PAR/pADPr Antibody
R&D Systems | Catalog # 4336-APC-050

Discontinued Product
4336-APC-050 has been discontinued.
An alternative/replacement product is available:
4335-MC-100. View all PAR/pADPr products.
Key Product Details
Species Reactivity
Validated:
Multi-Species
Cited:
Human, Mouse
Applications
Validated:
Western Blot
Cited:
Western Blot
Label
Unconjugated
Antibody Source
Polyclonal Rabbit IgG
Loading...
Product Specifications
Immunogen
Poly(ADP-ribose) polymer
Specificity
This polyclonal antibody detects free PAR and poly-ribosylated proteins.
Clonality
Polyclonal
Host
Rabbit
Isotype
IgG
Scientific Data Images for PAR/pADPr Antibody
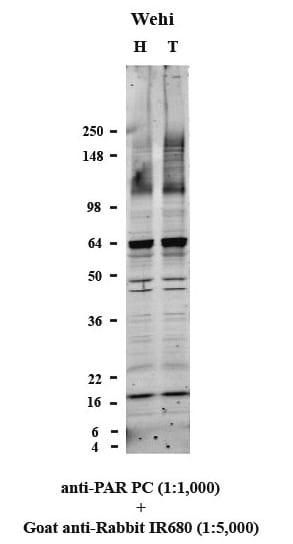
Detection of PAR/pADPr by Western Blot.
Western blot analysis of Wehi cells untreated (H) and treated (T) with 25 μM Etoposide for 4 hours at 37 °C. Cells were lysed in Tris-Glycine SDS sample buffer at the concentration of 1 x 107cells/ml, and 10 μl of clarified lysate were loaded per well of 4-20% Tris-Glycine gel. Proteins were transferred onto an Immobilon FL membrane and ribosylated proteins were detected using Trevigen's polyclonal anti-PAR antibody (cat# 4336-APC-050) followed by an IR680-conjugated secondary antibody (Licor). The membrane was scanned using an Odyssey Infrared Imaging System (Licor).Applications for PAR/pADPr Antibody
Application
Recommended Usage
Western Blot
1:1000 dilution
Sample: Wehi cells treated with Etoposide
Sample: Wehi cells treated with Etoposide
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Formulation
This antibody is an affinity purified IgG fraction in 1X PBS, containing 50%
glycerol.
Shipping
The product is shipped with polar packs. Upon receipt, store it immediately at the temperature recommended below.
Stability & Storage
-20 °C (manual defrost freezer)
Background: PAR/pADPr
Blotting buffer: 12 mM Tris base, 96 mM Glycine, and 15% MeOH.
Blocking solution: 5% (w/v) nonfat dry milk in TBS-0.1% Tween.
Antibody solution: 5% (w/v) nonfat dry milk, in TBS-0.1% Tween.
Transfer the electrophoresed proteins to a PVDF membrane and incubate the membrane for 1 hour at room temperature in blocking solution.
Incubate the membrane overnight at 4°C in antibody solution containing a 1:1000 dilution of anti-PAR rabbit polyclonal antibody. Empirical determination of primary antibody concentration will be required for optimal results.
Wash the membrane at room temperature for 5 minutes with 3 changes of TBS-0.1% Tween. Changing the membrane containers often reduces background.
Incubate the membrane at room temperature for 1 hour in antibody solution containing antirabbit conjugated to horseradish peroxidase.
Empirical determination of secondary antibody concentration will be required for optimal results.
Wash the membrane for 5 minutes with 4 changes of TBS-0.1% Tween.
Develop peroxidase reaction using chemiluminescence.
References
- Affar, E.B., et al. 1998. Immunodot blot method for the detection of poly(ADP-ribose) synthesized in vitro and in vivo. Anal Biochem 259:280-3.
- Shah, G.M., et al. 1995. Methods for biochemical study of poly(ADP-ribose) metabolism in vitro and in vivo. Anal Biochem 227:1-13.
- Gagné, J-P., et al. 2008. Proteome-wide identification of poly(ADP-ribose) binding proteins and poly(ADP-ribose)-associated protein complexes. Nucleic Acids Res 36:6959-76.
Long Name
Poly [ADP-ribose] Polymer
Alternate Names
PAR
Additional PAR/pADPr Products
Product Documents for PAR/pADPr Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for PAR/pADPr Antibody
For research use only
Related Research Areas
Citations for PAR/pADPr Antibody
Customer Reviews for PAR/pADPr Antibody
There are currently no reviews for this product. Be the first to review PAR/pADPr Antibody and earn rewards!
Have you used PAR/pADPr Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Cellular Response to Hypoxia Protocols
- R&D Systems Quality Control Western Blot Protocol
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...
