Distinguish Between Poly-ubiquitination and Multi-mono-ubiquitination
Ubiquitin can be attached to a protein substrate via two distinct mechanisms (Figure 1). Poly-ubiquitination occurs when Ubiquitin molecules are attached end-to-end to a single lysine residue on a substrate protein to form a poly-ubiquitin chain (Figure 1A). Alternatively, multi-mono-ubiquitination is the attachment of a single Ubiquitin molecule to multiple lysine residues on a substrate protein (Figure 1B). It is important to distinguish between poly-ubiquitination and multi-mono-ubiquitination because these different types of ubiquitination lead to different functions of the substrate protein. To complicate matters, a poly-ubiquitinated protein and a multi-mono-ubiquitinated protein look very similar by SDS-PAGE and Western blot. Fortunately, it is relatively simple to differentiate between poly-ubiquitination and multi-mono-ubiquitination by performing in vitro ubiquitin conjugation reactions. The protocol below describes in detail how to determine if your protein of interest is poly-ubiquitinated or multi-mono-ubiquitinated. Click on the links below to view listings of Boston Biochem® proteins, enzymes, and buffers.
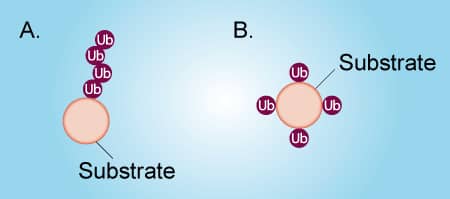
Figure 1. Poly-ubiquitination vs Multi-mono-ubiquitination
Materials and reagents:
| Material or Reagent | Stock Concentration |
|---|---|
| E1 Enzyme | 5 µM |
| E2 Enzymei | 25 µM |
| E3 Ligaseii | 10 µM |
| 10X E3 Ligase Reaction Buffer | 10X - (500 mM HEPES, pH 8.0, 500 mM NaCl, 10 mM TCEP) |
| Ubiquitin | 1.17 mM (10 mg/mL) |
| Ubiquitin No K | 1.17 mM (10 mg/mL) |
| MgATP Solution | 100 mM |
| SDS-PAGE sample buffer – if not using reaction products for downstream applications | 2X |
| EDTA or DTT – if using reaction products for downstream applications | 500 mM (EDTA);1 M (DTT) |
| Microcentrifuge tubes | |
| Water Bath (37 °C) | |
| Western Blot Equipment |
Procedure for distinguishing between poly-ubiquitination and multi-mono-ubiquitination
Two in vitro Ubiquitin conjugation reactions will need to be performed: 1) one that contains wild type Ubiquitin and 2) one that contains Ubiquitin No K, a mutant in which all 7 lysines have been mutated to arginines. Wild type Ubiquitin can be conjugated to substrate proteins and is capable of forming chains (Figure 2; left). Ubiquitin No K can also be conjugated to substrate proteins, but it is unable to form chains due to its lack of lysine residues (Figure 2; right). Therefore, if your substrate is poly-ubiquitinated, high molecular weight bands will be generated in reaction 1, but not reaction 2 (Figure 3A). Conversely, if your substrate is multi-mono-ubiquitinated, high molecular weight bands will be generated in both reaction 1 and reaction 2 (Figure 3B). Note that reaction 1 will look the same for both poly-ubiquitinated and multi-mono-ubiquitinated substrates.
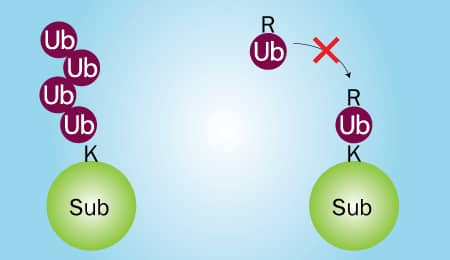
Figure 2. Wild Type Ubiquitin vs Ubiquitin No K
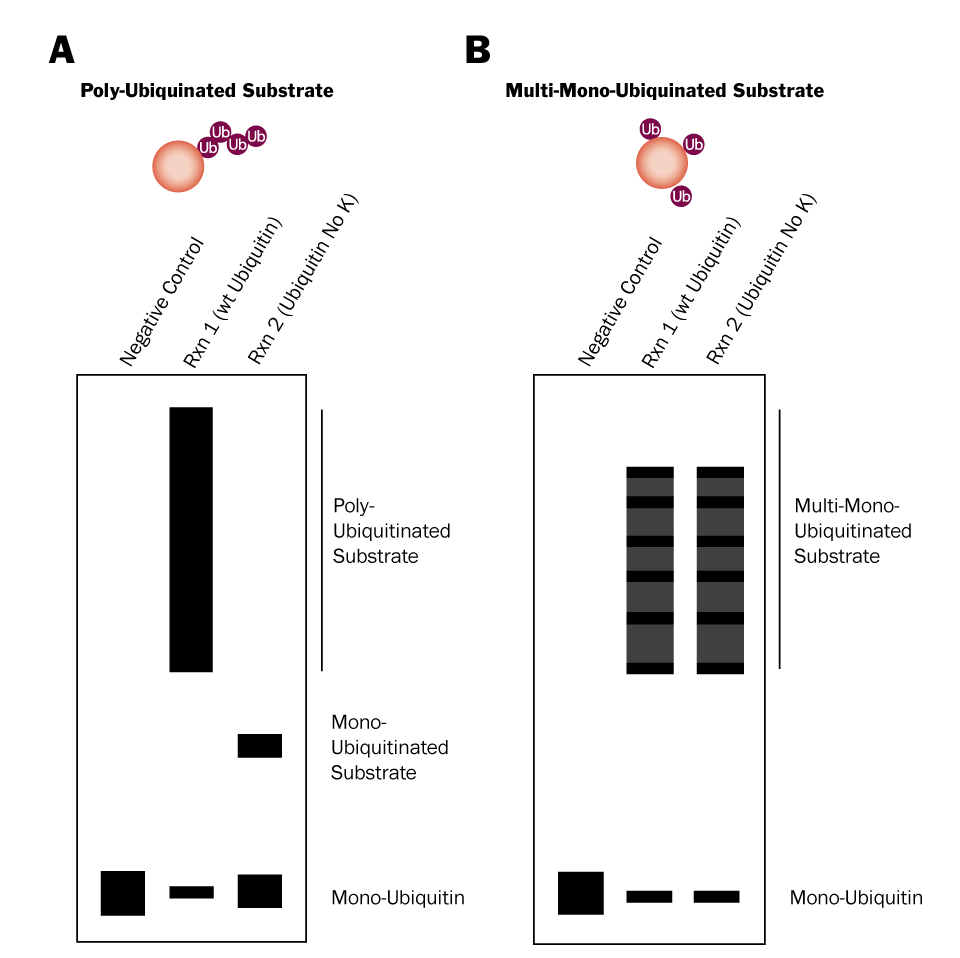
Figure 3. Differentiation Between Poly-Ubiquitination and Multi-Mono-Ubiquitination Using Ubiquitin No K
Procedure for 25 µL reactions (scale as needed):
For reaction 1, combine the indicated volume of each component listed in the table below, in the order shown, in a microcentrifuge tube. For a negative control reaction replace the MgATP Solution with dH2O.
Reaction 1
iiiThe volume needed will depend on the stock concentration of your substrate.
ivThe volume needed will depend on the stock concentration of your E3 ligase.Reagent Volume Working Concentration dH2O X µL (to 25 µL; dependent on volume of substrate and E3 ligase) N/A 10X E3 Ligase Reaction Buffer 2.5 µL 1X - (50 mM HEPES, pH 8.0, 50 mM NaCl, 1 mM TCEP) Ubiquitin 1 µL Approximately 100 µM MgATP Solution 2.5 µL 10 mM Substrate X µLiii 5-10 µM E1 Enzyme 0.5 µL 100 nM E2 Enzyme 1 µL 1 µM E3 Ligase X µLiv 1 µM For reaction 2, combine the indicated volume of each component listed in the table below, in the order shown, in a microcentrifuge tube. The only difference between reaction 1 and reaction 2 is the substitution of Ubiquitin No K for wild type Ubiquitin.
Reaction 2
iiiThe volume needed will depend on the stock concentration of your substrate.
ivThe volume needed will depend on the stock concentration of your E3 ligase.Reagent Volume Working Concentration dH2O X µL (to 25 µL; dependent on volume of substrate and E3 ligase) N/A 10X E3 Ligase Reaction Buffer 2.5 µL 1X - (50 mM HEPES, pH 8.0, 50 mM NaCl, 1 mM TCEP) Ubiquitin No K 1 µL Approximately 100 µM MgATP Solution 2.5 µL 10 mM Substrate X µLiii 5-10 µM E1 Enzyme 0.5 µL 100 nM E2 Enzyme 1 µL 1 µM E3 Ligase X µLiv 1 µM - Incubate reaction 1 and reaction 2 in a 37 °C water bath for 30-60 minutes.
Terminate the reactions. See the table below for the appropriate method of termination.
vEDTA and DTT are equally effective at terminating the reaction; determining which to use will depend on the intended downstream enzymatic application of the reaction products. Are you using reaction products for downstream enzymatic applications? Termination method Volume (final concentration) No SDS-PAGE sample buffer 25 µL (1X) Yes EDTA or DTTv 0.5 µL EDTA (20 mM) or 1 µL DTT (100 mM) Analyze Ubiquitin conjugation reactions.
- Separate the reaction products by SDS-PAGE and transfer them to a PVDF or nitrocellulose membrane.
- Perform a Western blot using an anti-Ubiquitin antibody.
- Compare your Western blot to figure 3 above to determine if your substrate is poly-ubiquitinated or multi-mono-ubiquitinated.*
*It is possible for a substrate to be both poly-ubiquitinated and multi-mono-ubiquitinated. If this is the case the Ubiquitin No K will still prevent poly-ubiquitination and the highest molecular weight protein species should disappear from reaction 2. Contact us for technical assistance!
