Spi-1 (SFFV proviral integration 1 protein; also PU.1 and Sfpi1) is a 37-41 kDa member of the ets family of transcription factors. It is both an RNA and DNA binding protein that is found in hematopoietic cells such as B cells, neutrophils, macrophages and dendritic cells (DC). Spi-1 can act as both a transcriptional activator and repressor. In DC, Spi-1 promotes the expression of CD80, CD86 and CD11b, while in proerythroblasts it blocks GATA-1 induced transcriptional activation. Spi-1 binds to DNA as a monomer and interacts with multiple factors such as Runx-1, IRF8, GATA-1 and c-Jun. Mouse Spi-1 is 272 amino acids (aa) in length. It contains a transactivation region (aa 7-117), a PEST domain (aa 118-164), and a DNA-binding domain (aa 165-266). Phosphorylation on Ser41, 142 and 148 increases activity. There is one alternative start site at Met7. Over aa 1-169, mouse Spi-1 shares 88% and 81% aa identity with rat and human Spi-1, respectively.
Mouse PU.1/Spi‑1 Antibody
R&D Systems | Catalog # MAB7124


Key Product Details
Species Reactivity
Validated:
Mouse
Cited:
Mouse
Applications
Validated:
Western Blot, Immunocytochemistry
Cited:
Immunohistochemistry
Label
Unconjugated
Antibody Source
Monoclonal Rat IgG2A Clone # 823123
Loading...
Product Specifications
Immunogen
E. coli-derived recombinant mouse Spi-1
Met1-Lys169
Accession # P17433
Met1-Lys169
Accession # P17433
Specificity
Detects mouse PU.1/Spi-1 in ELISAs. In direct ELISAs, no cross-reactivity with recombinant human PU.1/Spi-1 is observed. Detects mouse and rat PU.1/Spi-1 in Western Blots
Clonality
Monoclonal
Host
Rat
Isotype
IgG2A
Scientific Data Images for Mouse PU.1/Spi‑1 Antibody
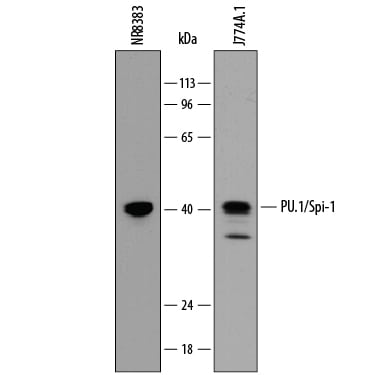
Detection of Mouse and Rat PU.1/Spi‑1 by Western Blot.
Western blot shows lysates of NR8383 rat alveolar macrophage cell line and J774A.1 mouse reticulum cell sarcoma macrophage cell line. PVDF membrane was probed with 0.1 µg/mL of Rat Anti-Mouse PU.1/Spi-1 Monoclonal Antibody (Catalog # MAB7124) followed by HRP-conjugated Anti-Rat IgG Secondary Antibody (Catalog # HAF005). A specific band was detected for PU.1/Spi-1 at approximately 40 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.PU.1/Spi‑1 in J774A.1 Mouse Cell Line.
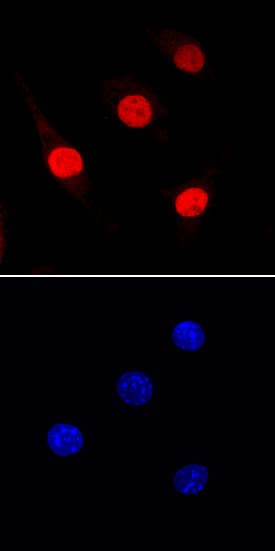
PU.1/Spi-1 was detected in immersion fixed J774A.1 mouse reticulum cell sarcoma macrophage cell line using Rat Anti-Mouse PU.1/Spi-1 Monoclonal Antibody (Catalog # MAB7124) at 10 µg/mL for 3 hours at room temperature. Cells were stained using the NorthernLights™ 557-conjugated Anti-Rat IgG Secondary Antibody (red, upper panel; Catalog # NL013) and counterstained with DAPI (blue, lower panel). Specific staining was localized to nuclei. View our protocol for Fluorescent ICC Staining of Non-adherent Cells.Applications for Mouse PU.1/Spi‑1 Antibody
Application
Recommended Usage
Immunocytochemistry
8-25 µg/mL
Sample: Immersion fixed J774A.1 mouse reticulum cell sarcoma macrophage cell line
Sample: Immersion fixed J774A.1 mouse reticulum cell sarcoma macrophage cell line
Western Blot
0.1 µg/mL
Sample: NR8383 rat alveolar macrophage cell line and J774A.1 mouse reticulum cell sarcoma macrophage cell line
Sample: NR8383 rat alveolar macrophage cell line and J774A.1 mouse reticulum cell sarcoma macrophage cell line
Formulation, Preparation, and Storage
Purification
Protein A or G purified from hybridoma culture supernatant
Reconstitution
Sterile PBS to a final concentration of 0.5 mg/mL. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: PU.1/Spi-1
Long Name
Hematopoietic transcription factor PU.1
Alternate Names
OF, SFPI1, Spi-1, SPI-A, SPI1
Gene Symbol
SPI1
UniProt
Additional PU.1/Spi-1 Products
Product Documents for Mouse PU.1/Spi‑1 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Mouse PU.1/Spi‑1 Antibody
For research use only
Citations for Mouse PU.1/Spi‑1 Antibody
Customer Reviews for Mouse PU.1/Spi‑1 Antibody
There are currently no reviews for this product. Be the first to review Mouse PU.1/Spi‑1 Antibody and earn rewards!
Have you used Mouse PU.1/Spi‑1 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars