FoxM1 (Forkhead box protein M1; also FKHL16, HFH11 and WIN) is a 90-100 kDa member of the winged helix transcription factor gene family. It is expressed in proliferating and tumor cells and promotes the transition from G1 to S phase and G2 to M phase within the cell cycle. Human FoxM1 is 763 amino acids (aa) in length. It contains three PEST sequences (aa 22-34, 460-472 and 478-515), a forkhead DNA binding domain (aa 235-327), a Pro/Ser/Thr-rich region (aa 480-513) and a phosphorylation site at Ser704. There are at least two splice variants. The first shows a 38 aa insertion after Val423. This blocks FoxM1 transcriptional activation activity. The second variant shows a deletion of aa 326-340. Over aa 642-763, human FoxM1 shares 73% aa identity with mouse FoxM1.


Key Product Details
Validated by
Biological Validation
Species Reactivity
Human
Applications
Western Blot, Immunocytochemistry
Label
Unconjugated
Antibody Source
Polyclonal Sheep IgG
Loading...
Product Specifications
Immunogen
E. coli-derived recombinant human FoxM1
Thr642-Gln763
Accession # Q08050
Thr642-Gln763
Accession # Q08050
Specificity
Detects human FoxM1 in direct ELISAs and Western blots. In direct ELISAs, less than 1% cross-reactivity with recombinant human (rh) FoxB2 and rhFoxJ1 is observed.
Clonality
Polyclonal
Host
Sheep
Isotype
IgG
Scientific Data Images for Human FoxM1 Antibody
FoxM1 in U2OS Human Cell Line.
FoxM1 was detected in immersion fixed U2OS human osteosarcoma cell line using 10 µg/mL Sheep Anti-Human FoxM1 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF3975) for 3 hours at room temperature. Cells were stained with the NorthernLights™ 557-conjugated Anti-Sheep IgG Secondary Antibody (red; Catalog # NL010) and counterstained (green). View our protocol for Fluorescent ICC Staining of Cells on Coverslips.Detection of Human FoxM1 by Western Blot.
Western blot shows lysates of U2OS human osteosarcoma cell line. PVDF membrane was probed with 1 µg/mL of Sheep Anti-Human FoxM1 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF3975) followed by HRP-conjugated Anti-Sheep IgG Secondary Antibody (Catalog # HAF016). A specific band was detected for FoxM1 at approximately 100 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 8.Detection of Human FoxM1 by Western Blot
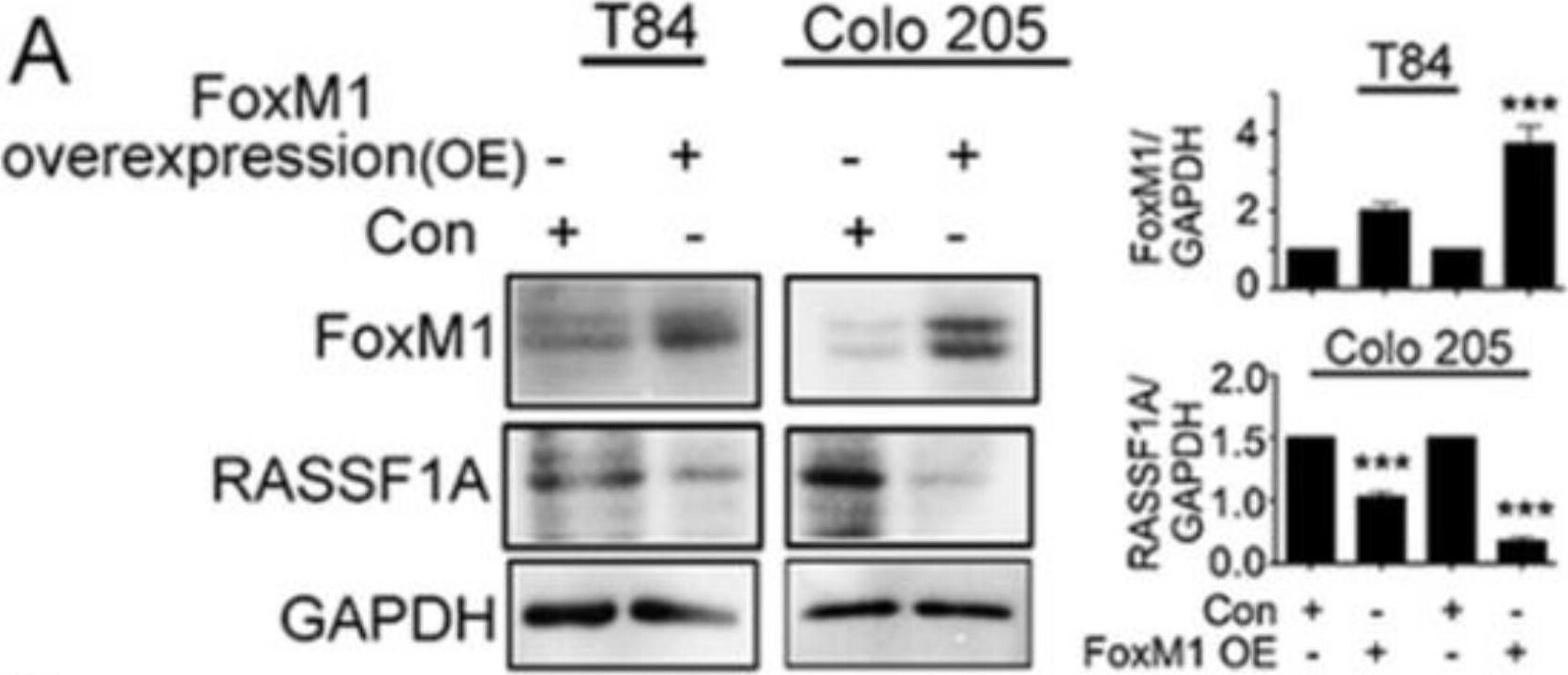
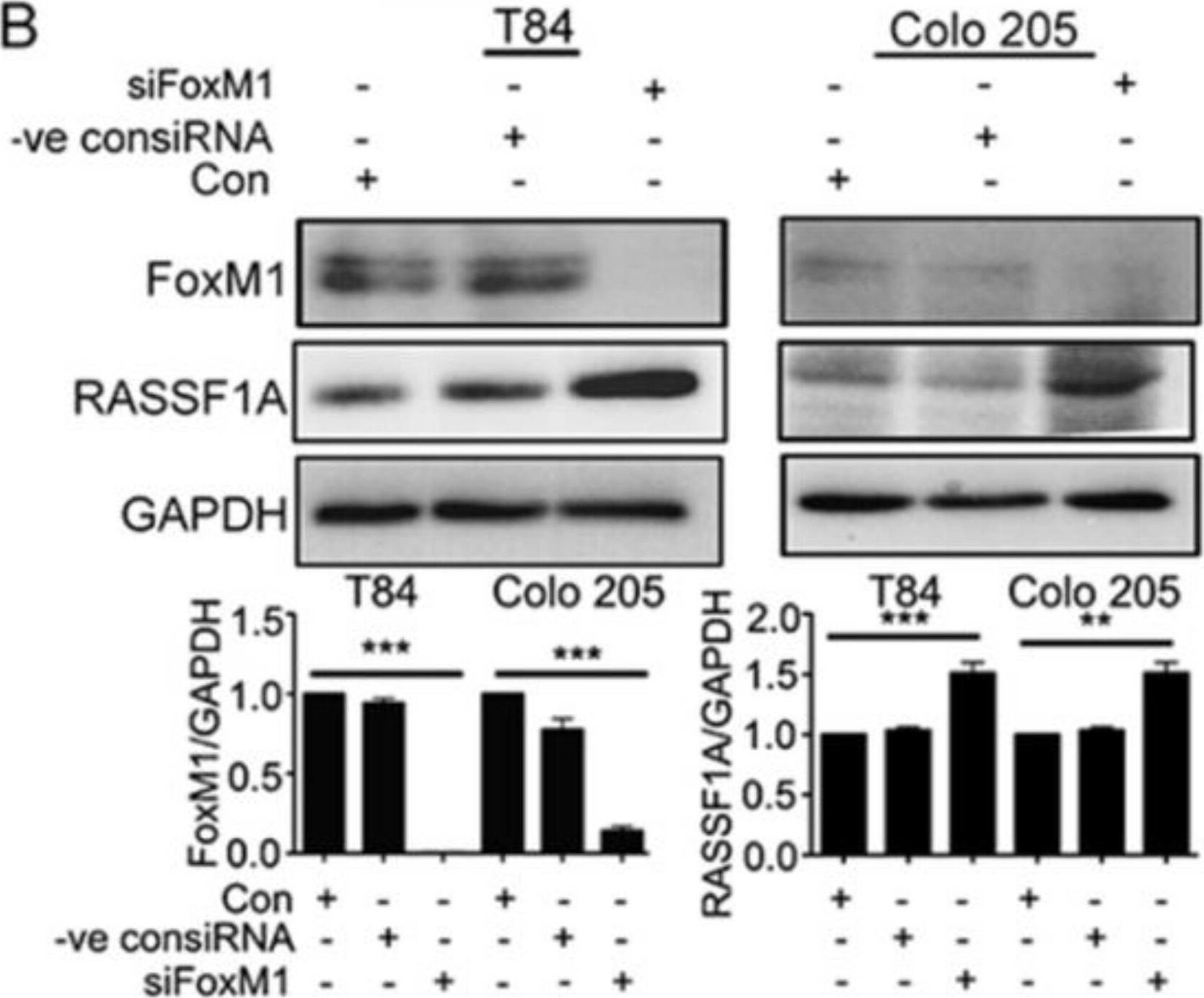
RASSF1A regulation is dependent on FoxM1. T84 and Colo 205 cells were transfected with plasmid for overexpression of FoxM1. (A) Cell lysates were analyzed by immunoblot and quantified by densitometry for expression of FoxM1, RASSF1A. Expression is normalized against GAPDH. The right panel of (A) represents the densitometric analysis of FoxM1 and RASSF1A. (B) T84 and Colo 205 cells were transfected with siRNA for FoxM1 or control siRNA for 48 h. Cell lysates were evaluated for FoxM1 ((B), 1st lane), RASSF1A ((B), 2nd lane) by immunoblot and quantified by densitometry. The results are from three independent experiments. (** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human FoxM1 by Western Blot
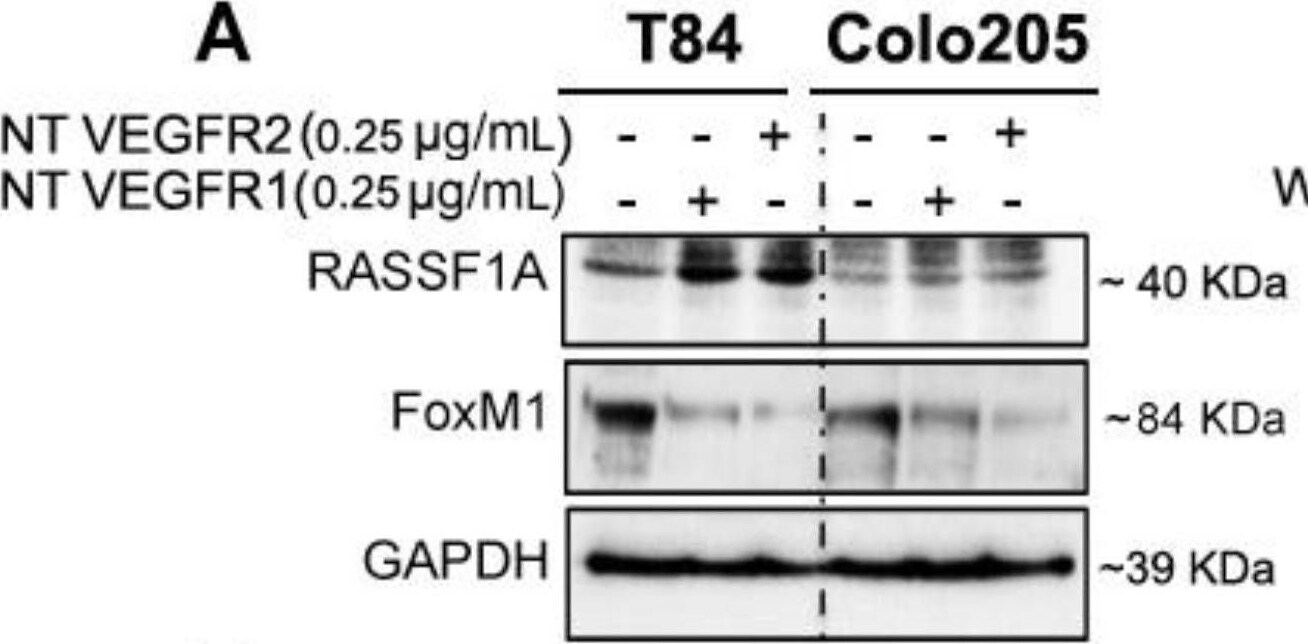
Neutralizing VEGF receptor antibodies and Akt inhibition upregulates RASSF1A and downregulates FoxM1 in mCRC. T84 and Colo 205 cells were treated with VEGF receptor 1 and VEGFR2 as indicated before (see Section 2.3). Protein expression from cell lysates was determined by immunoblotting for RASSF1A ((A), upper panel) and FoxM1 ((A), middle panel). Densitometry analysis was performed and normalyzed with GAPDH expression to demonstrate significant upregulation for RASSF1A (B) and downregulation of FoxM1 in the presence of neutralized (NT) VEGFR1 or VEGFR2 (C). (D,E) T84 and Colo 205 cells were treated with Akt inhibitor (wortmannin) with different doses (0–1 µM) for 24 h. Cell lysates were analyzed for pAkt (1st lane), total Akt (2nd lane), RASSF1A (3rd lane) expression by immunoblot analysis and quantified by densitometry (F,G). The results are from three independent experiments. (* p < 0.05, ** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human FoxM1 by Western Blot
RASSF1A regulation is dependent on FoxM1. T84 and Colo 205 cells were transfected with plasmid for overexpression of FoxM1. (A) Cell lysates were analyzed by immunoblot and quantified by densitometry for expression of FoxM1, RASSF1A. Expression is normalized against GAPDH. The right panel of (A) represents the densitometric analysis of FoxM1 and RASSF1A. (B) T84 and Colo 205 cells were transfected with siRNA for FoxM1 or control siRNA for 48 h. Cell lysates were evaluated for FoxM1 ((B), 1st lane), RASSF1A ((B), 2nd lane) by immunoblot and quantified by densitometry. The results are from three independent experiments. (** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human FoxM1 by Western Blot
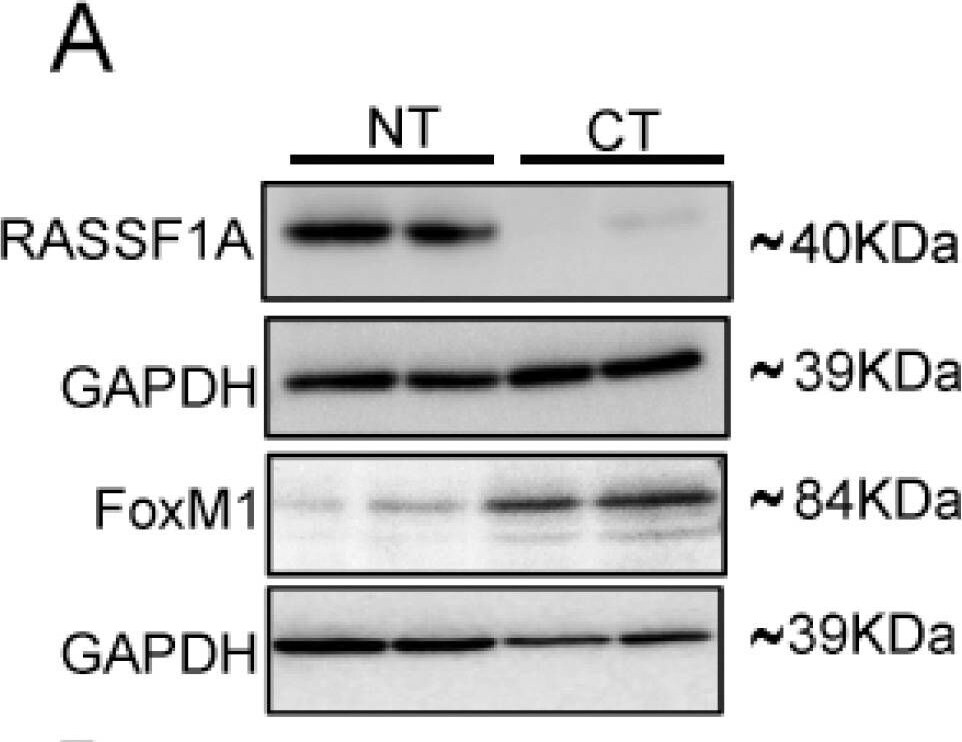
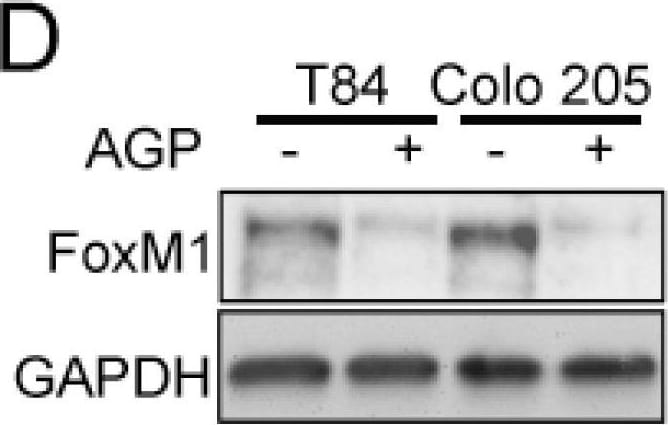
FoxM1 and RASSF1A expression is inversely co-related in colon cancer tissues. (A) Tissue extranction from both normal and colon cancer patient for monitoring the translational level of FoxM1 and RASSF1A by immunoblot. Quantification of RASSF1A (B) and FoxM1 (C) by scanning densitometry. GAPDH was used as a loading control. (D) T84 and Colo 205 cells were treated with or without AGP IC50 (45 µM) for 48 h. Cell lysates were analyzed by Western blot for FoxM1 and GAPDH expression. (E) Quantitative estimations of FoxM1 levels determined by densitometry measurements of western blots from three independent experiments after normalization with GAPDH (p < 0.001). T84 and Colo 205 cells were treated with or without AGP as indicated before and the transcriptional level were determined by qRT-PCR for (F) FoxM1 and (G) PTTG1. Bar graphs show quantitative results normalized to GAPDH mRNA levels. Results are from three independent experiments and statistical significance was determined using one way-ANOVA followed Bonferroni test. (* p < 0.05, ** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human FoxM1 by Western Blot
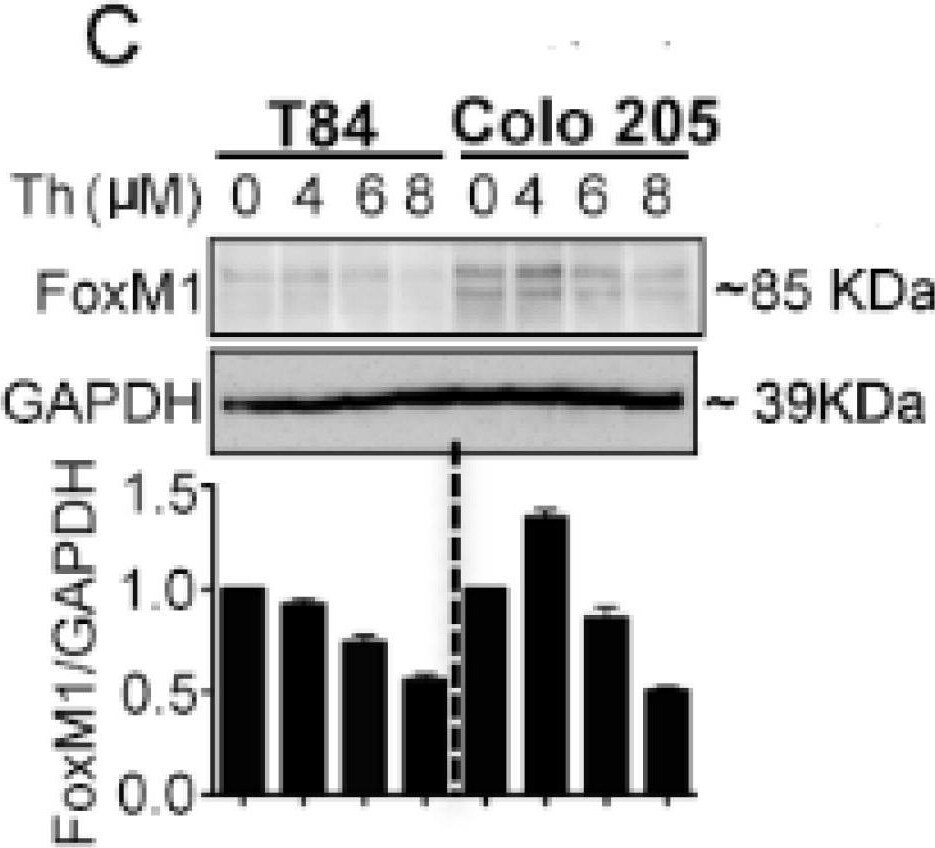
FoxM1 inhibitor induced RASSF1A and PTEN expression and YAP phosphorylation in mCRC cells. T84 and Colo 205 cells were treated with FoxM1 inhibitor, thiostrepton (Th) with different doses (0–8 µM) for 48 h. (A,B) mRNA was extracted after 24 h for detection of RASSF1A by qRT-PCR. Bar graphs show quantitative results normalized to GAPDH mRNA levels. Cell lysates were monitored by immunoblot for FoxM1(C), RASSF1A (D), p-YAP and total YAP (E), PTEN (F), and GAPDH as indicated. Immunoblots were quantified by scanning densitometry and normalized against GAPDH expression (lower panels of (C–E) for FoxM1, RASSF1A and p-YAP, respectively. Right panel of (F) represents the quantification of PTEN). (G) Patient derived organoids were treated with (8 µM) and without thiostrepton for 24 h and 48 h. Organoid cell lysates were analyzed by immunoblot for RASSF1A expression and quantified by densitometry ((G), lower panel). The results are from three independent experiments. (* p < 0.05, ** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human FoxM1 by Western Blot
FoxM1 and RASSF1A expression is inversely co-related in colon cancer tissues. (A) Tissue extranction from both normal and colon cancer patient for monitoring the translational level of FoxM1 and RASSF1A by immunoblot. Quantification of RASSF1A (B) and FoxM1 (C) by scanning densitometry. GAPDH was used as a loading control. (D) T84 and Colo 205 cells were treated with or without AGP IC50 (45 µM) for 48 h. Cell lysates were analyzed by Western blot for FoxM1 and GAPDH expression. (E) Quantitative estimations of FoxM1 levels determined by densitometry measurements of western blots from three independent experiments after normalization with GAPDH (p < 0.001). T84 and Colo 205 cells were treated with or without AGP as indicated before and the transcriptional level were determined by qRT-PCR for (F) FoxM1 and (G) PTTG1. Bar graphs show quantitative results normalized to GAPDH mRNA levels. Results are from three independent experiments and statistical significance was determined using one way-ANOVA followed Bonferroni test. (* p < 0.05, ** p < 0.01, *** p < 0.001). Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/30744076), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human FoxM1 Antibody
Application
Recommended Usage
Immunocytochemistry
5-15 µg/mL
Sample: Immersion fixed U2OS human osteosarcoma cell line
Sample: Immersion fixed U2OS human osteosarcoma cell line
Western Blot
1 µg/mL
Sample: U2OS human osteosarcoma cell line
Sample: U2OS human osteosarcoma cell line
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: FoxM1
Long Name
Forkhead Box M1
Alternate Names
FKHL16, HFH-11, HNF-3, INS-1, MPHOSPH2, MPP-2, PIG29, Trident
Gene Symbol
FOXM1
UniProt
Additional FoxM1 Products
Product Documents for Human FoxM1 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human FoxM1 Antibody
For research use only
Related Research Areas
Citations for Human FoxM1 Antibody
Customer Reviews for Human FoxM1 Antibody
There are currently no reviews for this product. Be the first to review Human FoxM1 Antibody and earn rewards!
Have you used Human FoxM1 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...