Human Laminin alpha 4 Antibody
R&D Systems | Catalog # AF7340


Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human
Applications
Validated:
Immunohistochemistry, Western Blot, Neutralization, Immunocytochemistry, Simple Western
Cited:
Western Blot, Neutralization, Immunocytochemistry, Bioassay
Label
Unconjugated
Antibody Source
Polyclonal Sheep IgG
Loading...
Product Specifications
Immunogen
Chinese hamster ovary cell line CHO-derived recombinant human Laminin alpha 4
Gln826-Ala1816, predicted
Accession # EAW48267
Gln826-Ala1816, predicted
Accession # EAW48267
Specificity
Detects human Laminin alpha 4 in direct ELISAs and Western blots. In direct ELISAs, approximately 15% cross-reactivity with recombinant mouse Laminin alpha 4 is observed.
Clonality
Polyclonal
Host
Sheep
Isotype
IgG
Endotoxin Level
<0.10 EU per 1 μg of the antibody by the LAL method.
Scientific Data Images for Human Laminin alpha 4 Antibody
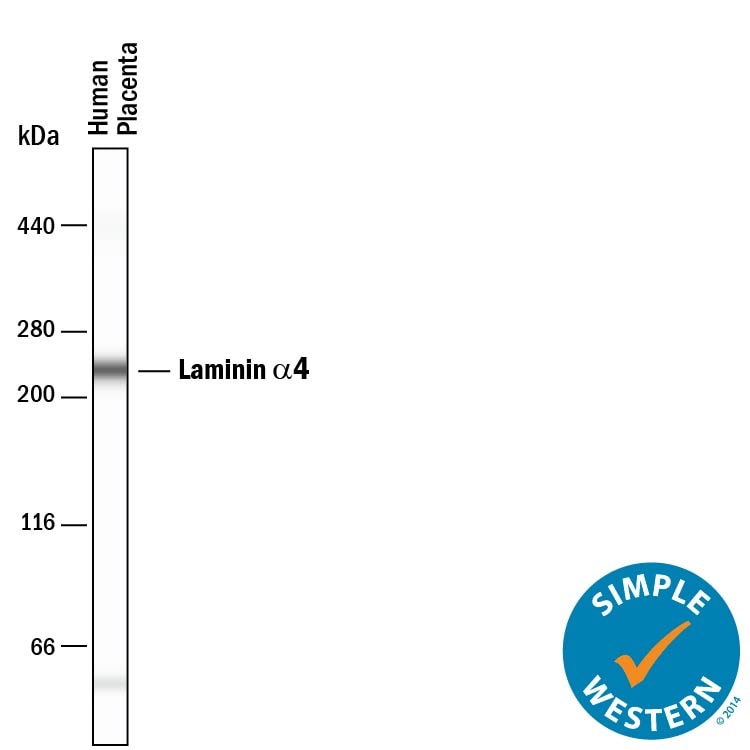
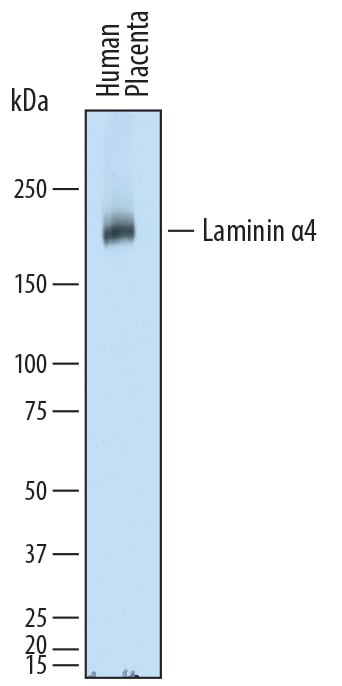
Detection of Human Laminin alpha 4 by Western Blot.
Western blot shows lysates of human placenta tissue. PVDF membrane was probed with 2 µg/mL of Sheep Anti-Human Laminin a4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF7340) followed by HRP-conjugated Anti-Sheep IgG Secondary Antibody (Catalog # HAF016). A specific band was detected for Laminin a4 at approximately 200-220 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.Laminin alpha 4 in T98G Human Cell Line.
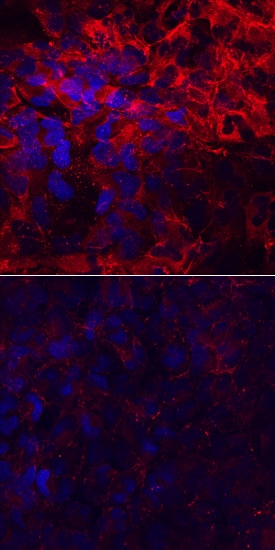
Laminin a4 was detected in immersion fixed T98G human glioblastoma cell line, induced (upper panel) or not (lower panel) with StemXVivo EMT Inducing Media Supplement (Catalog # CCM017), using Sheep Anti-Human Laminin a4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF7340) at 10 µg/mL for 3 hours at room temperature. Cells were stained using the NorthernLights™ 557-conjugated Anti-Sheep IgG Secondary Antibody (red; Catalog # NL010) and counterstained with DAPI (blue). Specific staining was localized to cytoplasm and cell surfaces. View our protocol for Fluorescent ICC Staining of Cells on Coverslips.Laminin alpha 4 in Human Placenta.
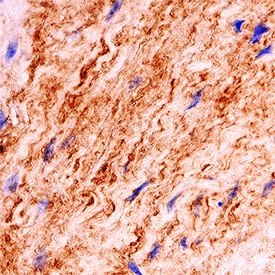
Laminin a4 was detected in immersion fixed paraffin-embedded sections of human placenta using Sheep Anti-Human Laminin a4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF7340) at 3 µg/mL for 1 hour at room temperature followed by incubation with the Anti-Sheep IgG VisUCyte™ HRP Polymer Antibody (Catalog # VC006). Before incubation with the primary antibody, tissue was subjected to heat-induced epitope retrieval using Antigen Retrieval Reagent-Basic (Catalog # CTS013). Tissue was stained using DAB (brown) and counterstained with hematoxylin (blue). Specific staining was localized to Golgi apparatus. View our protocol for IHC Staining with VisUCyte HRP Polymer Detection Reagents.Detection of Human Laminin alpha 4 by Simple WesternTM.
Simple Western lane view shows lysates of human placenta tissue, loaded at 0.2 mg/mL. A specific band was detected for Laminin a4 at approximately 233 kDa (as indicated) using 20 µg/mL of Sheep Anti-Human Laminin a4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF7340) followed by 1:50 dilution of HRP-conjugated Anti-Sheep IgG Secondary Antibody (Catalog # HAF016). This experiment was conducted under reducing conditions and using the 66-440 kDa separation system.Adhesion Induced by Laminin alpha 4 and Neutralization by Human Laminin alpha 4 Antibody.
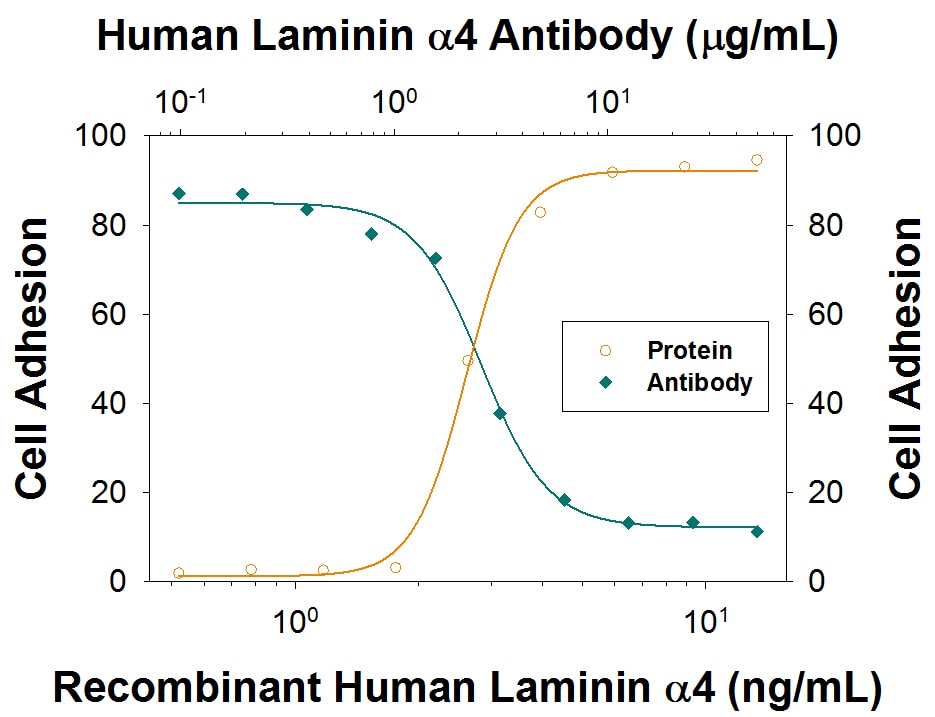
Recombinant Human Laminin a4 (Catalog # 7340-A4) supports adhesion in the HT1080 human fibrosarcoma cell line in a dose-dependent manner (orange line), as measured by Calcein AM. Adhesion elicited by Recombinant Human Laminin a4 (5 µg/mL) is neutralized (green line) by increasing concentrations of Sheep Anti-Human Laminin a4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF7340). The ND50 is typically 1.5-7.5 µg/mL.Detection of Human Laminin alpha 4 by Western Blot
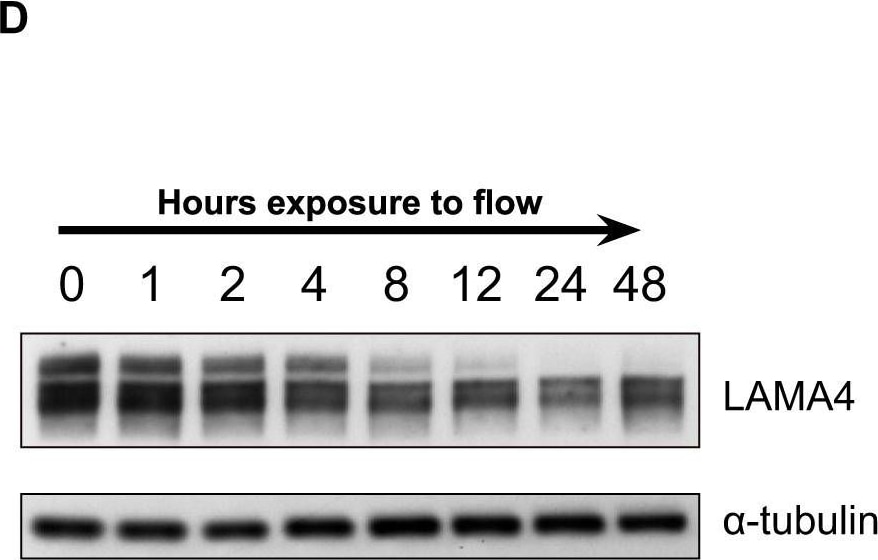
Flow induces remodeling of the endothelial extracellular matrix.A, Proteome and (B) cell surface proteome data of identified laminins. T or CT represents the enzyme used for protein digestion. The depicted numbers represent the biotin-modified lysine residue. C, Schematic representation of laminin 411 with the linker region between domains LG3 and LG4. The blue lysine residues were found to be decreased under flow conditions (as depicted in panel B). D, Immunoblot analysis of LAMA4 in ECs exposed to flow over time. alpha -tubulin was used as a loading control. E, Protein distribution of LAMA5 and LAMA4 after flow-exposure. Green represents LAMA4 or LAMA5, blue is HOECHST. The scale bar is 10 μm. Depicted are maximum intensity projections. F, Proteome and (G) cell surface proteome data of identified fibulins. H, Immunofluorescence analysis of EFEMP1 and FBLN2 abundance and distribution affected by flow. Red is VE-cadherin, blue is HOECHST. Depicted are maximum intensity projections. Micrographs are representative for 3 independent experiments. The scale bar is 10 μm. Image collected and cropped by CiteAb from the following publication (https://pubmed.ncbi.nlm.nih.gov/32332107), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
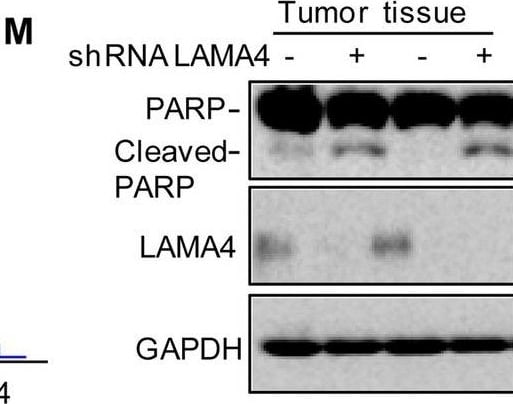
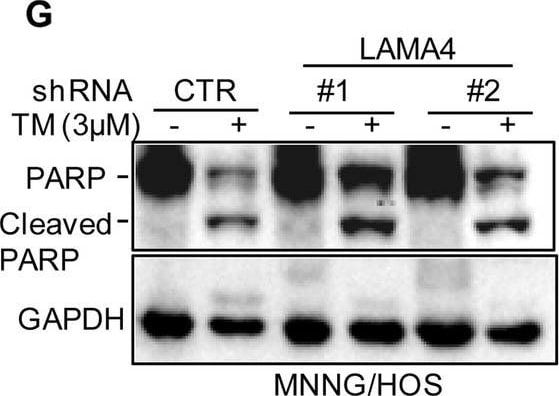
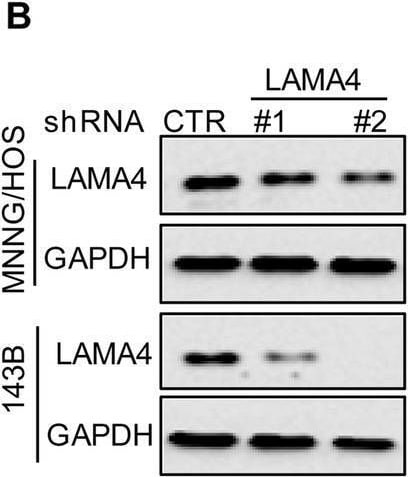
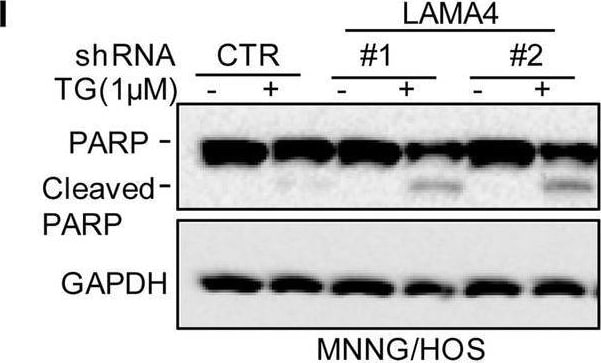
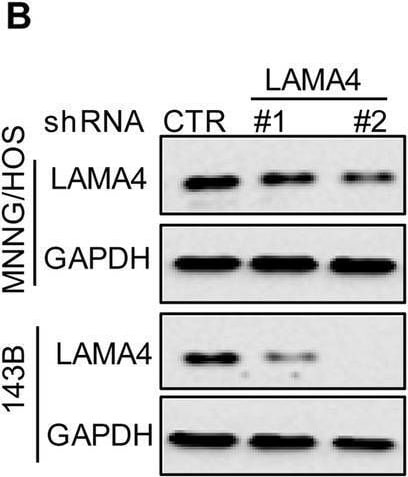
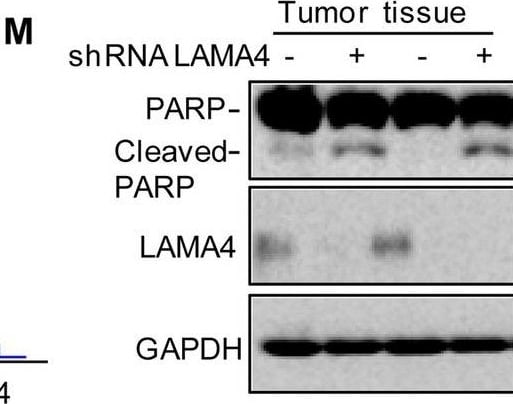
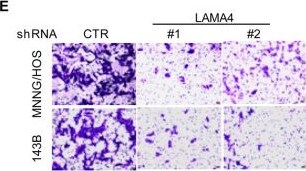
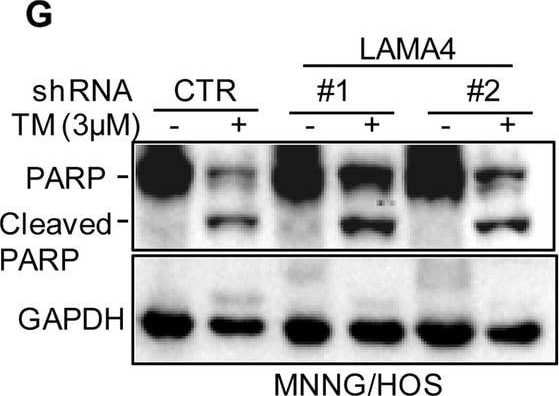
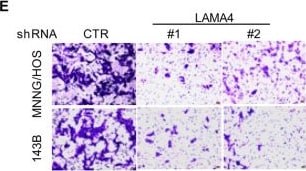
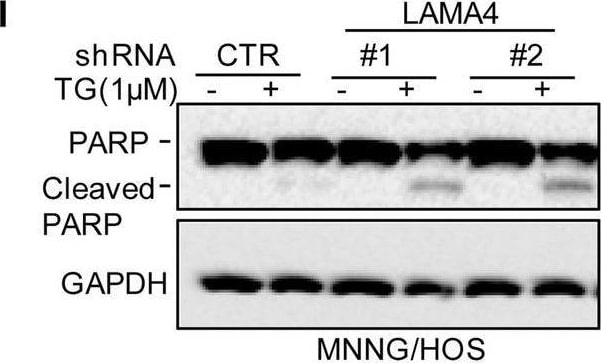
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
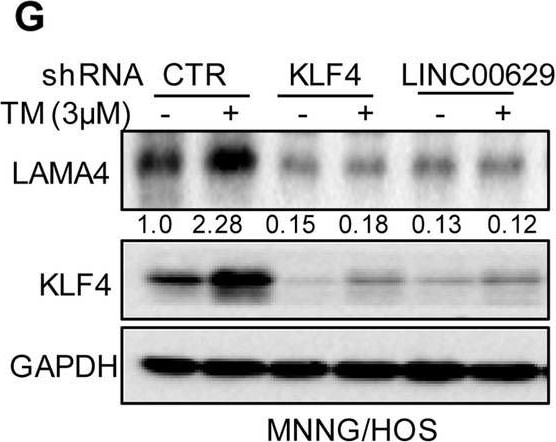
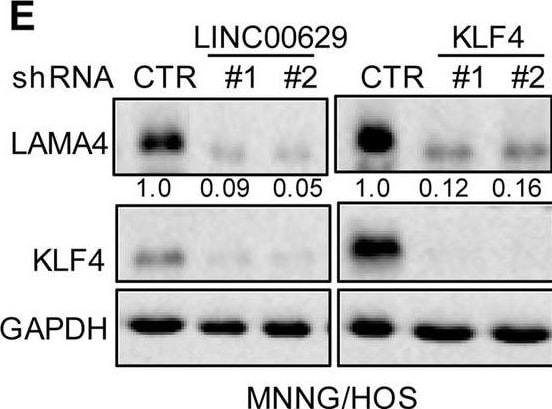
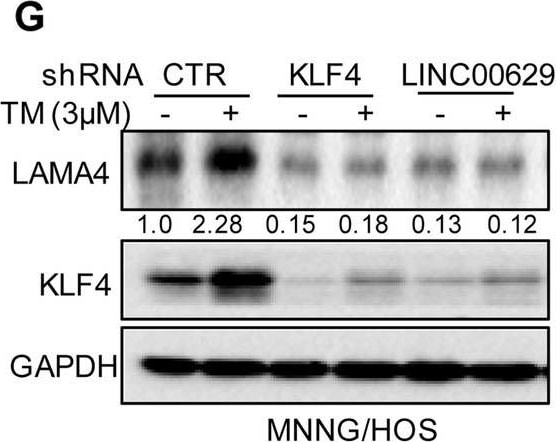
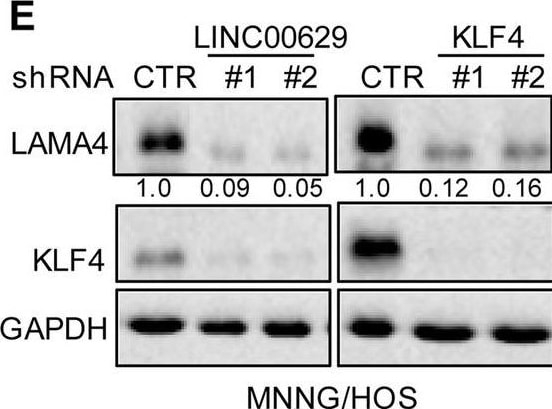
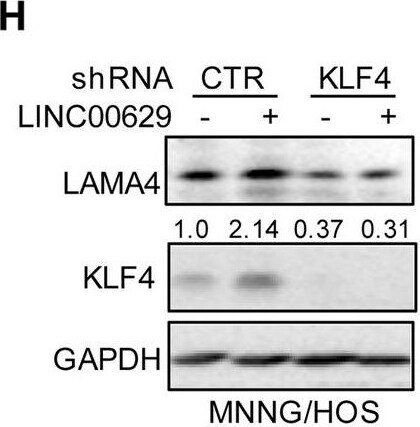
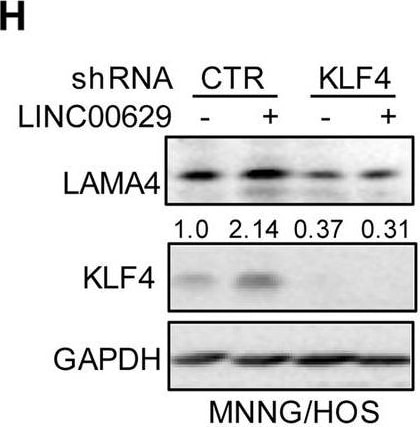
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Immunohistochemistry
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Immunohistochemistry
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
Knockdown of LAMA4 suppresses the malignant behaviour of osteosarcoma and accelerates ER stress-induced apoptosis. A The expression levels of LAMA4 in osteosarcoma (n = 88) and normal tissues (n = 564) were analysed from the TNMplot database. B LAMA4 was knocked down in MNNG/HOS and 143B cells. The expression of LAMA4 was detected by Western blot. GAPDH was used as the loading control. C-D The cells (3000 cells/well) with or without LAMA4 depletion were tested for cell growth in the colony formation assay. Viable colonies after 1 week were counted and are shown (C). Data are depicted as bar graphs (D). E-F The migration of the indicated cells was detected by Transwell assays. Representative images of crystal violet-stained culture plates are shown (E). Data are depicted as bar graphs (F). G-H MNNG/HOS cells with or without LAMA4 knockdown were treated with 3 μM TM for 36 h. Cell apoptosis and cell viability were analysed by Western blot (G) and CCK8 assays (H). GAPDH was used as the loading control. I-J MNNG/HOS cells with or without LAMA4 knockdown were treated with 1 μM TG for 36 h. Cell apoptosis and cell viability were analysed by Western blot (I) and CCK8 assays (J). GAPDH was used as the loading control. K-M MNNG/HOS cells (106 cells per mouse) with or without LAMA4 knockdown were injected subcutaneously into nude mice (n = 4). Representative images of xenograft tumours (K). The weights of the tumours were calculated and analysed (L). The expression levels of PARP, cleaved PARP and LAMA4 in tumours were measured by Western blot (M). Data in D, F, H, J, L were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Laminin alpha 4 by Western Blot
KLF4 upregulates LAMA4 expression. A-B Heatmap showing the altered genes identified by RNA sequencing analysis in LINC00629 (A)- or KLF4 (B)-depleted MNNG/HOS cells. C Volcano plots showing differentially expressed mRNAs for LINC00629 knockdown or KLF4 knockdown versus the controls (absolute log2-fold change > 1 and q value < 0.05). The green dot indicates the downregulated genes, and the red dot indicates the upregulated genes. D The mRNA levels of LAMA4 are shown from RNA sequencing data. E-F The protein and mRNA levels of LAMA4 were detected by Western blot (E) and qRT–PCR (F) in MNNG/HOS cells with or without LINC00629 or KLF4 knockdown. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. G MNNG/HOS cells with or without KLF4 or LINC00629 knockdown were treated with 3 μM TM for 36 h. The expression of LAMA4 was detected by Western blot. Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. H-I LINC00629 was overexpressed in MNNG/HOS cells with or without KLF4 knockdown. The protein and mRNA levels of LAMA4 were detected by Western blotting (H) and qRT–PCR (I). Numbers represent the relative intensities of western blot bands of LAMA4 to GAPDH. Data in F and M were analysed by Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36539799), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human Laminin alpha 4 Antibody
Application
Recommended Usage
Immunocytochemistry
5-15 µg/mL
Sample: Immersion fixed T98G human glioblastoma cell line induced with EMT
Sample: Immersion fixed T98G human glioblastoma cell line induced with EMT
Immunohistochemistry
3-25 µg/mL
Sample: Immersion fixed paraffin-embedded sections of human placenta
Sample: Immersion fixed paraffin-embedded sections of human placenta
Simple Western
20 µg/mL
Sample: Human placenta tissue
Sample: Human placenta tissue
Western Blot
2 µg/mL
Sample: Human placenta tissue
Sample: Human placenta tissue
Neutralization
Measured by its ability to neutralize Laminin alpha 4-induced adhesion in the HT1080 human fibrosarcoma cell line. The Neutralization Dose (ND50) is typically 1.5-7.5 µg/mL in the presence of 5 µg/mL Recombinant Human Laminin alpha 4.
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Sterile PBS to a final concentration of 0.2 mg/mL. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: Laminin alpha 4
Alternate Names
LAMA4
Gene Symbol
LAMA4
UniProt
Additional Laminin alpha 4 Products
Product Documents for Human Laminin alpha 4 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human Laminin alpha 4 Antibody
For research use only
Related Research Areas
Citations for Human Laminin alpha 4 Antibody
Customer Reviews for Human Laminin alpha 4 Antibody
There are currently no reviews for this product. Be the first to review Human Laminin alpha 4 Antibody and earn rewards!
Have you used Human Laminin alpha 4 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...
Associated Pathways
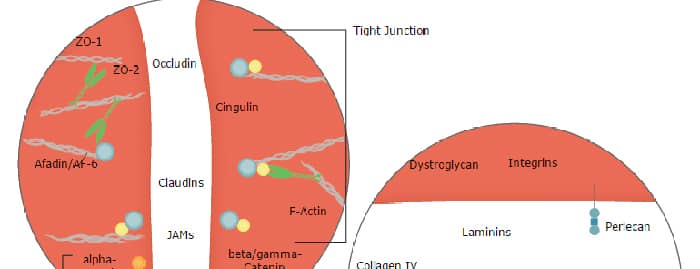
 Blood-Brain Barrier Pathway: Anatomy
Blood-Brain Barrier Pathway: Anatomy