OGG1 Antibody - BSA Free
Novus Biologicals | Catalog # NB100-106

![Immunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-Paraffin-NB100-106-img0023.jpg)
Key Product Details
Validated by
Species Reactivity
Validated:
Cited:
Applications
Validated:
Cited:
Label
Antibody Source
Format
Product Specifications
Immunogen
Reactivity Notes
Clonality
Host
Isotype
Theoretical MW
Disclaimer note: The observed molecular weight of the protein may vary from the listed predicted molecular weight due to post translational modifications, post translation cleavages, relative charges, and other experimental factors.
Scientific Data Images for OGG1 Antibody - BSA Free
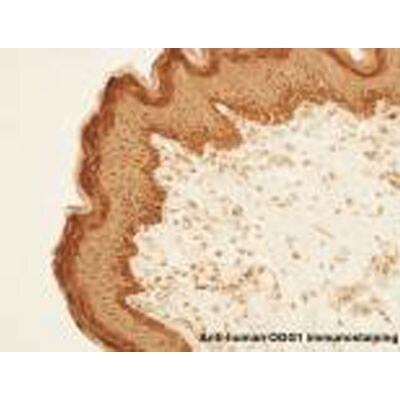
Immunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106]
Immunohistochemistry-Paraffin: OGG1 Antibody [NB100-106] - Analysis of OGG1 in human skin tissues. Image courtesy of product review by Arash Javeri.Western Blot: OGG1 AntibodyBSA Free [NB100-106]
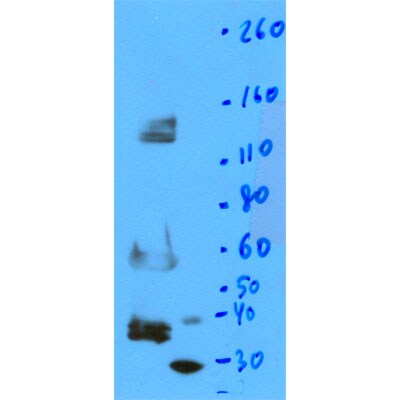
Western Blot: OGG1 Antibody [NB100-106] - - Analysis of Ogg1 expression in Jurkat whole cell lysate.Immunocytochemistry/ Immunofluorescence: OGG1 Antibody - BSA Free [NB100-106]
Immunocytochemistry/Immunofluorescence: OGG1 Antibody [NB100-106] - A431 cells were fixed for 10 minutes using 10% formalin and then permeabilized for 5 minutes using 1X TBS + 0.5% Triton-X100. The cells were incubated with anti-hOgg1 [NB100-106] at a 1:200 dilution overnight at 4C and detected with an anti-rabbit Dylight 488 (Green) at a 1:500 dilution. Alpha Tubulin (DM1A) [NB100-690] was used as a co-stain at a 1:1000 dilution and detected with an anti-mouse Dylight 550 (Red) at a 1:500 dilution. Nuclei were counterstained with DAPI (Blue). Cells were imaged using a 40X objective.Immunohistochemistry-Frozen: OGG1 Antibody - BSA Free [NB100-106]
Immunohistochemistry-Frozen: OGG1 Antibody [NB100-106] - Detection of OGG1 in formaldehyde-fixed frozen sections of the substantia nigra from a Rhesus macaque (Macaca mulatta) using NB100-106 at 10 ug/mL. Photo courtesy of Glen Kisby, Oregon Health Sciences University.Flow (Intracellular): OGG1 Antibody - BSA Free [NB100-106]
Flow (Intracellular): OGG1 Antibody [NB100-106] - An intracellular stain was performed on HeLa cells with NB100-106C (blue) and a matched isotype control (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 2.5 ug/mL for 30 minutes at room temperature. Both antibodies were conjugated to Dylight 650.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106]
OGG1-Antibody-Immunohistochemistry-NB100-106-img0033.jpgWestern Blot: OGG1 AntibodyBSA Free [NB100-106]
OGG1-Antibody-Western-Blot-NB100-106-img0034.jpgImmunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106]
OGG1-Antibody-Immunohistochemistry-Paraffin-NB100-106-img0035.jpgFlow Cytometry: OGG1 Antibody - BSA Free [NB100-106]
Flow Cytometry: OGG1 Antibody [NB100-106] - An intracellular stain was performed on HeLa cells with OGG1 Antibody NB100-106 (blue) and a matched isotype control NBP2-24891 (orange). Cells were fixed with 4% PFA and then permeabilized with 0.1% saponin. Cells were incubated in an antibody dilution of 1.0 ug/mL for 30 minutes at room temperature, followed by Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Dylight 550 (SA5-10033, Thermo Fisher).Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106]
Immunohistochemistry: OGG1 Antibody [NB100-106] - Staining of human tonsil, germinal center and mantle zone.Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106]
Flow Cytometry: OGG1 Antibody [NB100-106] - Baseline Ogg1 expression in human PBMC from a healthy donor. Image from verified.customer review.Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106]
Flow Cytometry: OGG1 Antibody [NB100-106] - An intracellular stain was performed on Jurkat Cells with NB100-106 and a matched isotype control. Cells were fixed with 4% PFA and then permeablized with 0.1% saponin. Cells were incubated in an antibody dilution of 2.5 ug/mL for 30 minutes at room temperature, followed by Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Dylight 550.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
The protein expression levels of HVL and LVL sorted cells by western blotting.(A) The ATM, phospho-ATM, GPX2, MRE11, OGG1, and XPC levels were measured by western blot analysis. The figure is representative of three independent experiments. (B) The relative fold activity was quantified using AlphaImage software and normalized to the expression of the internal actin control. *P < 0.05, **P < 0.01.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - The protein expression levels of HVL & LVL sorted cells by western blotting.(A) The ATM, phospho-ATM, GPX2, MRE11, OGG1, & XPC levels were measured by western blot analysis. The figure is representative of three independent experiments. (B) The relative fold activity was quantified using AlphaImage software & normalized to the expression of the internal actin control. *P < 0.05, **P < 0.01. Image collected & cropped by CiteAb from the following publication (https://dx.plos.org/10.1371/journal.pone.0164281), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - IHC-P analysis of hOGG1 protein expression in serous ovarian cancer: (A) Normal ovarian tissue treated with anti-hOGG1 antibody, biotinylated anti-rabbit IgG secondary antibody, & avidin-biotin-peroxidase complex, & counterstained with p-dimethylaminobenzaldehyde reagent (magnification 100 x); (B) serous cystadenoma with positive staining (magnification 200 x); (C) LG-SOC tissue with moderate positive staining (magnification 200 x); & (D) HG-SOC with negative staining (magnification 200 x). Black arrow pointed at the immunostained epithelial cells. Image collected & cropped by CiteAb from the following publication (https://ovarianresearch.biomedcentral.com/articles/10.1186/1757-2215-6-…), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - The H&E staining ((a)–(c)) & IHC staining for oxidative DNA damage, 8-oxo-dGuo ((d)–(g)), DNA repair enzyme, hOGG1 ((h)–(k)), & antioxidant proteins, CAT ((l)–(o)), GCLC ((p)–(s)), GPx ((t)–(w)), Nrf2 ((x)–(aa)), & MnSOD ((ab)–(ae)), in control subjects & tumor & nontumor lesions of BCC patients. Values given are mean ± SD. The statistical significance of differences between the control & case & between adjacent epidermis & tumor lesions of BCC patients was evaluated by nonparametric variables with Kruskal-Wallis test followed by Dunnett's post hoc test. ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001 compared to control; ##P < 0.01, ###P < 0.001 compared to tumor lesions of BCC patients. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/27057281), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - IHC-P analysis of hOGG1 protein expression in serous ovarian cancer: (A) Normal ovarian tissue treated with anti-hOGG1 antibody, biotinylated anti-rabbit IgG secondary antibody, & avidin-biotin-peroxidase complex, & counterstained with p-dimethylaminobenzaldehyde reagent (magnification 100 x); (B) serous cystadenoma with positive staining (magnification 200 x); (C) LG-SOC tissue with moderate positive staining (magnification 200 x); & (D) HG-SOC with negative staining (magnification 200 x). Black arrow pointed at the immunostained epithelial cells. Image collected & cropped by CiteAb from the following publication (https://ovarianresearch.biomedcentral.com/articles/10.1186/1757-2215-6-…), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - EBNA1 promotes the upregulation of oxidative DNA damage repair pathways in EBV converted cell lines & EBV positive BLs. a Representative western blots illustrating the expression of MTH1, OGG1, & MUTYH in pairs of EBV-negative & -positive cell lines. GAPDH was used as loading control. b Densitometry quantification of the specific bands. The intensity of the specific band in EBV positive cells relative the EBV-negative parental is shown. Mean ± SE of four independent experiments. c Representative western blots illustrating the expression of EBNA1, MTH1, OGG1, & MUTYH in the Mutu cell lines. d Densitometry quantification of expression in the EBV positive cell lines relative to the EBV-negative Mutu-30. Mean ± SE of four independent experiments Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/31511648), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - EBNA1 promotes the upregulation of oxidative DNA damage repair pathways in EBV converted cell lines & EBV positive BLs. a Representative western blots illustrating the expression of MTH1, OGG1, & MUTYH in pairs of EBV-negative & -positive cell lines. GAPDH was used as loading control. b Densitometry quantification of the specific bands. The intensity of the specific band in EBV positive cells relative the EBV-negative parental is shown. Mean ± SE of four independent experiments. c Representative western blots illustrating the expression of EBNA1, MTH1, OGG1, & MUTYH in the Mutu cell lines. d Densitometry quantification of expression in the EBV positive cell lines relative to the EBV-negative Mutu-30. Mean ± SE of four independent experiments Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/31511648), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - IHC-P analysis of hOGG1 protein expression in serous ovarian cancer: (A) Normal ovarian tissue treated with anti-hOGG1 antibody, biotinylated anti-rabbit IgG secondary antibody, & avidin-biotin-peroxidase complex, & counterstained with p-dimethylaminobenzaldehyde reagent (magnification 100 x); (B) serous cystadenoma with positive staining (magnification 200 x); (C) LG-SOC tissue with moderate positive staining (magnification 200 x); & (D) HG-SOC with negative staining (magnification 200 x). Black arrow pointed at the immunostained epithelial cells. Image collected & cropped by CiteAb from the following publication (https://ovarianresearch.biomedcentral.com/articles/10.1186/1757-2215-6-…), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - IHC-P analysis of hOGG1 protein expression in serous ovarian cancer: (A) Normal ovarian tissue treated with anti-hOGG1 antibody, biotinylated anti-rabbit IgG secondary antibody, & avidin-biotin-peroxidase complex, & counterstained with p-dimethylaminobenzaldehyde reagent (magnification 100 x); (B) serous cystadenoma with positive staining (magnification 200 x); (C) LG-SOC tissue with moderate positive staining (magnification 200 x); & (D) HG-SOC with negative staining (magnification 200 x). Black arrow pointed at the immunostained epithelial cells. Image collected & cropped by CiteAb from the following publication (https://ovarianresearch.biomedcentral.com/articles/10.1186/1757-2215-6-…), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - Immunoblot analysis of kidney lysates.(A) Blot membranes were incubated with the antibodies against Nrf2, p53 & 8-oxoguanine glycosylase alpha (OGG1 alpha ). The mean integrated optical density is related to actin for Nrf2 (B), p53 (C) & mature 8-oxoguanine glycosylase alpha (D) levels. Results are given as a mean ± S.E.M. KD4, KD6, KD8 – groups of Eker rats (Tsc2+/−) treated with HFKD for four, six or eight mo., respectively; ST – Eker rats fed with a standard diet; LE ST – wild-type Long Evans rats treated with a standard diet; LE KD – wild-type Long Evans rats treated with a ketogenic diet similarly to the KD6 group. *P < 0.05, **p < 0.01, ***p < 0.001 as compared to ST unless otherwise stated (horizontal line) for ANOVA with a Fisher post hoc test. Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/26892894), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - EBNA1 expression is associated with upregulation of MTH1 & oxidative damage repair pathways. The expression of MTH1, OGG1, & MUTYH was investigated by western blots & qPCR in BJAB-tTAE1 cells upon addition & withdrawal of doxycycline. a Representative western blots illustrating the correlation between the reversible down- & upregulation of MTH1 & EBNA1 in BJAB-tTAE1 cells upon addition & withdrawal of doxycycline. GAPDH was used as loading control. b Densitometry quantification of MTH1 & EBNA1 expression in BJAB-tTAE1 cells. The mean intensity of the MTH1 & EBNA1 specific bands relative to GAPDH in three independent experiments is shown in the figure. c Regression analysis of the relationship between expression levels of MTH1 & EBNA1. The data from three independent experiments were used for the plot. d Representative western blots illustrating the expression of MTH1, MUTYH, & OGG1 in BJAB-tTAE1 cells cultured for two weeks in the presence or absence of doxycycline. e Fold change is the ratio between the intensity of the specific band in cells cultured without or with doxycycline. The mean ± SD of four independent experiments is shown. f qPCR analysis of the levels of MTH1, OGG1, & MUTYH transcripts in BJAB-tTAE1 cells cultured for 2 weeks in the presence or absence of doxycycline. The mean ± SE of the fold change in six independent experiments each performed in triplicate is shown Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/31511648), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - The antioxidant pathways are activated during EBV infection & are required for growth transformation. Freshly isolated B-lymphocytes infected with the transforming B95-8 strain of EBV were cultured for up to 2 weeks in the presence or absence of MTH1 inhibitors. Protein expression was monitored by western blots, cell proliferation & activation of the DDR were assessed by 3H-Thy incorporation & staining for gamma H2AX, respectively. a representative western blots illustrating the parallel increase of MTH1 & EBNA1 expression following EBV infection & mean ± SE of the intensity of the MTH1 specific band in five independent experiments. b Representative western blots illustrating the expression of MTH1, MUTYH & OGG1 in ex vivo untreated B-cell & freshly EBV infected & SAC induced B blasts cultured for comparable times & showing similar levels of cell proliferation. c Quantification of the specific bands. Relative expression is the ration between the intensity in the treated cells versus freshly harvested cells. The mean ± SE of three to four independent experiments is shown. d Inhibition of MTH1 prevents the establishment of EBV transformed lymphoblastoid cell lines. 3H-Thy incorporation was measured after culture of freshly EBV infected B-lymphocytes in the presence of the indicated amounts of MTH1 inhibitors. Depending on the condition of the cultures harvesting was done after ten to fifteen days. e Inhibition of MTH1 strongly enhances the induction of DNA damage in freshly EBV infected cells. DNA damage was detected by gamma H2AX staining. f Mean ± SE of the % gamma H2AX positive cells in three independent experiments. *P < 0.05; **P < 0.01 Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/31511648), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] -
Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - Comparison of the expression of OGG1 (A) & PARP-1 (B) protein in normal colon tissue, polyp & cancer tissue of adenoma (AD, n = 68) & carcinoma (CRC, n = 103) patients.Immunohistochemical detection in paraffin embedded sections stained with hematoxylin & eosin. Center mark in the box indicates the medians of the samples. The length of each box (IQR, interquartile range) represents the range within which the central 50% of the values fell, with the vertical edges placed at the first & third quartiles. Whiskers show variability outside the upper & lower quartiles. P was obtained with the Mann-Whitney test. Representative examples of the levels of PARP-1 protein in tissues of CRC patients determined by Western analysis. The analysis was performed on tumor & normal tissues of 41 CRC patients (C). Image collected & cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/25526641), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Western Blot: OGG1 Antibody - BSA Free [NB100-106] -
Western Blot: OGG1 Antibody - BSA Free [NB100-106] - HDAC1 modulates OGG1 activity.a, b Representative images & quantification of OGG1 acetylation assessed by WB analysis using anti-acetylated lysine (Ac-Lysine, 4G12). Recombinant OGG1 (100 ng) first acetylated by p300 (25 ng), & incubated w/ titrated amount of HDAC1 (25 & 50 ng). a, b Four replicates from four independent experiments. c Reaction scheme of OGG1 cleavage assay. 5′IRDye800CW-labeled oligonucleotides (33 nt) contain an 8-oxoG base at position 17 & complementary unlabeled strand used as OGG1 substrate. Recombinant OGG1 proteins incubated w/ OGG1 substrate & resolved by 20% denaturing urea PAGE. d, e Representative images & quantification of OGG1 cleavage activity on reaction samples shown in Fig. 3a. d, e Seven replicates from four independent experiments. Nuclear extracts from HEK cells overexpressing OGG1 used as a positive control. f Co-immunoprecipitation of OGG1 w/ HDAC1 in cortical nuclear extracts from 2-month-old wild-type mice (n = 2) from one experiment. The asterisks indicate nonspecific cross-reaction w/ IgG heavy & light chains. g, h Representative images & quantification of acetylation status of endogenous OGG1 in hippocampal lysates of 13M control (n = 4) & Hdac1 cKO (n = 4) animals. g, h Two independent experiments. i Representative images of OGG1 cleavage assay using hippocampal nuclear extracts of 13M control & Hdac1 cKO mice. j Quantification of cleaved 17-nt products after OGG1 cleavage assay using hippocampal nuclear extracts of 13M control (n = 6) & Hdac1 cKO (n = 4) animals. i, j Two independent experiments. All values are shown as mean ± SEM. Statistical analysis: b, e one-way ANOVA w/ Tukey’s post hoc test; h, j two-tailed Student’s t test. Source data are provided as a Source Data file. Image collected & cropped by CiteAb from following publication (https://pubmed.ncbi.nlm.nih.gov/32424276), licensed under a CC-BY license. Not internally tested by Novus Biologicals.Applications for OGG1 Antibody - BSA Free
ELISA
Flow Cytometry
Immunoblotting
Immunocytochemistry/ Immunofluorescence
Immunohistochemistry-Frozen
Immunoprecipitation
Proximity Ligation Assay
Western Blot
Reviewed Applications
Read 6 reviews rated 4 using NB100-106 in the following applications:
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Formulation
Format
Preservative
Concentration
Shipping
Stability & Storage
Background: OGG1
Long Name
Alternate Names
Gene Symbol
UniProt
Additional OGG1 Products
Product Documents for OGG1 Antibody - BSA Free
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Product Specific Notices for OGG1 Antibody - BSA Free
This product is for research use only and is not approved for use in humans or in clinical diagnosis. Primary Antibodies are guaranteed for 1 year from date of receipt.
Related Research Areas
Citations for OGG1 Antibody - BSA Free
Customer Reviews for OGG1 Antibody - BSA Free (6)
Have you used OGG1 Antibody - BSA Free?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
-
Application: Western BlotSample Tested: 20 ug whole cell lysateSpecies: RatVerified Customer | Posted 12/12/2022OGG1
-
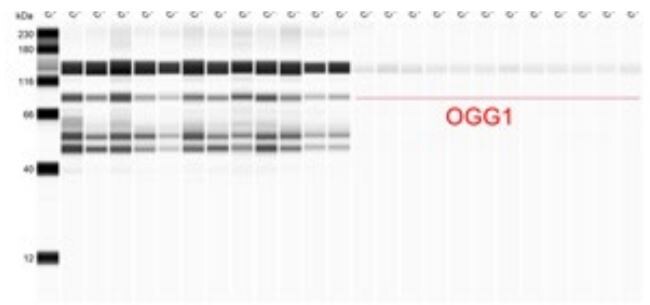
Application: Simple WesternSample Tested: mouse liver, Mouse Heart Tissue, Sample Amount: 100 ug and Mouse brainSpecies: MouseVerified Customer | Posted 05/19/2020Bio-Techne ResponseThis review was submitted through the legacy Novus Innovators Program, reflecting a new species or application tested on a primary antibody.
-
Application: ImmunoprecipitationSample Tested: MRC5 WCE- IP with Ogg1 ab and immunoblot with Ogg1 abSpecies: HumanVerified Customer | Posted 11/25/20141st lane-IP Ogg1, 2nd lane- Input (WCE), 3rd lane- IP Ogg1
-
Application: Flow CytometrySample Tested: Human PBMCSpecies: HumanVerified Customer | Posted 06/30/2014Baseline Ogg1 expression in human PBMC from a healthy donor.
-
Application: Western BlotSample Tested: human recombinant Ogg1, mouse skeletal muscle whole lysate, mouse spinal cord whole lysateSpecies: HumanVerified Customer | Posted 11/12/2012
-
Application: ImmunohistochemistrySample Tested: Human foreskin, adult ski, engineered human skin, keratinocytes, HaCaT cellsSpecies: HumanVerified Customer | Posted 05/17/2011
There are no reviews that match your criteria.
Protocols
View specific protocols for OGG1 Antibody - BSA Free (NB100-106):
Sample Preparation.
1. Grow cells to 60-85% confluency. Flow cytometry requires between 2 x 105 and 1 x 106 cells for optimal performance.
2. If cells are adherent, harvest gently by washing once with staining buffer and then scraping. Avoid using trypsin as this can disrupt certain epitopes of interest. If enzymatic harvest is required, use Accutase, Collagenase, or TrypLE Express for a less damaging option.
3. Reserve 100 uL for counting, then transfer cell volume into a 50 mL conical tube and centrifuge for 8 minutes at 400 RCF.
a. Count cells using a hemocytometer and a 1:1 trypan blue exclusion stain to determine cell viability before starting the flow protocol. If cells appear blue, do not proceed.
4. Re-suspend cells to a concentration of 1 x 106 cells/mL in staining buffer (NBP2-26247).
5. Aliquot out 1 mL samples in accordance with your experimental samples.
Tip: When cell surface and intracellular staining are required in the same sample, it is advisable that the cell surface staining be performed first since the fixation and permeablization steps might reduce the availability of surface antigens.
Intracellular Staining.
Tip: When performing intracellular staining, it is important to use appropriate fixation and permeabilization reagents based upon the target and its subcellular location. Generally, our Intracellular Flow Assay Kit (NBP2-29450) is a good place to start as it contains an optimized combination of reagents for intracellular staining as well as an inhibitor of intracellular protein transport (necessary if staining secreted proteins). Certain targets may require more gentle or transient permeabilization protocols such as the commonly employed methanol or saponin-based methods.
Protocol for Cytoplasmic Targets:
Optional: Perform cell surface staining as described in the previous section.
1. Fix the cells by adding 100 uL fixation solution (such as 4% PFA) to each sample for 10-15 minutes.
2. Permeabilize cells by adding 100 uL of a permeabization buffer to every 1 x 106 cells present in the sample. Mix well and incubate at room temperature for 15 minutes.
a. For cytoplasmic targets, use a gentle permeabilization solution such as 1X PBS + 0.5% Saponin or 1X PBS + 0.5% Tween-20.
b. To maintain the permeabilized state throughout your experiment, use staining buffer + 0.1% of the permeabilization reagent (i.e. 0.1% Tween-20 or 0.1% Saponin).
3. Following the 15 minute incubation, add 2 mL of the staining buffer + 0.1% permeabilizer to each sample.
4. Centrifuge for 5 minutes at 400 RCF.
5. Discard supernatant and re-suspend in 1 mL of staining buffer + 0.1% permeabilizer.
6. Stain each sample at 1 uL/ 1 x 106 cells of primary antibody or 1-3 uL/ 1 x 106 cells for directly conjugated antibodies. Mix well and incubate at room temperature for 30 minutes- 1 hour. Gently mix samples every 10-15 minutes.
7. Following the primary/conjugate incubation, add 2 mL/sample of staining buffer +0.1% permeabilizer and centrifuge for 5 minutes at 400 RCF.
8. Remove supernatant and re-suspend each sample in 2 mL staining buffer + 0.1% permeabilizer, repeat wash for 5 minutes at 400 RCF.
9. If using a directly conjugated antibody, after the second wash, re-suspend cell pellet to a final volume of 500 uL per sample and proceed with flow analysis.
Immunocytochemistry Protocol
Culture cells to appropriate density on suitable glass coverslips in 35 mm culture dishes or 6-well plates.
1. Remove culture medium and add 10% formalin to the dish. Fix at room temperature for 5-10 minutes.
2. Remove the formalin and add 0.5% Triton-X 100 in TBS to permeabilize the cells. Incubate for 5-10 minutes.
3. Remove the permeabilization buffer and add wash buffer (i.e. PBS or PBS with 0.1% Tween-20). Be sure to not let the specimen dry out. Gently wash three times for 10 minutes.
4. Alternatively, cells can be fixed with -20C methanol for 10 min at room temperature. Remove the methanol and rehydrate in PBS for 10 min before proceeding.
5. To block nonspecific antibody binding incubate in 10% normal goat serum for 1 hour at room temperature.
6. Add primary antibody at appropriate dilution and incubate at room temperature for 1 hour or at 4 degrees C overnight.
7. Remove primary antibody and replace with wash buffer. Gently wash three times for 10 minutes.
8. Add secondary antibody at the appropriate dilution. Incubate for 1 hour at room temperature.
9. Remove antibody and replace with wash buffer. Gently wash three times for 10 minutes.
10. Nuclei can be staining with 4',6' diamino phenylindole (DAPI) at 0.1 ug/ml, or coverslips can be directly mounted in media containing DAPI.
11. Cells can now be viewed with a fluorescence microscope.
*The above information is only intended as a guide. The researcher should determine what protocol best meets their needs. Please follow proper laboratory procedures for the disposal of formalin.
Immunohistochemistry - FFPE sections
I. Deparaffinization:
A. Treat slides with Xylene: 3 changes for 5 minutes each. Drain slides for 10 seconds between changes.
B. Treat slides with 100% Reagent Alcohol: 3 changes for 5 minutes each. Drain slides for 10 seconds between changes.
II. Quench Endogenous Peroxidase:
A. Place slides in peroxidase quenching solution: 15-30 minutes.
To Prepare 200 ml of Quenching Solution:
Add 3 ml of 30% Hydrogen Peroxide to 200 ml of Methanol.
Use within 4 hours of preparation
B. Place slides in distilled water: 2 changes for 2 minutes each.
III. Retrieve Epitopes:
A. Preheat Citrate Buffer. Place 200 ml of Citrate Buffer Working Solution into container, cover and place into steamer. Heat to 90-96 degrees Celcius.
B. Place rack of slides into hot Citrate Buffer for 20 minutes. Cover.
C. Carefully remove container with slides from steamer and cool on bench, uncovered, for 20 minutes.
D. Slowly add distilled water to further cool for 5 minutes.
E. Rinse slides with distilled water. 2 changes for 2 minutes each.
IV. Immunostaining Procedure:
A. Remove each slide from rack and circle tissue section with a hydrophobic barrier pen (e.g. Liquid Blocker-Super Pap Pen).
B. Flood slide with Wash Solution. Do not allow tissue sections to dry for the rest of the procedure.
C. Drain wash solution and apply 4 drops of Blocking Reagent to each slide and incubate for 15 minutes.
D. Drain Blocking Reagent (do not wash off the Blocking Reagent), apply 200 ul of primary antibody solution to each slide, and incubate for 1 hour.
E. Wash slides with Wash Solution: 3 changes for 5 minutes each.
F. Drain wash solution, apply 4 drops of Secondary antibody to each slide and incubate for 1 hour.
G. Wash slides with Wash Solution: 3 changes for 5 minutes each.
H. Drain wash solution, apply 4 drops of DAB Substrate to each slide and develop for 5-10 minutes. Check development with microscope.
I. Wash slides with Wash Solution: 3 changes for 5 minutes each.
J. Drain wash solution, apply 4 drops of Hematoxylin to each slide and stain for 1-3 minutes. Increase time if darker counterstaining is desired.
K. Wash slides with Wash Solution: 2-3 changes for 2 minutes each.
L. Drain wash solution and apply 4 drops of Bluing Solution to each slide for 1-2 minutes.
M. Rinse slides in distilled water.
N. Soak slides in 70% reagent alcohol: 3 minutes with intermittent agitation.
O. Soak slides in 95% reagent alcohol: 2 changes for 3 minutes each with intermittent agitation.
P. Soak slides in 100% reagent alcohol: 3 changes for 3 minutes each with intermittent agitation. Drain slides for 10 seconds between each change.
Q. Soak slides in Xylene: 3 changes for 3 minutes each with intermittent agitation. Drain slides for 10 seconds between each change.
R. Apply 2-3 drops of non-aqueous mounting media to each slide and mount coverslip.
S. Lay slides on a flat surface to dry prior to viewing under microscope.
NOTES:
Use treated slides (e.g. HistoBond) to assure adherence of FFPE sections to slide.
Prior to deparaffinization, heat slides overnight in a 60 degrees Celcius oven.
All steps in which Xylene is used should be performed in a fume hood.
For Epitope Retrieval, a microwave or pressure cooker may be substituted for the steamer method. Adjust times as necessary depending on conditions.
For the initial IHC run with a new primary antibody, test tissues with and without Epitope Retrieval. In some instances, Epitope Retrieval may not be necessary.
200 ul is the recommended maximum volume to apply to a slide for full coverage. Using more than 200 ul may allow solutions to wick off the slide and create drying artifacts. For small tissue sections less than 200 ul may be used.
5 minutes of development with DAB Substrate should be sufficient. Do not develop for more than 10 minutes. If 5 minutes of development causes background staining, further dilution of the primary antibody may be necessary.
Hematoxylin should produce a light nuclear counterstain so as not to obscure the DAB staining. Counterstain for 1-1 1/2 minutes for nuclear antigens. Counterstain for 2-3 minutes for cytoplasmic and membranous antigens. If darker counterstaining is desired increase time (up to 10 minutes).
Western Blot Procedure
1. Run 50 ug of protein on a 4-20% Tris-glycine mini-gel at 125V for 90 minutes.
2. Equilibrate gel, nitrocellulose membrane, Whatman paper, and blotting pads in transfer buffer for 15 minutes.
3. Transfer protein to the membrane at 25V for 90 minutes.
4. Allow membrane to air-dry.
5. Block membrane with 1XPBS/3% BSA for 1 hour at room temperature (23-27 degrees C).
6. Wash membrane twice, for 5 minutes each, with 1XPBS/0.05% Tween-20 (PBST).
7. Incubate membrane with NB100-106 (anti-hOGG1), diluted in 1XPBS/1% BSA, for 1 hour at room temperature.
8. Wash membrane once for 15 minutes, then four times for 5 minutes each, with PBST.
9. Incubate membrane with goat anti-rabbit IgG-HRP, diluted in 1XPBS/1% BSA, for 1 hour at room temperature.
10. Wash membrane once for 15 minutes, then four times for 5 minutes each, with PBST.
11. Detect cross-reacting proteins using Renaissance Chemiluminescence Reagent Plus kit from NEN Life Sciences.
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ELISA Sample Preparation & Collection Guide
- ELISA Troubleshooting Guide
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- How to Run an R&D Systems DuoSet ELISA
- How to Run an R&D Systems Quantikine ELISA
- How to Run an R&D Systems Quantikine™ QuicKit™ ELISA
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Immunoprecipitation Protocol
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Liperfluo
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- Quantikine HS ELISA Kit Assay Principle, Alkaline Phosphatase
- Quantikine HS ELISA Kit Principle, Streptavidin-HRP Polymer
- R&D Systems Quality Control Western Blot Protocol
- Sandwich ELISA (Colorimetric) – Biotin/Streptavidin Detection Protocol
- Sandwich ELISA (Colorimetric) – Direct Detection Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: ELISA
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
FAQs for OGG1 Antibody - BSA Free
-
Q: Is there a difference between Lot#J1 and Lot#L1 for Immunohistochemistry ? What does this product change for product specification between Lot#J1 and Lot#L1?
A: No, there should not have any significant differences between these two lots, J-1 and L-1 applying to any application. The different is the production time for the lots, as they could be bled, purified, and/or gone through other manufacturing processes at the different time points. However sometimes some minor issues do happen, and we call it "lot-to-lot" variations. These minor changes could readily due to how close the breeding time respective to the boosting injections (the closer, the higher titers of IgG), the healthy conditions of the hosts (infection or others), etc. But the specificities nor applicability of the different lots should not be changed.
-
Q: Would you please help confirm if NB100-106 would work with human OGG1 alpha?
A: NB100-106 has been validated for use in human, primate and rat for the following applications: Western Blot, IHC-Paraffin and IHC-Frozen. The immunogen used for this antibody maps to the N-terminal region between amino acids 1-100. I believe OGG1 isoforms differ in the C-terminal regional and all have a conserved N-terminus which would indicate to me that our antibody would recognize all isoforms. To be sure, take a close look at the isoform of interest in and compare its sequence to OGG1. If the differences lie outside the 1-100 amino acid range then we are confident the antibody will work for OGG1 alpha.
-
Q: Is there a difference between Lot#J1 and Lot#L1 for Immunohistochemistry ? What does this product change for product specification between Lot#J1 and Lot#L1?
A: No, there should not have any significant differences between these two lots, J-1 and L-1 applying to any application. The different is the production time for the lots, as they could be bled, purified, and/or gone through other manufacturing processes at the different time points. However sometimes some minor issues do happen, and we call it "lot-to-lot" variations. These minor changes could readily due to how close the breeding time respective to the boosting injections (the closer, the higher titers of IgG), the healthy conditions of the hosts (infection or others), etc. But the specificities nor applicability of the different lots should not be changed.
-
Q: Would you please help confirm if NB100-106 would work with human OGG1 alpha?
A: NB100-106 has been validated for use in human, primate and rat for the following applications: Western Blot, IHC-Paraffin and IHC-Frozen. The immunogen used for this antibody maps to the N-terminal region between amino acids 1-100. I believe OGG1 isoforms differ in the C-terminal regional and all have a conserved N-terminus which would indicate to me that our antibody would recognize all isoforms. To be sure, take a close look at the isoform of interest in and compare its sequence to OGG1. If the differences lie outside the 1-100 amino acid range then we are confident the antibody will work for OGG1 alpha.

![Western Blot: OGG1 AntibodyBSA Free [NB100-106] Western Blot: OGG1 AntibodyBSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Western-Blot-NB100-106-img0024.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-NB100-106-img0032.jpg)
![Immunocytochemistry/ Immunofluorescence: OGG1 Antibody - BSA Free [NB100-106] Immunocytochemistry/ Immunofluorescence: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunocytochemistry-Immunofluorescence-NB100-106-img0020.jpg)
![Immunohistochemistry-Frozen: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry-Frozen: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-Frozen-NB100-106-img0021.jpg)
![Flow (Intracellular): OGG1 Antibody - BSA Free [NB100-106] Flow (Intracellular): OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Flow-Intracellular-NB100-106-img0028.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-NB100-106-img0033.jpg)
![Western Blot: OGG1 AntibodyBSA Free [NB100-106] Western Blot: OGG1 AntibodyBSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Western-Blot-NB100-106-img0034.jpg)
![Immunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry-Paraffin: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-Paraffin-NB100-106-img0035.jpg)
![Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106] Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Flow-Cytometry-NB100-106-img0036.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Immunohistochemistry-NB100-106-img0022.jpg)
![Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106] Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Flow-Cytometry-NB100-106-img0026.jpg)
![Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106] Flow Cytometry: OGG1 Antibody - BSA Free [NB100-106]](https://resources.rndsystems.com/images/products/OGG1-Antibody-Flow-Cytometry-NB100-106-img0029.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-271220231253388.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415345339.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415291933.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-31020241538247.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415392553.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415363736.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-31020241535637.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415293311.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415175230.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-31020241537192.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-3102024155529.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-310202415523940.jpg)
![Immunohistochemistry: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-3102024165727.jpg)
![Western Blot: OGG1 Antibody - BSA Free [NB100-106] - OGG1 Antibody - BSA Free](https://resources.rndsystems.com/images/products/nb100-106_rabbit-polyclonal-ogg1-antibody-31020241682186.jpg)



-(01-ml)_NB100-106_8496.png)