CHREBP Antibody - BSA Free
Novus Biologicals | Catalog # NB400-135

![Western Blot: CHREBP AntibodyBSA Free [NB400-135] Western Blot: CHREBP AntibodyBSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0023.jpg)
Key Product Details
Validated by
Species Reactivity
Validated:
Cited:
Applications
Validated:
Cited:
Label
Antibody Source
Format
Product Specifications
Immunogen
Reactivity Notes
Clonality
Host
Isotype
Theoretical MW
Disclaimer note: The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Scientific Data Images for CHREBP Antibody - BSA Free
Western Blot: CHREBP AntibodyBSA Free [NB400-135]
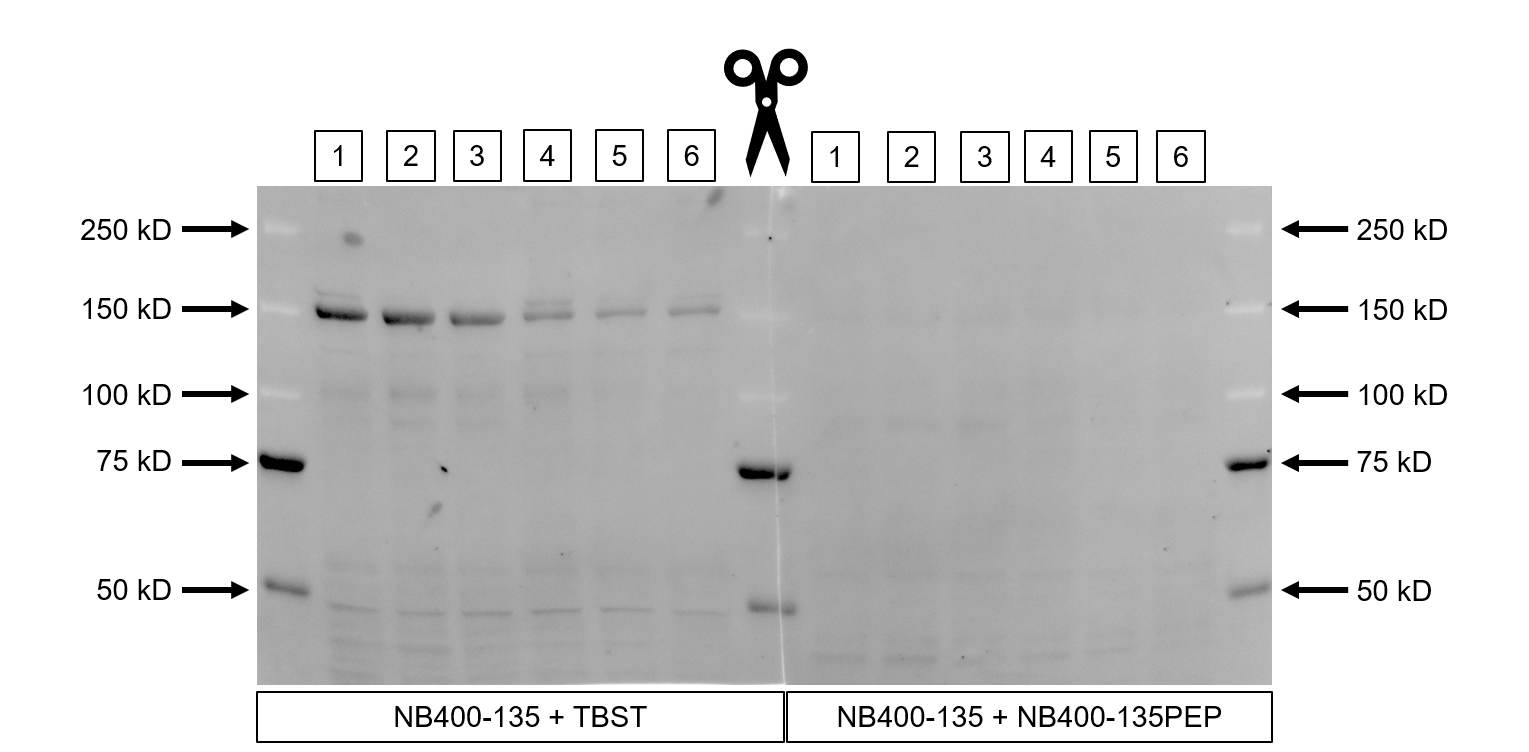
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - Mouse liver tissue lysate, 10 ug total protein per lane, probed with CHREBP antibody. Specific bands at ~150 kDa (some wells as doublets). Weaker bands at ~45 kDa, very faint bands at ~100 kDa. These bands were absent after exposure to ChREBP blocking peptide, although some faint non-specific banding remained at ~90, ~60, and ~40 kDa. WB image submitted by a verified customer review.Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135]
Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - Staining of human colon, epithelium at a 40X magnification.Western Blot: CHREBP AntibodyBSA Free [NB400-135]
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - Detection of ChREBP in 20 ug of human hepatocyte lysate using NB400-135. 5-10 second film exposure.Western Blot: CHREBP AntibodyBSA Free [NB400-135]
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - Detection of ChREBP in liver nuclear extracts from well-fed rats. 7% SDS-PAGE gel, 1:1000 dilution of NB400-135. Photo courtesy of Dr. Uyeda, UT Southwestern University.Western Blot: CHREBP AntibodyBSA Free [NB400-135]
CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0019.jpgWestern Blot: CHREBP AntibodyBSA Free [NB400-135]
CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0020.jpgImmunocytochemistry/ Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135]
Immunocytochemistry/Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - HeLa cells were fixed for 10 minutes using 10% formalin and permeabilized for 5 minutes using 1X TBS + 0.5% Triton X-100. The cells were incubated with anti-ChREBP (NB400-135) at a 1:200 dilution overnight at 4C and detected with an anti-rabbit DyLight 488 (Green) at a 1:500 dilution. Alpha tubulin was used as a co-stained at 1:1000 and detected with an anti-mouse DyLight 550 (Red) at a 1:500. Nuclei were counterstained with DAPI (Blue). Cells were imaged using a 40X objective.Immunohistochemistry: CHREBP Antibody - BSA Free [NB400-135]
Immunohistochemistry: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody [NB400-135] - Analysis of CHREBP in mouse liver using DAB with hematoxylin counterstain.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Analysis of protein levels in livers transfected with GK and GKA456V. Livers excised from mice in postabsorptive state were homogenized and resolved by Western blot. (a, b) Representative blots from three independent experiments. The densitometric analysis is presented (N = 4, *P < 0.05 versus pControl, ***P < 0.001 versus pGKA456V). (c) Liver sections of 5-hour fasted mice injected with pControl, pGK, or pGKA456V were immunostained with Glc6Pase antibody. TO-PRO-3 was used to visualize nuclei.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Immunoblot analysis of livers from control & L-Chrebp−/− mice subjected to fasting & refeeding with a high-sucrose diet. Littermate control & L-Chrebp−/− mice (same as those described in supplemental Tables S2A & S2B) were subjected to fasting & refeeding. The nonfasted (N) groups were fed chow diet ad libitum. The fasted (F) group was fasted 12 h, & the refed (R) group was fasted for 12 h & then refed with 60% (w/w) high-sucrose diet for 12 h prior to study. Liver whole-cell lysates & membrane fractions were prepared individually, & equal amounts of protein from each mouse of the same group (four per group) were pooled. Aliquots (40 μg for whole-cell lysates & 30 μg for membrane fractions) of the pooled protein were subjected to SDS-PAGE & immunoblot analysis. Immunoblot analysis of Insig-1 & Insig-2 were carried out using membrane fractions. Whole-cell lysates were used to detect other proteins. The precursor & nuclear forms of SREBPs are denoted as P & N, respectively. Calnexin was used as loading control. Image collected & cropped by CiteAb from the following publication (https://linkinghub.elsevier.com/retrieve/pii/S0022227520331369), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Raffinose stimulates transcription of genes involved in epidermal barrier function in HaCaT cells.(a & b) HaCaT cells were treated with vehicle, 1 μM or 10 μM raffinose (Raf), or 1 μM TO901317 (T17) for 24 h. Expressions of transcripts (a) & proteins (b) were analyzed by qRT-PCR or western blotting, respectively. FLG; filaggrin. IVL; involucrin, LOR; loricrin, AQP3; aquaporin3. (c) HaCaT cells were transfected with siGFP control, siLXR alpha, or siLXR beta, & then treated with 1 μM raffinose for 24 h. Expression of proteins was analyzed by western blotting. The original blots are shown in Supplementary Fig. S9. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/28266648), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] -
Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] - Impaired hepatic SIRT1-FoxO signaling in L-G6pc-/- mice.(A) Western blots & densitometry analysis (n = 5), & quantification of mRNA for hepatic SIRT1 & FoxO3a (n = 8). (B) Hepatic NAD+ levels (n = 9). (C) Western blots & densitometry analysis of PPAR-gamma, PPAR-alpha & beta -actin (n = 5). (D) Immunohistochemical analysis of hepatic ChREBP & quantification of nuclear ChREBP-translocated cells (n = 4). Scale bar, 25 μm. (E) Quantification of mRNA for hepatic Acaca, Fasn & Elovl6 by real-time RT-PCR (n = 6). (F) Western blots of acetylated & total FoxO3a after immunoprecipitation of nuclear extracts using anti-FoxO3a, & quantification of the acetylated FoxO3a/total FoxO3a (n = 5). Data represent the mean ± SEM. *P < 0.05, **P < 0.005. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/28558013), licensed under a CC0-1.0 license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Expressing recombinant ChREBP beta in Rictor-deficient adipocytes rescues expression of DNL enzymes.Western blot of indicated proteins in differentiated adipocytes with or without Rictor deletion transfected with various rescue constructs that were stably expressed in cells before differentiation. ‘*' indicates ChREBP beta based on molecular weight. Image collected & cropped by CiteAb from the following publication (https://www.nature.com/articles/ncomms11365), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Hepatic expression of mRNAs encoding carbohydrate response element-binding protein (ChREBP) (a), liver pyruvate kinase (LPK) (c), & microsomal triglyceride transfer protein (MTTP) (d), & nuclear ChREBP protein (b) in water-control, 10% fructose solution-control, & fructose pair-fed oleanolic acid- (OA-) treated rats at week 10. Animals were administered with OA (25 mg/kg/day) or vehicle (OA: 0 mg/kg, 5% Gum Arabic) by oral gavage daily for 10 weeks. mRNA was determined by real-time PCR. Protein expression was determined by Western blot. Data are means ± SEM (n = 6 each group). *P < 0.05. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/23737835), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - LXR mediate the induction of lipogenesis induced by an oleic acid-rich diet.(A) Hepatic Acly, Acaca, Acacb, Fasn, Elovl6, Scd1 mRNA levels quantified by qPCR. (B) Cytoplasmic protein expression levels of P-ACLY, ACLY, ACC, ELOVL6, SCD1, FASN & beta -ACTIN assayed by Western Blotting. (C) Fads1, Fads2, Elovl5, Gpat, Pnpla3 & Lpk mRNA quantification assayed by qPCR. (D) Srebp-1c & Chrebp mRNA quantification assayed by qPCR. (E) Cytoplasmic & nuclear expression levels of LXR, SREBP-1c & ChREBP assayed by Western Blotting. Data are the mean±SEM of values measured in LXR+/+ & LXR-/- mice fed REF or OLIV diet. a Significant genotype effect. b Significant difference versus REF diet (n = 6 mice per group). Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/28732092), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Analysis of protein levels in livers transfected with GK & GKA456V. Livers excised from mice in postabsorptive state were homogenized & resolved by Western blot. (a, b) Representative blots from three independent experiments. The densitometric analysis is presented (N = 4, *P < 0.05 versus pControl, ***P < 0.001 versus pGKA456V). (c) Liver sections of 5-hour fasted mice injected with pControl, pGK, or pGKA456V were immunostained with Glc6Pase antibody. TO-PRO-3 was used to visualize nuclei. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/22194744), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - ChREBP knockdown inhibited glycolysis, lipogenesis & p21 in HT29 cells. (A) Inhibited relative mRNA expression of ChREBP alpha & ChREBP beta normalized to B2M after control or ChREBP siRNA transfection for 48 hours. (B) Western-blot of ChREBP, phospho-p53 & p21 & their quantification on the right. beta -actin was served as a loading control. Proteins were extracted from cells transfected with sicontrol & siChREBP after 48 hours. The quantification of western blot was normalized to beta -actin. (C) Decreased relative mRNA expression of glycolytic & lipogenic genes, normalized to B2M. (D) Relative mRNA expression of p53 & p21, normalized to B2M. (E) Relative mRNA expression of cell Cyclins, normalized to B2M. n = 4. *p < 0.05; **p < 0.01; ***p < 0.001. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/32144313), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Ligand-activated LXR reduces ChREBP binding to chromatin. (A). Left panels: Local pattern of LXR-ChREBP co-occupancy. Browser view of LXR & ChREBP tracks in the promoter region of Lpk (Pklr), Txnip, Fasn & Scd1. Square brackets indicate the scale maxima of ChIP/input ratios. Arrows indicate the genomic locations of quantitative RT-PCR primers. Right panel: AML12 cells transfected with ChREBP alpha /Mlx gamma & LXR alpha /RXR alpha were treated with DMSO (0.1%) or GW3965 (10 µM) for 18 h. ChREBP or LXR binding to genomic location indicated in the right panels were detected by ChIP using antibodies against ChREBP, LXR or IgG as negative control. Data are presented as mean ± SEM (n = 3–5). Significant differences are shown as * p < 0.05, ** p < 0.01, *** p < 0.001 compared to ChIP-IgG & #p < 0.05, ##p < 0.01, ###p < 0.001 between DMSO & GW3965 groups. ns, not significant. (B). CoIP of LXR alpha & ChREBP alpha, expressed in COS-1 cells cultured in 25 mM glucose, treated with DMSO (0.1%) or GW3965 (1 µM) for 18 h. Lysates were immunoprecipitated with ChREBP antibody (n = 3). Input & immunoprecipitated proteins were immunoblotted with ChREBP or LXR alpha antibodies. One representative western blot is shown. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/32414201), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Induction of hepatic Srebp-1 & carbohydrate response element-binding protein (Chrebp) beta expression by dietary glucose is reduced in LXR alpha −/− mice. (A) Hepatic gene expression of Lxr alpha / beta, Srebp-1, & Chrebp alpha / beta was analyzed by quantitative RT-PCR & normalized to Tbp; (B) Cytosolic & nuclear lysates were immunoblotted with antibodies against LXR, SREBP-1, & ChREBP with alpha -Tubulin & Lamin A as loading controls. Each lane represents independent mice from each group. One representative western blot is shown (n = 3). Data represent the mean ± SEM (n = 5). Significant differences were found using two-way ANOVA followed by Tukey’s multiple comparison test (fasted vs. fructose fed & fasted vs. glucose fed RNA data were analyzed separately). * p < 0.05, ** p < 0.01, *** p < 0.001 compared to fasted. #p < 0.05, ##p < 0.01, ###p < 0.001 compared to LXR alpha +/+ mice. Image collected & cropped by CiteAb from the following publication (http://www.mdpi.com/2072-6643/9/7/678), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Decreased glucose uptake & Chrebp beta -driven DNL is a primary consequence of Rictor loss in adipocytes.(a) Western blots of indicated proteins at different days of differentiation using Rictor iKO primary adipocytes (described in Methods). (b) Oil Red O staining of differentiated adipocytes. (c) The relative mRNA level of ChREBP beta in differentiated cells at various time points. n=3. (d) 2-DG uptake in differentiated adipocytes without or with insulin stimulation. n=3. (e) The C14-glucose-derived FA in differentiated adipocytes. n=3. (f) The C14-glucose derived triglyceride (TAG) in differentiated adipocytes. n=3. (g) Relative glut4 expression in differentiated cells at various time points. n=3. (h) Relative glut4 expression in sWAT. n=8. (i) Glycerol release in differentiated adipocytes under basal & isoproterenol (Iso) stimulation. n=4. (j) Glycerol release in ex vivo pgWAT under basal & isoproterenol stimulation. n=6. Data were analysed by Student's t-test. Values are expressed as mean+s.e.m. *P<0.05; **P<0.01; ***P<0.001. Image collected & cropped by CiteAb from the following publication (https://www.nature.com/articles/ncomms11365), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
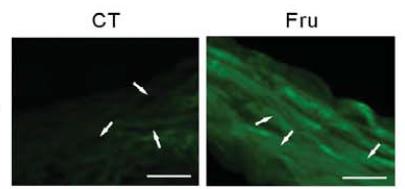
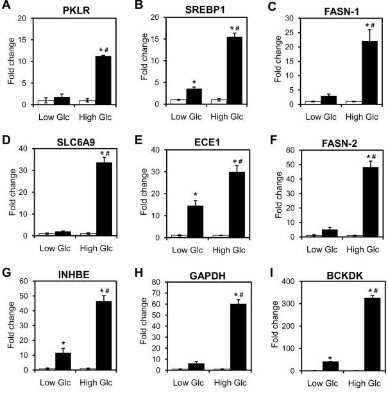
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - RetSat controls ChREBP activity & glucose sensing in primary hepatocytes. Primary mouse hepatocytes were treated with Control or RetSat siRNA for 48 h, (a, left) ChREBP mRNA expression determined by qPCR, & RetSat & ChREBP protein levels determined by immunoblotting (a, right). left, Data are shown as mean ± 11s.d., n = 3 independent transfections of hepatocyte cultures from the same mouse. Two independent experiments yielded similar results. b Hepatocytes were treated as described in a & mRNA expression of a selection of known ChREBP target genes visualized in a heatmap. c Primary hepatocytes were depleted of RetSat using two siRNA’s targeting different sites of the RetSat transcript for 48 h, & expression of the indicated genes analyzed by qPCR. Data are shown as mean ± s.d., n = 6 independent transfections of hepatocyte cultures from two different mice; *P < 0.05 between siControl und siRetSat by one-way ANOVA with Bonferroni post test. An independent experiment yielded similar results. d Hepatocytes treated with Control or RetSat siRNA were transfected with a ChoRE-Luc reporter, exposed to low & high glucose concentrations as indicated, & analyzed for luciferase activity. e Hepatocytes treated with Control or RetSat siRNA were exposed to low & high glucose concentrations as indicated, & mRNA expression determined by qPCR. In d, e, data are shown as mean ± s.d., n = 6 independent transfections of hepatocyte cultures from two mice; two-way ANOVA with Bonferroni post test revealed significances between low & high glucose concentrations (#P < 0.05) & between siControl & siRetSat (*P < 0.05). An independent experiment yielded similar results Image collected & cropped by CiteAb from the following publication (https://www.nature.com/articles/s41467-017-00430-w), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - LXR alpha & ChREBP alpha interact via key activation domains. (A). Top panel: Schematic representation of the ChREBP alpha full-length (FL), ChREBP beta & the low glucose inhibitory domain (LID) protein. Bottom panel: CoIP of LXR alpha & ChREBP alpha, ChREBP beta or LID, expressed in COS-1 cells cultured in 25 mM glucose. The ChREBP expression plasmids were transfected with a DNA ratio of ChREBP alpha :ChREBP beta :LID = 1:6:1, to obtain comparable protein levels. Lysates were immunoprecipitated with ChREBP, FLAG (for LID) or LXR alpha antibodies & input & immunoprecipitated proteins immunoblotted with the same antibodies (n = 3). One representative western blot is shown. LID, low-glucose inhibitory domain; GRACE, glucose-response activation conserved element; bHLH, basic helix-loop-helix domain: ZIP, leucine zipper. (B). Top panel: Schematic representation of the LXR alpha FL & truncations. Bottom panel: CoIP of ChREBP alpha & LXR alpha FL or truncations expressed in COS-1 cells cultured in 25 mM glucose. Lysates were immunoprecipitated with ChREBP antibody (n = 3). Input & immunoprecipitated proteins were immunoblotted with ChREBP or FLAG (for LXR alpha FL & truncations) antibodies. One representative western blot is shown. NTD, N-terminal domain; DBD, DNA-binding domain; LBD, ligand-binding domain. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/32414201), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - FXR inhibits glucose-induced proglucagon expression(a) Proglucagon qPCR on cDNA from GLUTag cells starved for 12h with lactate (10 mmol L−1) & then incubated for 24h in lactate (10 mmol L−1), glucose (5.6 mmol L−1) or 2-deoxyglucose (5.6 mmol L−1) media containing DMSO, GW4064 (5 μmol L−1) or CDCA (100 μmol L−1) (n=3; performed 3 times). Data are represented as mean +/− SD. Two-Way ANOVA analysis followed by Bonferronni’s posthoc test. ***P≤0.001: effect of GW4064 & CDCA on proglucagon mRNA levels in each medium conditions. §§§P≤0.001: effect of glucose on proglucagon mRNA levels in DMSO, GW4064 & CDCA conditions. (b) Chrebp qPCR on cDNA from FACS-sorted proglucagon-negative & proglucagon-positive cells from the ileum (ileum L−; ileum L+) & colon (colon L−; colon L+) of GLU-VENUS mice (lower panel; n=3) & ChREBP protein expression from cytoplasm & nucleus extract from GLUTag cells (upper panel; performed 3 times). Data are represented as mean +/− SD. Student t-test. *P≤0.05 & **P≤0.01 (c) ChREBP & FXR western-blots after FXR immunoprecipitation on lysates from cytoplasm & nucleus of GLUTag cells treated or not with GW4064 (5 μmol L-1) in presence or not of glucose (5.6 mmol L−1) (performed 2 times). (d) Proglucagon qPCR on cDNA from GLUTag cells electroporated with a siCtrl or siChrebp, starved for 12h with lactate (10 mmol L−1) & then incubated for 24h in lactate 10 mmol L−1 (Glc −) or glucose 5.6 mmol L−1 (Glc +) media supplemented with DMSO or GW4064 (5 μmol L−1) (n=3; performed 3 times). Data are represented as mean +/− SD. Two-Way ANOVA analysis followed by Bonferronni’s posthoc test. *P≤0.05 & **P≤0.01: effect of treatments on each transfection condition. §§§P≤0.001: effect of siChrebp in each treatment condition. Image collected & cropped by CiteAb from the following publication (https://www.nature.com/articles/ncomms8629), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Dietary fructose induces nuclear O-GlcNAc signaling. (A) Simplified schematic overview of the hexosamine signaling pathway including the rate limiting enzyme GFAT & O-GlcNAc transferase (OGT); (B) High-performance liquid chromatography (HPLC) analysis of hepatic UDP-GlcNAc levels. Left panel: Example of a typical running profile w/ or w/out injection of UDP-GlcNAc standard (Std.) is shown. Right panel: Quantification of the HPLC data normalized to protein concentration; (C) Hepatic expression of the Ogt gene & cytosolic & nuclear OGT protein analyzed by quantitative RT-PCR & western blotting/Image J & normalized to Tbp, alpha -tubulin, & Lamin A, respectively. Data represent the mean ± SEM (n = 5); (D) Cytosolic & nuclear lysates immunoblotted w/ anti-O-GlcNAc antibody (RL2) w/ alpha -Tubulin & Lamin A as loading controls. Each lane represents independent mice from each group. One representative western blot is shown (n = 2); (E) Nuclear lysates subjected to wheat germ agglutinin (WGA) beads to precipitate O-GlcNAcylated proteins. WGA enriched samples (upper panel) & input lysates (bottom panel) immunoblotted w/ antibodies detecting O-GlcNAcylated proteins (RL2), LXR & ChREBP. Each lane represents independent mice from experimental groups. Representative western blots are shown (n = 2); (F) ChREBP binding to carbohydrate response element (ChoRE) containing region of the L-pk promoter & negative control sequence (NC) 2216–2288 bp into L-pk gene after ChoRE sequence detected by chromatin immunoprecipitation (ChIP) using antibodies against ChREBP or IgG as a control. Data represent the mean ± SEM (n = 5). Significant differences found using two-way ANOVA followed by Tukey’s multiple comparison test. ** p < 0.01 compared to fasted. Image collected & cropped by CiteAb from the following publication (http://www.mdpi.com/2072-6643/9/7/678), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] -
Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] - Correction of hepatic G6Pase-alpha deficiency normalizes autophagy.L-G6pc-/- mice were treated with 1 x 1012 vp/kg of rAAV-G6PC at 4 WP & analyzed at 12 WP. (A) Hepatic G6Pase-alpha activity in control (n = 7), L-G6pc-/- (n = 7), & rAAV-treated L-G6pc-/- (AAV/ L-G6pc-/-, n = 8) mice. (B) Liver weights in control (n = 10), L-G6pc-/- (n = 5), & AAV/ L-G6pc-/- (n = 8) mice. (C) The levels of hepatic metabolites in control, L-G6pc-/- & AAV/ L-G6pc-/- (n = 8) mice. (D) Fasting glucose test (FGT) profile of control (n = 13), L-G6pc-/- (n = 6) & AAV/ L-G6pc-/- (n = 8) mice. (E) Western blots of hepatic SIRT1, FoxO3a, LC3B, p62 & beta -actin & densitometry analysis (n = 8). (F) Hematoxylin & eosin (H&E) stained liver sections, & immunohistochemical analysis of hepatic ChREBP & quantification of nuclear ChREBP-translocated cells in control, L-G6pc-/-, & rAAV-treated L-G6pc-/- (AAV/ L-G6pc-/-) mice (n = 4). The insets present higher magnification views. Scale bar, 25 μm. Data represent the mean ± SEM. *P < 0.05, **P < 0.005. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/28558013), licensed under a CC0-1.0 license. Not internally tested by Novus Biologicals.Western Blot: CHREBP Antibody - BSA Free [NB400-135] -
Western Blot: CHREBP Antibody - BSA Free [NB400-135] - Phf2 expression is increased in the liver of obese patients with benign hepatic steatosis & positively correlates with insulin sensitivity. Human liver biopsies from lean subjects with no fatty liver (NoFL) or obese patients with simple steatosis or NASH were obtained from the ABOS cohort. a Representative Western blot analysis of Phf2, ChREBP, SCD1, Nrf2, Gpx4, & alpha -SMA expression (n = 10 per group). b ChIP experiments for H3K9me2 levels & for the recruitment of ChREBP & the RNA polII at the SCD1 & Nrf2 promoter (n = 10 per group). c MUFA/SFA ratio reflecting SCD1 activity (n = 10 per group). d Expression of Nrf2 & Nrf2-regulated genes (n = 10 per group). e Levels of carbonylated proteins (n = 10 per group). f Expression of coll-Ia1, alpha -SMA, & TIMP-1 (n = 10 per group). All error bars represent mean ± SEM. Statistical analyses were made using Anova, followed by Bonferonni’s test. *Obese with steatosis compared to lean with noFL; P < 0.01. **Obese with steatosis compared to obese with NASH; P < 0.05 Image collected & cropped by CiteAb from the following publication (https://www.nature.com/articles/s41467-018-04361-y), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for CHREBP Antibody - BSA Free
Chromatin Immunoprecipitation
Chromatin Immunoprecipitation (ChIP)
Gel Super Shift Assays
Immunoblotting
Immunocytochemistry/ Immunofluorescence
Immunohistochemistry
Immunohistochemistry-Paraffin
Immunoprecipitation
Knockout Validated
SDS-Page
Western Blot
Reviewed Applications
Read 7 reviews rated 4 using NB400-135 in the following applications:
Formulation, Preparation, and Storage
Purification
Formulation
Format
Preservative
Concentration
Shipping
Stability & Storage
Background: CHREBP
Alternate Names
Gene Symbol
Additional CHREBP Products
Product Documents for CHREBP Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for CHREBP Antibody - BSA Free
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Citations for CHREBP Antibody - BSA Free
Customer Reviews for CHREBP Antibody - BSA Free (7)
Have you used CHREBP Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Application: Western BlotSample Tested: Liver, mouse liver lysate, Mouse Liver Tissue, Mouse Liver, Mouse Liver Lysates and C57BL/6J LiverSpecies: MouseVerified Customer | Posted 07/01/2021Specific bands at ~150kD (some wells as doublets). Weaker bands at ~45kD, very faint bands at ~100kD. These bands were absent after exposure to ChREBP blocking peptide, although some faint non-specific banding remained at ~90, ~60, & ~40kD.SDS-PAGE was performed using a Biorad mini-PROTEAN TGX 4-20% precast gel and separated proteins were transferred to a nitrocellulose membrane. Precision Plus Duel Standard ladder was added to either end and the middle of the gel. For samples, 10ug protein was loaded per well. After transfer the membrane was cut down the middle and each half was blocked for 1h at room temperature in TBST containing 3% BSA. The left half of the membrane was incubated at 4C overnight in NB400-135 (1:1000 prepared in TBST w/ 2.5% BSA), whereas the right half of the membrane was incubated at 4C overnight in both NB400-135 and blocking peptide (NB400-135PEP, 1ug/mL, prepared in TBST w/ 2.5% BSA). Membranes were washed in TBST, incubated in Alexa Fluor 488 anti-rabbit secondary antibody for 1h at room temperature (1:15,000 prepared in TBST w/ 3% BSA), washed again, and imaged using a BioRad ChemiDoc MP Imager.
-
Application: Western BlotSample Tested: Rat Liver Nuclear Protein ExtractSpecies: RatVerified Customer | Posted 06/16/2020ChREBP1:2000, overnight Secondary antibody: 1:20000
-
Application: Immunohistochemistry-FrozenSample Tested: aortic tissuesSpecies: MouseVerified Customer | Posted 09/01/2017Localization CHREBP protein in mice aortic tissuesThe frozen sections were permeabilized with 0.1% TritonX-100 and block with 10% goat serum. Sections were incubated with FITC-conjugated secondary antibodies.
-
Application: Western BlotSample Tested: human fibroblast whole cell extractSpecies: HumanVerified Customer | Posted 01/15/2013
-
Application: Western BlotVerified Customer | Posted 12/13/2012
-
Application: Chromatin ImmunoprecipitationSample Tested: Rat hepatocytesSpecies: RatVerified Customer | Posted 11/28/2012
-
Application: Western BlotSample Tested: Aorta whole tissue, Sample Amount: 20ugSpecies: MouseVerified Customer | Posted 02/02/2011
There are no reviews that match your criteria.
Protocols
View specific protocols for CHREBP Antibody - BSA Free (NB400-135):
Culture cells to appropriate density in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and wash the cells briefly in PBS. Add 10% formalin to the dish and fix at room temperature for 10 minutes.
2. Remove the formalin and wash the cells in PBS.
3. Permeablize the cells with 0.1% Triton X100 or other suitable detergent for 10 min.
4. Remove the permeablization buffer and wash three times for 10 minutes each in PBS. Be sure to not let the specimen dry out.
5. To block nonspecific antibody binding, incubate in 10% normal goat serum from 1 hour to overnight at room temperature.
6. Add primary antibody at appropriate dilution and incubate overnight at 4C.
7. Remove primary antibody and replace with PBS. Wash three times for 10 minutes each.
8. Add secondary antibody at appropriate dilution. Incubate for 1 hour at room temperature.
9. Remove secondary antibody and replace with PBS. Wash three times for 10 minutes each.
10. Counter stain DNA with DAPi if required.
Antigen Unmasking:
Bring slides to a boil in 10 mM sodium citrate buffer (pH 6.0) then maintain at a sub-boiling temperature for 10 minutes. Cool slides on bench-top for 30 minutes (keep slides in the sodium citrate buffer all the time).
Staining:
1. Wash sections in deionized water three times for 5 minutes each.
2. Wash sections in PBS for 5 minutes.
3. Block each section with 100-400 ul blocking solution (1% BSA in PBS) for 1 hour at room temperature.
4. Remove blocking solution and add 100-400 ul diluted primary antibody. Incubate overnight at 4 C.
5. Remove antibody solution and wash sections in wash buffer three times for 5 minutes each.
6. Add 100-400 ul HRP polymer conjugated secondary antibody. Incubate 30 minutes at room temperature.
7. Wash sections three times in wash buffer for 5 minutes each.
8. Add 100-400 ul DAB substrate to each section and monitor staining closely.
9. As soon as the sections develop, immerse slides in deionized water.
10. Counterstain sections in hematoxylin.
11. Wash sections in deionized water two times for 5 minutes each.
12. Dehydrate sections.
13. Mount coverslips.
1. Perform SDS-PAGE on samples to be analyzed, loading 10-25 ug of total protein per lane.
2. Transfer proteins to PVDF membrane according to the instructions provided by the manufacturer of the membrane and transfer apparatus.
3. Stain the membrane with Ponceau S (or similar product) to assess transfer success, and mark molecular weight standards where appropriate.
4. Rinse the blot TBS -0.05% Tween 20 (TBST).
5. Block the membrane in 5% Non-fat milk in TBST (blocking buffer) for at least 1 hour.
6. Wash the membrane in TBST three times for 10 minutes each.
7. Dilute primary antibody in blocking buffer and incubate overnight at 4C with gentle rocking.
8. Wash the membrane in TBST three times for 10 minutes each.
9. Incubate the membrane in diluted HRP conjugated secondary antibody in blocking buffer (as per manufacturer's instructions) for 1 hour at room temperature.
10. Wash the blot in TBST three times for 10 minutes each (this step can be repeated as required to reduce background).
11. Apply the detection reagent of choice in accordance with the manufacturers instructions.
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ChIP Protocol Video
- Chromatin Immunoprecipitation (ChIP) Protocol
- Chromatin Immunoprecipitation Protocol
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Immunoprecipitation Protocol
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
FAQs for CHREBP Antibody - BSA Free
-
Q: Pplease I want to block ChREBP Ab (NB400-135) using ChREBP peptide I need help to no the procedure and how much I need to add from the peptide to the ChREBP Ab
A: Here is a protocol for a blocking peptide control for a western blot: Materials and Reagents Blocking buffer (usually TBST plus either 5% non-fat dry milk or 3% BSA for Western blot, or PBS plus 1% BSA for IHC) - Antibody - Blocking (immunizing) peptide - Two tubes - Two identical samples (e.g. a Western blot with two identical lanes, cut in half; two slides containing the cells of interest; etc) Method 1. Determine the optimal concentration of antibody that consistently gives a positive result in your particular protocol. Using that concentration, determine how much antibody you will need for two experiments. a. For example, an antibody is being used successfully in Western blot at 0.5 ug/ml. You will need 2 ml of antibody solution to stain one strip of a Western blot. Thus, you would use 1 ug of antibody in 2 ml buffer for each strip. 2. Dilute the necessary amount of antibody in blocking buffer to the final volume needed for the two experiments. Divide this equally into two tubes. 3. In the first tube, labeled locked, add the blocking peptide to a final concentration of 1 ug/ml (2 ug total peptide in this example). In the second tube, labeled Control, add an equivalent amount of buffer. 4. Incubate both tubes, with agitation, at room temperature for 30 minutes, or overnight at 4 degrees C. 5. Perform the staining protocol on the two identical samples, using the blocked antibody for one and the control for the other. Be careful not to mix up the strips using the blocked and control antibodies! 6. Observe the staining. The staining that disappears when using the blocked antibody is specific to the antibody. (See note i) Notes i. If more than one band disappears in Western blot by peptide/antigen competition, those bands contain the antigenic determinants and could be fragments of the full antigen or a complex containing the antigen.

![Immunocytochemistry/ Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135] Immunocytochemistry/ Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Immunocytochemistry-Immunofluorescence-NB400-135-img0021.jpg)
![Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Immunohistochemistry-Paraffin-NB400-135-img0016.jpg)
![Western Blot: CHREBP AntibodyBSA Free [NB400-135] Western Blot: CHREBP AntibodyBSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0007.jpg)
![Western Blot: CHREBP AntibodyBSA Free [NB400-135] Western Blot: CHREBP AntibodyBSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0010.jpg)
![Western Blot: CHREBP AntibodyBSA Free [NB400-135] Western Blot: CHREBP AntibodyBSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0019.jpg)
![Western Blot: CHREBP AntibodyBSA Free [NB400-135] Western Blot: CHREBP AntibodyBSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Western-Blot-NB400-135-img0020.jpg)
![Immunocytochemistry/ Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135] Immunocytochemistry/ Immunofluorescence: CHREBP Antibody - BSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Immunocytochemistry-Immunofluorescence-NB400-135-img0017.jpg)
![Immunohistochemistry: CHREBP Antibody - BSA Free [NB400-135] Immunohistochemistry: CHREBP Antibody - BSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP-Antibody---BSA-Free-Immunohistochemistry-NB400-135-img0013.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] Knockout Validated: CHREBP Antibody - BSA Free [NB400-135]](https://resources.rndsystems.com/images/products/CHREBP Antibody - BSA Free-Knockout Validated-NB400-135-img0024.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-271220231253379.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-210202423452438.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202415291928.jpg)
![Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202415171345.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202415171318.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-31020241529337.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-31020241534316.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202415371929.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-31020241528455.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416171431.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-3102024161891.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416163714.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416165648.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416175271.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-31020241616057.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416171465.jpg)
![Immunohistochemistry-Paraffin: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416175247.jpg)
![Western Blot: CHREBP Antibody - BSA Free [NB400-135] - CHREBP Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb400-135_rabbit-polyclonal-chrebp-antibody-310202416163772.jpg)