Granulysin (formerly NKG5 or Lymphokine LAG-2) is a member of the saposin-like protein (SAPLIP) family of membrane disrupting proteins (1). Granulysin is expressed in granules of natural killer and activated cytotoxic T cells. It exhibits cytolytic activity against intracellular or extracellular microbes and also tumors, either alone or in synergy with perforin (2). Human granulysin has structural similarity and 30 - 40% aa identity to granulysins and NK-lysins of other mammals such as bovine, porcine and canine; similar peptides in rodents have not been identified (1). The 15 kDa unglycosylated protein contains five helical domains; helix 2 and 3 contain 9 arginines and one cysteine critical for activity. Peptides of either helix 2 or 3 will lyse bacteria, while helix 3 is needed to lyse tumor targets (3, 4). One isoform designated 519 uses a different start codon, has no signal peptide sequence and is poorly expressed (5). A post-translationally processed 9 kDa form is present in acidified granules and is less lytic than the 15 kDa form at granule pH (6). IL-15 is necessary and sufficient for granulysin upregulation in CD8 T cells (2). Nanomolar granulysin promotes chemotaxis and increases production of chemokines by monocytic cells, while micromolar local concentrations are needed for lysis (7). Experimental evidence supports the following mechanism for activity against intracellular pathogens (8). First, granulysin forms clusters in lipid rafts due to interaction of positive charges in helices 2-3 with acidic sphingolipids. After endocytosis, early endosomes fuse with phagosomes, probably regulated by small GTPase rab5. Granulysin binds microbial membranes through charge interactions and disrupts them, probably via scissoring actions of granulysin molecules (9, 10).
Human Granulysin Biotinylated Antibody
R&D Systems | Catalog # BAF3138

Key Product Details
Validated by
Biological Validation
Species Reactivity
Human
Applications
Western Blot, Intracellular Staining by Flow Cytometry
Label
Biotin
Antibody Source
Polyclonal Goat IgG
Loading...
Product Specifications
Immunogen
Mouse myeloma cell line NS0-derived recombinant human Granulysin
Arg23-Leu145
Accession # P22749
Arg23-Leu145
Accession # P22749
Specificity
Detects human Granulysin in Western blots. In Western blots, less than 1% cross-reactivity with recombinant human NKG-2D is observed.
Clonality
Polyclonal
Host
Goat
Isotype
IgG
Scientific Data Images for Human Granulysin Biotinylated Antibody
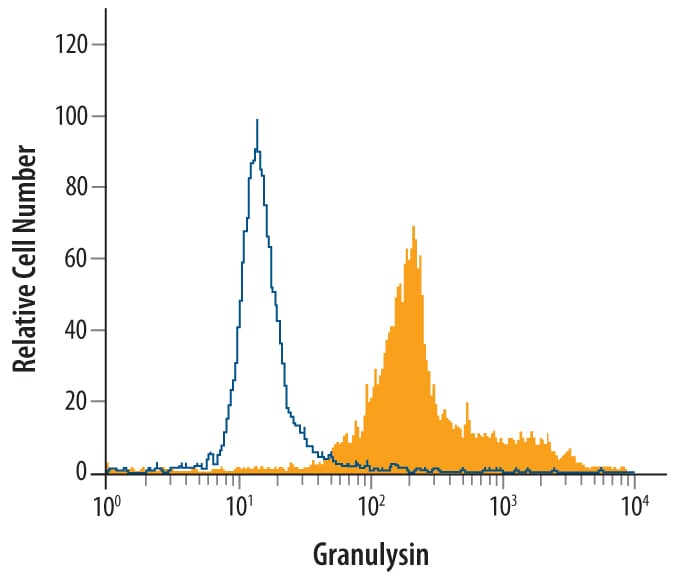
Detection of Granulysin in Human PBMCs by Flow Cytometry.
Human peripheral blood mononuclear cells were treated for 24 hr with 50 ng/mL PMA and 500 ng/mL Ca2+ionomycin, then stained with Goat Anti-Human Granulysin Biotinylated Antigen Affinity-purified Polyclonal Antibody (Catalog # BAF3138, filled histogram) or control antibody (Catalog # BAF108, open histogram), followed by Streptavidin-Phycoerythrin (Catalog # F0040).Applications for Human Granulysin Biotinylated Antibody
Application
Recommended Usage
Intracellular Staining by Flow Cytometry
2.5 µg/106 cells
Sample: Human peripheral blood mononuclear cells activated with PMA and Ca2+ ionomycin
Sample: Human peripheral blood mononuclear cells activated with PMA and Ca2+ ionomycin
Western Blot
0.1 µg/mL
Sample: Recombinant Human Granulysin
Sample: Recombinant Human Granulysin
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with BSA as a carrier protein.
Shipping
The product is shipped at ambient temperature. Upon receipt, store it immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: Granulysin
References
- Clayberger, C. and A.M. Krensky (2003) Curr. Opin. Immunol. 15:560.
- Ma, L.L. et al. (2002) J. Immunol. 169:5787.
- Wang, Z. et al. (2000) J. Immunol. 165:1486.
- Linde, C.M.A. et al. (2005) Infect. Immun. 73:6332.
- Yabe, T. et al. (1990) J. Exp. Med. 172:1159.
- Hanson, D.A. et al. (1999) Mol. Immunol. 36:413.
- Deng, A. et al. (2005) J. Immunol. 174:5243.
- Walch, M. et al. (2005) J. Immunol. 174:4220.
- Ernst, W.A. et al. (2000) J. Immunol. 165:7102.
- Anderson, D.H. et al. (2003) J. Mol. Biol. 325:355.
Alternate Names
GNLY, LAG2, Lymphokine, NKG5, TLA519
Entrez Gene IDs
10578 (Human)
Gene Symbol
GNLY
UniProt
Additional Granulysin Products
Product Documents for Human Granulysin Biotinylated Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human Granulysin Biotinylated Antibody
For research use only
Customer Reviews for Human Granulysin Biotinylated Antibody
There are currently no reviews for this product. Be the first to review Human Granulysin Biotinylated Antibody and earn rewards!
Have you used Human Granulysin Biotinylated Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Cellular Response to Hypoxia Protocols
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Liperfluo
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- R&D Systems Quality Control Western Blot Protocol
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...

