PIEZO1 Antibody - BSA Free
Novus Biologicals | Catalog # NBP1-78446

![Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446] Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446]](https://resources.rndsystems.com/images/products/PIEZO1-Antibody-Immunocytochemistry-NBP1-78446-img0007.jpg)
Key Product Details
Species Reactivity
Validated:
Cited:
Applications
Validated:
Cited:
Label
Antibody Source
Format
Product Specifications
Immunogen
Reactivity Notes
Localization
Clonality
Host
Isotype
Scientific Data Images for PIEZO1 Antibody - BSA Free
Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446]
Immunocytochemistry/Immunofluorescence: PIEZO1 Antibody [NBP1-78446] - PIEZO1 antibody (Goat anti-rabbit 568) was tested in MDCK cells. Nuclei and B-actin were counterstained with DAPI (blue) and phalloidin (green). Image from verified customer review.Immunohistochemistry: PIEZO1 Antibody - BSA Free [NBP1-78446]
Immunohistochemistry: PIEZO1 Antibody [NBP1-78446] - Analysis of PIEZ01 in mouse epidermis using DAB with hematoxylin counterstain.Flow Cytometry: PIEZO1 Antibody - BSA Free [NBP1-78446]
Flow Cytometry: PIEZO1 Antibody [NBP1-78446] - An intracellular stain was performed on MCF7 cells with PIEZO1 Antibody NBP1-78446 (blue) and a matched isotype control NBP2-24891 (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 1.0 ug/mL for 30 minutes at room temperature, followed by Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Dylight 550 (SA5-10033, Thermo Fisher).Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446]
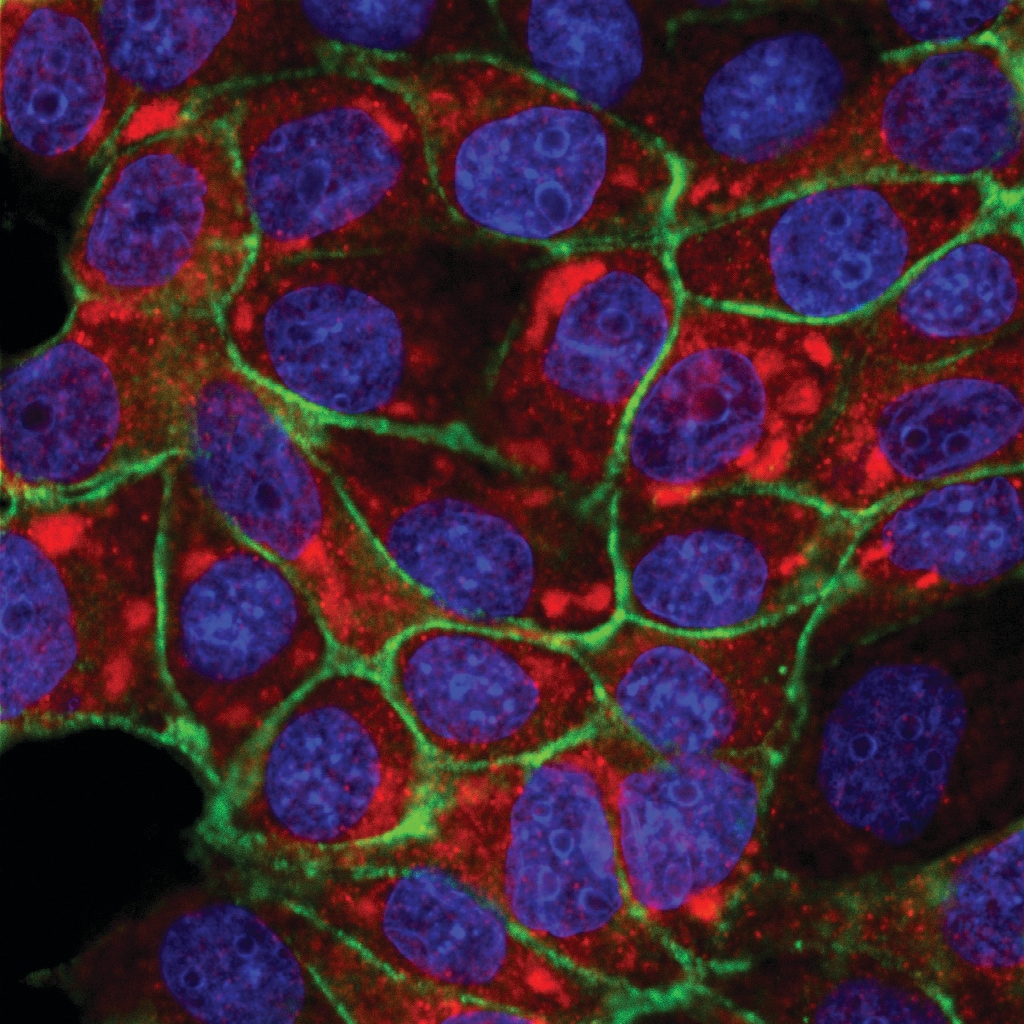
Immunocytochemistry/Immunofluorescence: PIEZO1 Antibody [NBP1-78446] - PIEZO1 antibody was tested in A431 cells with FITC (green). Nuclei and alpha-tubulin were counterstained with DAPI (blue) and Dylight 550 (red).PIEZO1 in MCF7 Human Cell Line.
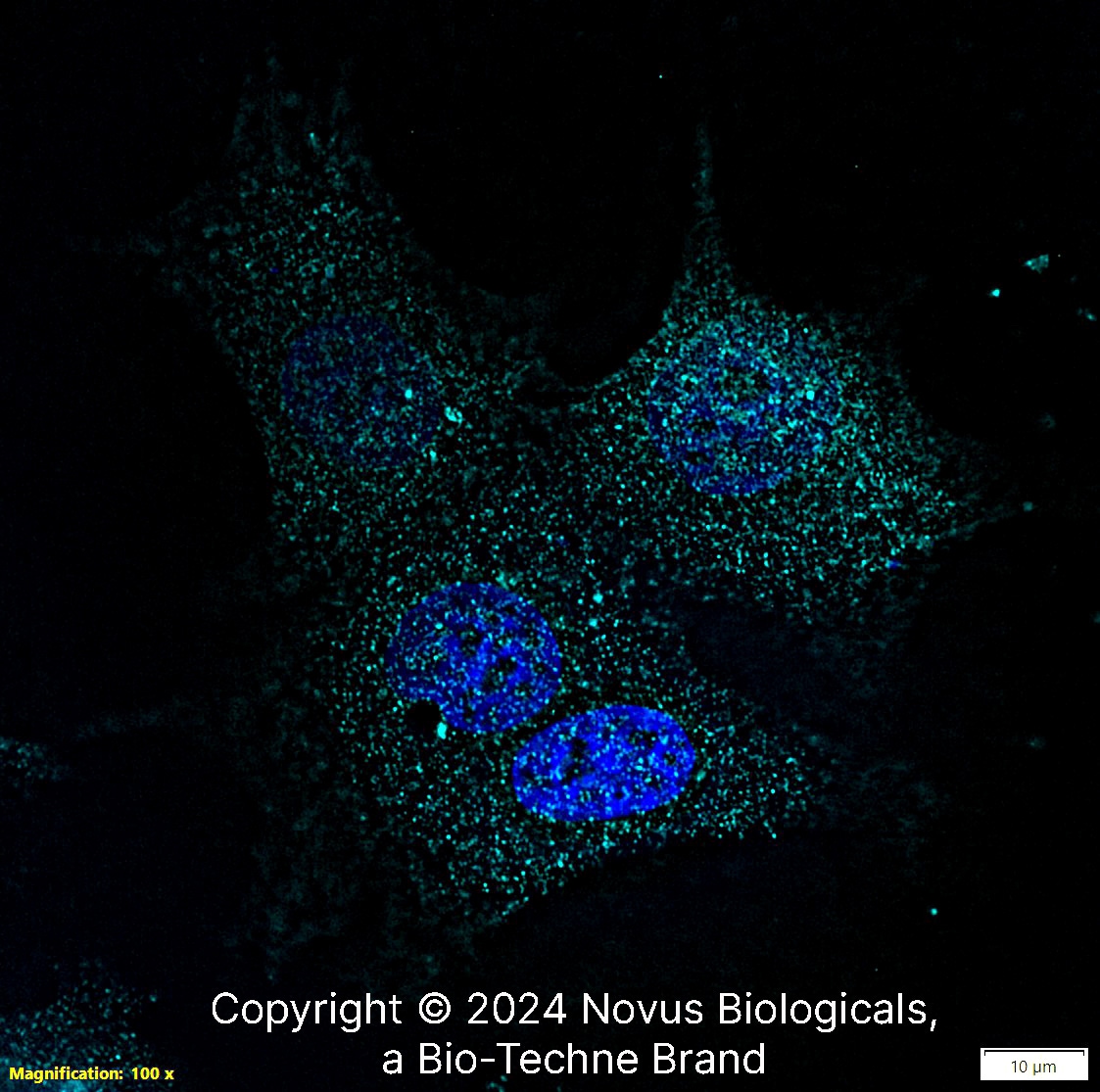
PIEZO1 was detected in immersion fixed MCF7 human breast cancer cell line using Rabbit anti-PIEZO1 Affinity Purified Polyclonal Antibody conjugated to DyLight 650 (Catalog # NBP1-78446C) (light blue) at 10 µg/mL overnight at 4C. Cells were stained counterstained with DAPI (blue). Cells were imaged using a 100X objective and digitally deconvolved.PIEZO1 in MCF7 Human Cell Line.
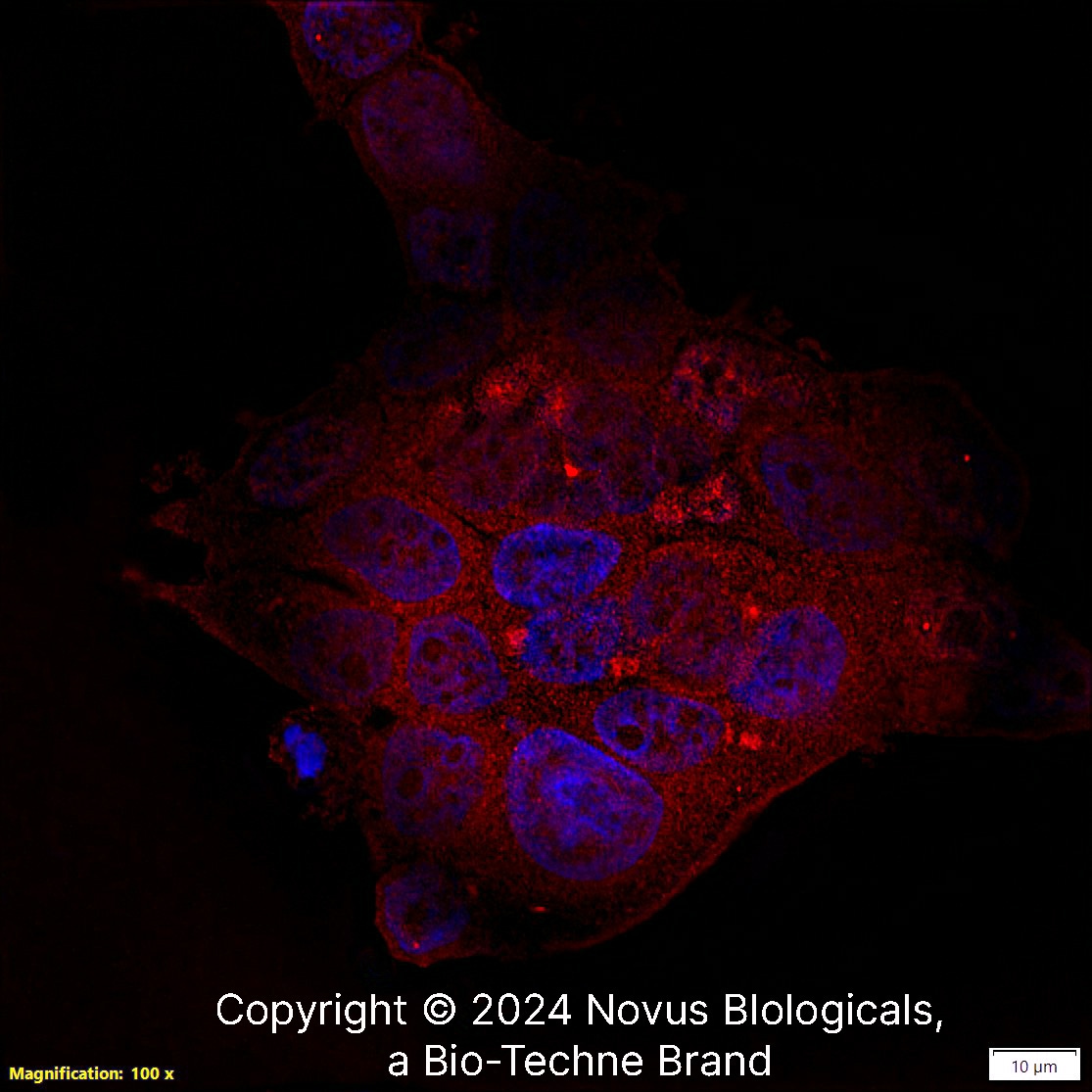
PIEZO1 was detected in immersion fixed MCF7 human breast cancer cell line using Rabbit anti-PIEZO1 Affinity Purified Polyclonal Antibody conjugated to Janelia Fluor® 549 (Catalog # NBP1-78446JF549) (red) at 10 µg/mL overnight at 4C. Cells were stained counterstained with DAPI (blue). Cells were imaged using a 100X objective and digitally deconvolved.Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] -
Immune response of macrophages after blocking Piezo1. (A) IF staining of CCR7 and CD206 in the Control, LPS and GsMTx4+LPS groups after 4-day-cultured. M1 macrophages were marked with CCR7 (red), M2 macrophages with CD206 (green), and nuclei with DAPI (blue). (B) Semi-quantitative analysis of CCR7 and CD163 in each group. (C) Flow cytometry analysis of RAW264. 7 cells in the Control, LPS and GsMTx4+LPS groups. Q6 represents M1 types (F4/80+/iNOS+) and Q10 represents M2 types (F4/80+/CD206+). (D) Expressions of Piezo1 and inflammation-related genes (Tnfa and Il1b) in macrophages cultured for 4 days. (E) Western blotting analysis of PIEZO1, IL-1B and TNF-A in RAW264. 7 cultured for 4 days. (F) Intracellular ROS levels of RAW264. 7 cultured for 4 days. (G) Concentration of inflammatory cytokines in macrophage medium detected by ELISA. *P < 0. 05; **P < 0. 01; ***P < 0. 001. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37261355), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] -
Ultrasound induces Piezo1 expression and localization to peripheral adhesion in MDA‐MB‐231 cancer cells. (a) Fluorescent images showing spatial distribution of microtubule, paxillin, and Piezo1 in the wild‐type (control, ultrasound‐ and nocodazole‐treated) and myosin IIA KD (control and ultrasound‐treated) cells. (b) Corresponding fluorescent intensity plot displaying paxillin and Piezo1 distribution in cells under different experimental condition described in panel a. (c) Western blot measuring the Piezo1 expression level in control and US‐treated cancer cells. (d) Western blot showing myosin IIA expression in control siRNA and myosin IIA KD cancer cells. (e) Western blot showing Piezo1 expression in control siRNA and Piezo1 KD cancer cells. (f) Fluorescent images showing microtubule network in myosin IIA KD cells and bar diagram demonstrating number of intact microtubules with and without US treatment. n = 15 cells. (g) Fluorescent images showing microtubule network and bar diagram showing number of intact microtubules in Piezo1 KD cells with and without US treatment, n = 25 cells. (h) Representative images illustrating microtubule network and Pieoz1 localization in control and US‐treated cells and bar diagram estimating cell population with peripheral Piezo1 in control and US‐treated cells, n > 204 cells. ****p < 0.0001. In all image panels, scale bar: 10 μm Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34589605), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] -
Ultrasound induces Piezo1 expression and localization to peripheral adhesion in MDA‐MB‐231 cancer cells. (a) Fluorescent images showing spatial distribution of microtubule, paxillin, and Piezo1 in the wild‐type (control, ultrasound‐ and nocodazole‐treated) and myosin IIA KD (control and ultrasound‐treated) cells. (b) Corresponding fluorescent intensity plot displaying paxillin and Piezo1 distribution in cells under different experimental condition described in panel a. (c) Western blot measuring the Piezo1 expression level in control and US‐treated cancer cells. (d) Western blot showing myosin IIA expression in control siRNA and myosin IIA KD cancer cells. (e) Western blot showing Piezo1 expression in control siRNA and Piezo1 KD cancer cells. (f) Fluorescent images showing microtubule network in myosin IIA KD cells and bar diagram demonstrating number of intact microtubules with and without US treatment. n = 15 cells. (g) Fluorescent images showing microtubule network and bar diagram showing number of intact microtubules in Piezo1 KD cells with and without US treatment, n = 25 cells. (h) Representative images illustrating microtubule network and Pieoz1 localization in control and US‐treated cells and bar diagram estimating cell population with peripheral Piezo1 in control and US‐treated cells, n > 204 cells. ****p < 0.0001. In all image panels, scale bar: 10 μm Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34589605), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] -
Immune response of macrophages after activating Piezo1. (A) IF staining of CCR7 and CD206 in the Control, LPS and Yoda1+LPS groups after 4-day-cultured. M1 macrophages were marked with CCR7 (red), M2 macrophages with CD206 (green), and nuclei with DAPI (blue). (B) Semi-quantitative analysis of CCR7 and CD206. (C) Flow cytometry analysis of RAW264. 7 cells in the Control, LPS and Yoda1+LPS groups. Q6 represents M1 types (F4/80+/iNOS+) and Q10 represents M2 types (F4/80+/CD206+). (D)Piezo1, Tnfa and Il1b genes expressions in macrophages cultured for 4 days. (E) Western blotting analysis of PIEZO1, TNF-A and IL-1B in RAW264. 7 cultured for 4 days. (F) Intracellular ROS levels of RAW264. 7 cultured for 4 days. (G) Concentration of inflammatory cytokines in macrophages medium detected by ELISA. *P < 0. 05; **P < 0. 01; ***P < 0. 001. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37261355), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446] -
Ultrasound induces Piezo1 expression and localization to peripheral adhesion in MDA‐MB‐231 cancer cells. (a) Fluorescent images showing spatial distribution of microtubule, paxillin, and Piezo1 in the wild‐type (control, ultrasound‐ and nocodazole‐treated) and myosin IIA KD (control and ultrasound‐treated) cells. (b) Corresponding fluorescent intensity plot displaying paxillin and Piezo1 distribution in cells under different experimental condition described in panel a. (c) Western blot measuring the Piezo1 expression level in control and US‐treated cancer cells. (d) Western blot showing myosin IIA expression in control siRNA and myosin IIA KD cancer cells. (e) Western blot showing Piezo1 expression in control siRNA and Piezo1 KD cancer cells. (f) Fluorescent images showing microtubule network in myosin IIA KD cells and bar diagram demonstrating number of intact microtubules with and without US treatment. n = 15 cells. (g) Fluorescent images showing microtubule network and bar diagram showing number of intact microtubules in Piezo1 KD cells with and without US treatment, n = 25 cells. (h) Representative images illustrating microtubule network and Pieoz1 localization in control and US‐treated cells and bar diagram estimating cell population with peripheral Piezo1 in control and US‐treated cells, n > 204 cells. ****p < 0.0001. In all image panels, scale bar: 10 μm Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34589605), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for PIEZO1 Antibody - BSA Free
Flow (Intracellular)
Flow Cytometry
Immunocytochemistry/ Immunofluorescence
Immunohistochemistry
Immunohistochemistry-Paraffin
Western Blot
Reviewed Applications
Read 1 review rated 3 using NBP1-78446 in the following applications:
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Formulation
Format
Preservative
Concentration
Shipping
Stability & Storage
Background: PIEZO1
Long Name
Alternate Names
Entrez Gene IDs
Gene Symbol
UniProt
Additional PIEZO1 Products
Product Documents for PIEZO1 Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for PIEZO1 Antibody - BSA Free
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Citations for PIEZO1 Antibody - BSA Free
Customer Reviews for PIEZO1 Antibody - BSA Free (1)
Have you used PIEZO1 Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Application: ImmunocytochemistrySample Tested: MDCK canine kidney epithelial cell lineSpecies: CanineVerified Customer | Posted 06/26/2017PIEZO1 antibody (Goat anti-rabbit 568) was tested in MDCK cells. Nuclei and B-actin were counterstained with DAPI (blue) and phalloidin (green). The new lot (A-2) doesn't stain this well.
There are no reviews that match your criteria.
Protocols
View specific protocols for PIEZO1 Antibody - BSA Free (NBP1-78446):
Sample Preparation.
1. Grow cells to 60-85% confluency. Flow cytometry requires between 2 x 105 and 1 x 106 cells for optimal performance.
2. If cells are adherent, harvest gently by washing once with staining buffer and then scraping. Avoid using trypsin as this can disrupt certain epitopes of interest. If enzymatic harvest is required, use Accutase, Collagenase, or TrypLE Express for a less damaging option.
3. Reserve 100 uL for counting, then transfer cell volume into a 50 mL conical tube and centrifuge for 8 minutes at 400 RCF.
a. Count cells using a hemocytometer and a 1:1 trypan blue exclusion stain to determine cell viability before starting the flow protocol. If cells appear blue, do not proceed.
4. Re-suspend cells to a concentration of 1 x 106 cells/mL in staining buffer (NBP2-26247).
5. Aliquot out 1 mL samples in accordance with your experimental samples.
Tip: When cell surface and intracellular staining are required in the same sample, it is advisable that the cell surface staining be performed first since the fixation and permeablization steps might reduce the availability of surface antigens.
Intracellular Staining.
Tip: When performing intracellular staining, it is important to use appropriate fixation and permeabilization reagents based upon the target and its subcellular location. Generally, our Intracellular Flow Assay Kit (NBP2-29450) is a good place to start as it contains an optimized combination of reagents for intracellular staining as well as an inhibitor of intracellular protein transport (necessary if staining secreted proteins). Certain targets may require more gentle or transient permeabilization protocols such as the commonly employed methanol or saponin-based methods.
Protocol for Cytoplasmic Targets:
Optional: Perform cell surface staining as described in the previous section.
1. Fix the cells by adding 100 uL fixation solution (such as 4% PFA) to each sample for 10-15 minutes.
2. Permeabilize cells by adding 100 uL of a permeabization buffer to every 1 x 106 cells present in the sample. Mix well and incubate at room temperature for 15 minutes.
a. For cytoplasmic targets, use a gentle permeabilization solution such as 1X PBS + 0.5% Saponin or 1X PBS + 0.5% Tween-20.
b. To maintain the permeabilized state throughout your experiment, use staining buffer + 0.1% of the permeabilization reagent (i.e. 0.1% Tween-20 or 0.1% Saponin).
3. Following the 15 minute incubation, add 2 mL of the staining buffer + 0.1% permeabilizer to each sample.
4. Centrifuge for 5 minutes at 400 RCF.
5. Discard supernatant and re-suspend in 1 mL of staining buffer + 0.1% permeabilizer.
6. Stain each sample at 1 uL/ 1 x 106 cells of primary antibody or 1-3 uL/ 1 x 106 cells for directly conjugated antibodies. Mix well and incubate at room temperature for 30 minutes- 1 hour. Gently mix samples every 10-15 minutes.
7. Following the primary/conjugate incubation, add 2 mL/sample of staining buffer +0.1% permeabilizer and centrifuge for 5 minutes at 400 RCF.
8. Remove supernatant and re-suspend each sample in 2 mL staining buffer + 0.1% permeabilizer, repeat wash for 5 minutes at 400 RCF.
9. If using a directly conjugated antibody, after the second wash, re-suspend cell pellet to a final volume of 500 uL per sample and proceed with flow analysis.
Culture cells to appropriate density in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and wash the cells briefly in PBS. Add 10% formalin to the dish and fix at room temperature for 10 minutes.
2. Remove the formalin and wash the cells in PBS.
3. Permeablize the cells with 0.1% Triton X100 or other suitable detergent for 10 min.
4. Remove the permeablization buffer and wash three times for 10 minutes each in PBS. Be sure to not let the specimen dry out.
5. To block nonspecific antibody binding, incubate in 10% normal goat serum from 1 hour to overnight at room temperature.
6. Add primary antibody at appropriate dilution and incubate overnight at 4C.
7. Remove primary antibody and replace with PBS. Wash three times for 10 minutes each.
8. Add secondary antibody at appropriate dilution. Incubate for 1 hour at room temperature.
9. Remove secondary antibody and replace with PBS. Wash three times for 10 minutes each.
10. Counter stain DNA with DAPi if required.
Immunohistochemistry-Paraffin Embedded Sections
Antigen Unmasking:
Bring slides to a boil in 10 mM sodium citrate buffer (pH 6.0) then maintain at a sub-boiling temperature for 10 minutes. Cool slides on bench-top for 30 minutes.
Staining:
1. Wash sections in deionized water three times for 5 minutes each.
2. Wash sections in wash buffer for 5 minutes.
3. Block each section with 100-400 ul blocking solution for 1 hour at room temperature.
4. Remove blocking solution and add 100-400 ul diluted primary antibody. Incubate overnight at 4C.
5. Remove antibody solution and wash sections in wash buffer three times for 5 minutes each.
6. Add 100-400 ul biotinylated diluted secondary antibody. Incubate 30 minutes at room temperature.
7. Remove secondary antibody solution and wash sections three times with wash buffer for 5 minutes each.
8. Add 100-400 ul Streptavidin-HRP reagent to each section and incubate for 30 minutes at room temperature.
9. Wash sections three times in wash buffer for 5 minutes each.
10. Add 100-400 ul DAB substrate to each section and monitor staining closely.
11. As soon as the sections develop, immerse slides in deionized water.
12. Counterstain sections in hematoxylin.
13. Wash sections in deionized water two times for 5 minutes each.
14. Dehydrate sections.
15. Mount coverslips.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow safe laboratory procedures.
1. Perform SDS-PAGE on samples to be analyzed, loading 10-25 ug of total protein per lane.
2. Transfer proteins to PVDF membrane according to the instructions provided by the manufacturer of the membrane and transfer apparatus.
3. Stain the membrane with Ponceau S (or similar product) to assess transfer success, and mark molecular weight standards where appropriate.
4. Rinse the blot TBS -0.05% Tween 20 (TBST).
5. Block the membrane in 5% Non-fat milk in TBST (blocking buffer) for at least 1 hour.
6. Wash the membrane in TBST three times for 10 minutes each.
7. Dilute primary antibody in blocking buffer and incubate overnight at 4C with gentle rocking.
8. Wash the membrane in TBST three times for 10 minutes each.
9. Incubate the membrane in diluted HRP conjugated secondary antibody in blocking buffer (as per manufacturer's instructions) for 1 hour at room temperature.
10. Wash the blot in TBST three times for 10 minutes each (this step can be repeated as required to reduce background).
11. Apply the detection reagent of choice in accordance with the manufacturer's instructions.
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Liperfluo
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
FAQs for PIEZO1 Antibody - BSA Free
-
Q: I am interested in purchasing the PIEZO 1 and PIEZO 2 antibodies and their blocking peptides from Novus. I have read your data sheets that in WBs the bands seen are approximately 286 kDa and 318 kDa, respectively. I would greatly appreciate if you could show me these WBs on human cells For PIEZO1 and 2 and the effects of the blocking peptides (or point me to publications using your Abs).
A: We offer a PIEZO1 blocking peptide specific to product NBP1-78446. We unfortunately do not offer a PIEZO2 blocking peptide. Our western blot data for NBP1-78446, NBP1-78624, and NBP1-78538 was performed by an outside collaborator and they have not given permission to us to release their images. That is why we do not have them available on our website. We will fully guarantee all of our products for any listed application or species. We unfortunately do not have any publications for these products either.

![Immunohistochemistry: PIEZO1 Antibody - BSA Free [NBP1-78446] Immunohistochemistry: PIEZO1 Antibody - BSA Free [NBP1-78446]](https://resources.rndsystems.com/images/products/PIEZO1-Antibody-Immunohistochemistry-NBP1-78446-img0005.jpg)
![Flow Cytometry: PIEZO1 Antibody - BSA Free [NBP1-78446] Flow Cytometry: PIEZO1 Antibody - BSA Free [NBP1-78446]](https://resources.rndsystems.com/images/products/PIEZO1-Antibody-Flow-Cytometry-NBP1-78446-img0008.jpg)
![Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446] Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446]](https://resources.rndsystems.com/images/products/PIEZO1-Antibody-Immunocytochemistry-Immunofluorescence-NBP1-78446-img0006.jpg)
![Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] - PIEZO1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-78446_rabbit-polyclonal-piezo1-antibody-western-blot-13220261444210.jpg)
![Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] - PIEZO1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-78446_rabbit-polyclonal-piezo1-antibody-western-blot-13220261531185.jpg)
![Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] - PIEZO1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-78446_rabbit-polyclonal-piezo1-antibody-western-blot-132202615255418.jpg)
![Western Blot: PIEZO1 Antibody - BSA Free [NBP1-78446] - PIEZO1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-78446_rabbit-polyclonal-piezo1-antibody-western-blot-132202615335823.jpg)
![Immunocytochemistry/ Immunofluorescence: PIEZO1 Antibody - BSA Free [NBP1-78446] - PIEZO1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nbp1-78446_rabbit-polyclonal-piezo1-antibody-immunocytochemistry-immunofluorescence-132202619442319.jpg)