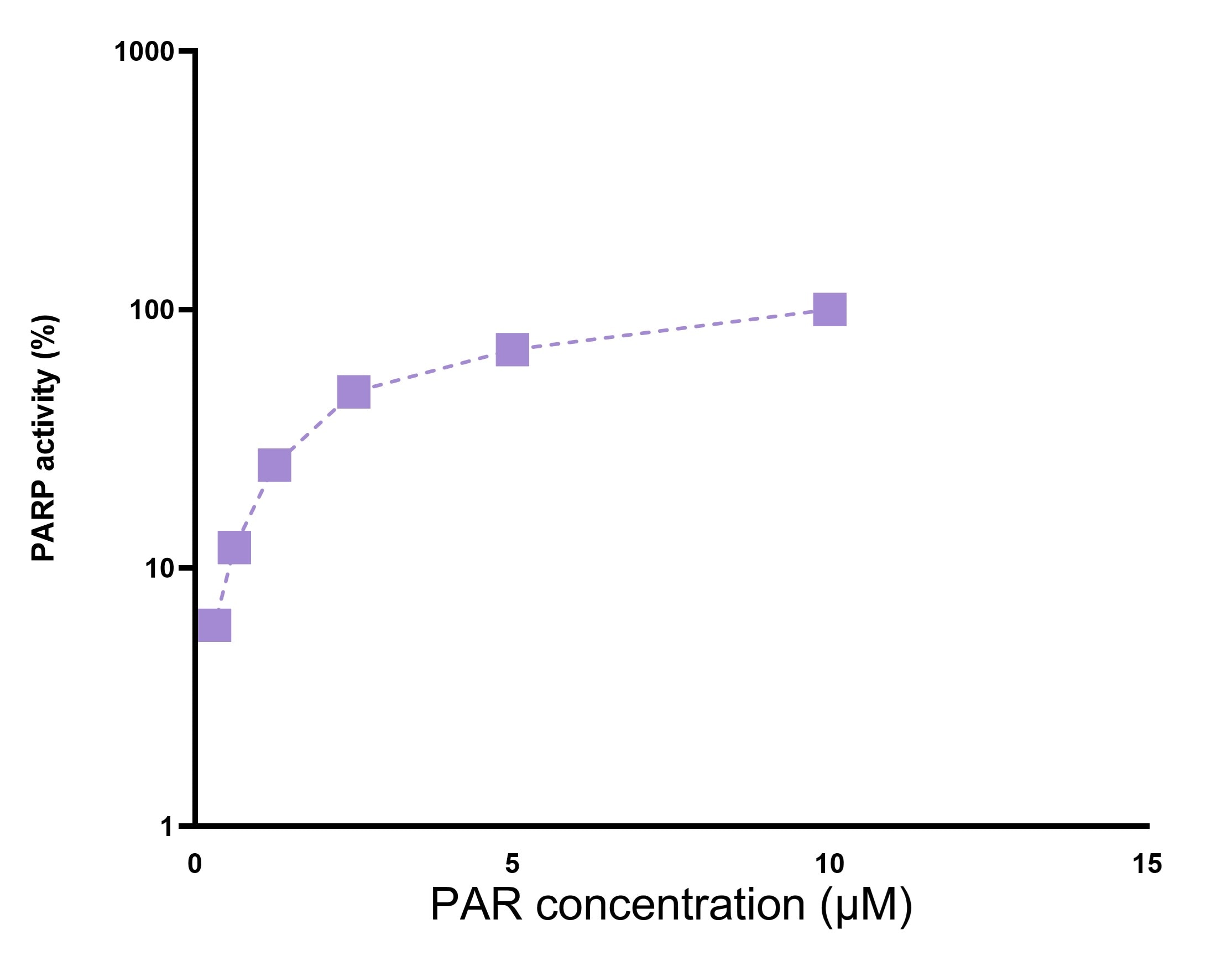
PARP [Poly(ADP-ribose) Polymerase], also known as ADPRT and PPOL, is a 118-kDa enzyme that uses NAD as a substrate to catalyze the covalent transfer of ADP-ribose to a variety of nuclear protein acceptors. ADP ribosyltransferase is required for cellular repair, and PARP expression is induced by single-strand breaks in DNA. PARP is proteolytically cleaved by Caspase-3 into two fragments of 89- and 24-kDa in one of the hallmark events of apoptosis.
Poly(ADP-ribose) Polymer/pADPr
R&D Systems | Catalog # 4336-100-01

Key Product Details
Features
Formulation, Preparation, and Storage
Shipping
Storage
Background: PAR/pADPr
Long Name
Alternate Names
Additional PAR/pADPr Products
Product Documents for Poly(ADP-ribose) Polymer/pADPr
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Related Research Areas
Citations for Poly(ADP-ribose) Polymer/pADPr
Customer Reviews for Poly(ADP-ribose) Polymer/pADPr (3)
Have you used Poly(ADP-ribose) Polymer/pADPr?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Verified Customer | Posted 03/21/2025
-
Verified Customer | Posted 07/28/2021works good
-
Verified Customer | Posted 12/15/2020
There are no reviews that match your criteria.
FAQs for Poly(ADP-ribose) Polymer/pADPr
-
Q: What is the concentration of this polymer in units of mass/volume?
A: The concentration of poly(ADP-ribose) Polymer/pADPr is determined by measuring light absorbance at 258 nm using the molar extinction coefficient of PAR. This measurement yields concentration in units of μM. The chain length of this polymer ranges between 2 and 300 ADP-ribose units. Because we are unable to evaluate the variation in polymer length and polymer branching, we are unable to calculate a concentration based on mass. As such, this product is bottled and sold by the concentration in molarity.