa, X‑shaped dodecamer through N‑terminal disulfide bonds, allowing for even finer discrimination of self vs. nonself carbohydrate patterns and facilitating binding to complex antigens (1). SP‑D also binds SIRP alpha and the calreticulin/CD91 complex on macrophages (9, 10). When the ratio of antigen/pathogen to available CRDs is low, antigen can be bound without occupying all available CRDs. The free CRDs will bind to SIRP alpha, generating a signal that downmodulates the inflammatory response. During high CRD ligand binding (low SIRP alpha binding), the dodecamer rearranges to expose N‑termini that bind the calreticulin/CD91 complex, an event that initiates inflammation (1). Also, direct and indirect binding of neutrophil defensins and macrophage CD14 and TLRs to SP‑D can modulate response to viruses and bacterial lipopolysaccharides (1‑3, 11‑15). Thus, SP‑D allows for a graded response to environmental challenge and clearance of small antigenic insults without the need for a damaging inflammatory response (1‑3).

Key Product Details
Species Reactivity
Mouse, Rat
Applications
Western Blot, Simple Western
Label
Unconjugated
Antibody Source
Polyclonal Sheep IgG
Loading...
Product Specifications
Immunogen
Chinese hamster ovary cell line CHO-derived recombinant mouse SP-D
Ala20-Phe374
Accession # P50404
Ala20-Phe374
Accession # P50404
Specificity
Detects mouse and rat SP-D in Western blots. In direct ELISAs, approximately 20% cross-reactivity with recombinant human SP-D is observed.
Clonality
Polyclonal
Host
Sheep
Isotype
IgG
Scientific Data Images for Mouse/Rat SP‑D Antibody
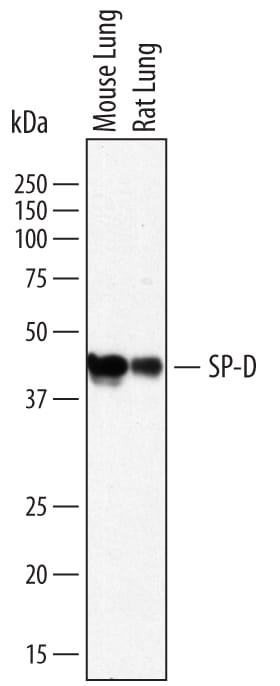
Detection of Mouse and Rat SP‑D by Western Blot.
Western blot shows lysates of mouse lung tissue and rat lung tissue. PVDF membrane was probed with 2 µg/mL of Sheep Anti-Mouse/Rat SP-D Antigen Affinity-purified Polyclonal Antibody (Catalog # AF6839) followed by HRP-conjugated Anti-Sheep IgG Secondary Antibody (Catalog # HAF016). A specific band was detected for SP-D at approximately 43 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.Detection of Rat and Mouse SP‑D by Simple WesternTM.
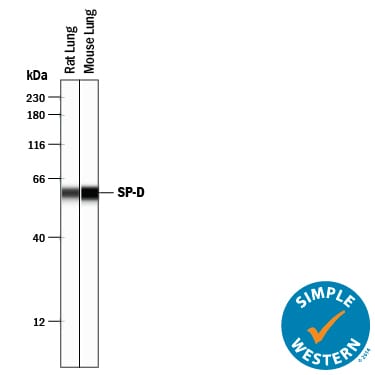
Simple Western lane view shows lysates of rat lung tissue and mouse lung tissue, loaded at 0.2 mg/mL. A specific band was detected for SP-D at approximately 60 kDa (as indicated) using 20 µg/mL of Sheep Anti-Mouse/Rat SP-D Antigen Affinity-purified Polyclonal Antibody (Catalog # AF6839) followed by 1:50 dilution of HRP-conjugated Anti-Sheep IgG Secondary Antibody (Catalog # HAF016). This experiment was conducted under reducing conditions and using the 12-230 kDa separation system.Applications for Mouse/Rat SP‑D Antibody
Application
Recommended Usage
Simple Western
20 µg/mL
Sample: Rat lung tissue and mouse lung tissue
Sample: Rat lung tissue and mouse lung tissue
Western Blot
2 µg/mL
Sample: Mouse lung tissue and rat lung tissue
Sample: Mouse lung tissue and rat lung tissue
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Sterile PBS to a final concentration of 0.2 mg/mL. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: SP-D
References
- Forbes, L.R. and A. Haczku (2010) Clin. Exp. Allergy 40:547.
- Kishore, U. et. al. (2006) Mol. Immunol. 43:1293.
- Hartl, D. and M. Griese (2006) Eur. J. Clin. Invest. 36:423.
- Sim, R.B. et. al. (2006) Novartis Found Symp. 279:170.
- Motwani, M. et al. (1995) J. Immunol. 155:5671.
- Honda, Y. et al. (1995) Am. J. Respir. Crit. Care Med. 152:1860.
- Hakansson, K. et. al. (1999) Structure 7:225.
- Crouch, E.C. et. al. (2006) Am. J. Respir. Cell Mol. Biol. 35:84.
- Janssen, W.J. et al. (2008) Am. J. Respir. Crit. Care Med. 178:158.
- Gardai, S.J. et al. (2003) Cell 115:13.Ohya, M. et. al. (2006) Biochemistry 45:8657.
- Ohya, M. et. al. (2006) Biochemistry 45:8657.
- Pastva, A.M. et al. (2007) Proc. Am. Thorac. Soc. 4:252.
- Sano, H. and Y. Kuroki (2005) Mol. Immunol. 42:279.
- Hartshorn, K.L. et al. (2006) J. Immunol. 176:6962.
- Yamazoe, M. et al. (2008) J. Biol. Chem. 283:35878.
Long Name
Surfactant Pulmonary Associated Protein D
Alternate Names
Collectin 7, PSP-D, SFTP4, SFTPD, SPD
Gene Symbol
SFTPD
UniProt
Additional SP-D Products
Product Documents for Mouse/Rat SP‑D Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Mouse/Rat SP‑D Antibody
For research use only
Related Research Areas
Customer Reviews for Mouse/Rat SP‑D Antibody
There are currently no reviews for this product. Be the first to review Mouse/Rat SP‑D Antibody and earn rewards!
Have you used Mouse/Rat SP‑D Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Cellular Response to Hypoxia Protocols
- R&D Systems Quality Control Western Blot Protocol
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...