The human Macrophage Mannose Receptor (MMR), also known as CD206 and MRC1 (mannose receptor C, type 1), is a 190 kDa scavenger receptor that is expressed on tissue macrophages, myeloid dendritic cells, and liver and lymphatic endothelial cells (1). It belongs to a family of receptors sharing similar protein structure that also includes DEC205, phospholipase A2 receptor, and Endo180 (2, 3). The human MMR protein is synthesized as a 1456 amino acid (aa) precursor that contains an 18 aa signal sequence, a 1371 aa extracellular region, a 21 aa transmembrane segment and a 46 aa cytoplasmic domain (4). Its extracellular region is composed of an N-terminal cysteine-rich domain, followed by a single fibronectin type II repeat, and eight C-type lectin carbohydrate recognition domains (CRD) (3, 4). Human to mouse, the extracellular region is 82% aa identical. The cysteine-rich domain mediates recognition of sulfated N-acetylgalactosamine, which occurs on some extracellular matrix proteins and is the terminal sugar of the unusual oligosaccharides present on pituitary hormones such as lutropin and thyrotropin (5). Several of the CRDs participate in the Ca2+-dependent recognition of carbohydrates showing a preference for branched sugars with terminal mannose, fucose or N‑acetylglucosamine (6). The cytoplasmic domain of MMR includes a tyrosine-based motif for internalization in clathrin-coated vesicles. Once internalized, ligands are released following acidification of phagosomes or endosomes, and the receptor is recycled to the cell surface (3, 7). MMR mediates phagocytosis upon binding to target structures that occur on a variety of pathogenic microorganisms including Gram-negative and Gram-positive bacteria, yeasts, parasites, and mycobacteria. MMR also functions to maintain homeostasis through the endocytosis of potentially harmful glycoproteins associated with inflammation (2, 3).

Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human
Applications
Validated:
Immunocytochemistry
Cited:
Immunohistochemistry, Immunocytochemistry
Label
Unconjugated
Antibody Source
Monoclonal Rat IgG2A Clone # 309210
Loading...
Product Specifications
Immunogen
Mouse myeloma cell line NS0-derived recombinant human MMR
Leu19-Lys1383 (Thr399Ala) & (Leu407Phe)
Accession # P22897
Leu19-Lys1383 (Thr399Ala) & (Leu407Phe)
Accession # P22897
Specificity
Detects human MMR in direct ELISAs. In direct ELISAs, this antibody does not cross-react with recombinant mouse MMR.
Clonality
Monoclonal
Host
Rat
Isotype
IgG2A
Scientific Data Images for Human MMR/CD206 Antibody
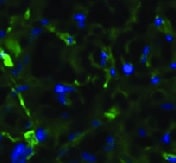
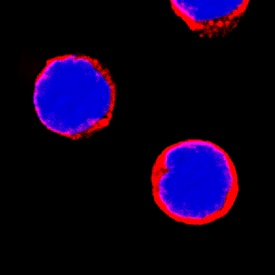
MMR/CD206 in Human PBMCs.
MMR/CD206 was detected in immersion fixed human peripheral blood mononuclear cells (PBMCs) using Rat Anti-Human MMR/CD206 Monoclonal Antibody (Catalog # MAB2534) at 8 µg/mL for 3 hours at room temperature. Cells were stained using the NorthernLights™ 557-conjugated Anti-Rat IgG Secondary Antibody (red; Catalog # NL013) and counterstained with DAPI (blue). Specific staining was localized to cell surfaces. View our protocol for Fluorescent ICC Staining of Non-adherent Cells.Applications for Human MMR/CD206 Antibody
Application
Recommended Usage
Immunocytochemistry
8-25 µg/mL
Sample: Immersion fixed human peripheral blood mononuclear cells
Sample: Immersion fixed human peripheral blood mononuclear cells
Reviewed Applications
Read 2 reviews rated 4.5 using MAB2534 in the following applications:
Formulation, Preparation, and Storage
Purification
Protein A or G purified from hybridoma culture supernatant
Reconstitution
Reconstitute at 0.5 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: MMR/CD206
References
- East, L. and C. Isake (2002) Biochim. Biophys. Acta 1572:364.
- Chieppa, M. et al. (2003) J. Immunol. 171:4552.
- Figdor, C. et al. (2002) Nat. Rev. Immunol. 2:77.
- Taylor, M. et al. (1990) J. Biol. Chem. 265:12156.
- Leteux, C. et al. (2000) J. Exp. Med. 191:1117.
- Martinez-Pomares, L. et al. (2001) Immunobiology 204:527.
- Feinberg, H. et al. (2000) J. Biol. Chem. 275:21539.
Long Name
Macrophage Mannose Receptor
Alternate Names
CD206, CLEC13D, MRC1
Gene Symbol
MRC1
UniProt
Additional MMR/CD206 Products
Product Documents for Human MMR/CD206 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human MMR/CD206 Antibody
For research use only
Citations for Human MMR/CD206 Antibody
Customer Reviews for Human MMR/CD206 Antibody (2)
4.5 out of 5
2 Customer Ratings
Have you used Human MMR/CD206 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
2 of
2 reviews
Showing All
Filter By:
-
Application: Immunocytochemistry/ImmunofluorescenceSample Tested: MacrophagesSpecies: HumanVerified Customer | Posted 11/02/2021
-
Application: Immunohistochemistry-FrozenSample Tested: See PMID 21943944Species: HumanVerified Customer | Posted 02/19/2015
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...