Pappalysins belong to a fifth family of metzincins that consists of ADAMs/ADAMTSs, MMPs, astacins and serrylysins (1). PAPP-A is an important pregnancy protein and increases in plasma by a factor of about 150 during pregnancy as compared to the nonpregnant state. PAPP-A is also a major marker of Down syndrome in the first trimester of pregnancy because maternal serum levels of PAPP-A are significantly reduced when a fetus affected by Down syndrome is present (2). PAPP-A cleaves Insulin-like Growth Factor-Binding Protein-4 and -5 (IGFBP-4 and -5) at a single site, resulting in the release of bioactive IGF (3). Lack of IGFBP-4 cleavage in embryonic fibroblasts derived from PAPP-A knockout mice indicates that PAPP-A functions as a physiological IGFBP-4 protease (4). Three Lin12-Notch repeats (LNR) in the PAPP-A protein bind Ca2+ and are required for the cleavage of IGFBP-4, not IGFBP-5, by PAPP-A (5). The C-terminal LNR (residues 1476 to 1503) is not present in rhPAPP-A (residues 82 to 1214), which starts at the N-terminus of the mature chain and ends before the five Sushi (SCR) modules. As an active protease, rhPAPP-A cleaves IGFBP-5, which can be inhibited by 1,10-phenanthroline.
Human Pappalysin‑1/PAPP‑A Antibody
R&D Systems | Catalog # AF2487


Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human
Applications
Validated:
Western Blot, Immunoprecipitation
Cited:
Immunohistochemistry, Western Blot
Label
Unconjugated
Antibody Source
Polyclonal Goat IgG
Loading...
Product Specifications
Immunogen
Mouse myeloma cell line NS0-derived recombinant human Pappalysin‑1/PAPP‑A
Glu82-Asp1214
Accession # Q13219
Glu82-Asp1214
Accession # Q13219
Specificity
Detects human Pappalysin‑1/PAPP‑A in direct ELISAs and Western blots. In direct ELISAs, less than 5% cross-reactivity with recombinant human PAPP-A2 is observed.
Clonality
Polyclonal
Host
Goat
Isotype
IgG
Scientific Data Images for Human Pappalysin‑1/PAPP‑A Antibody
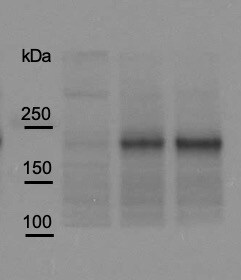
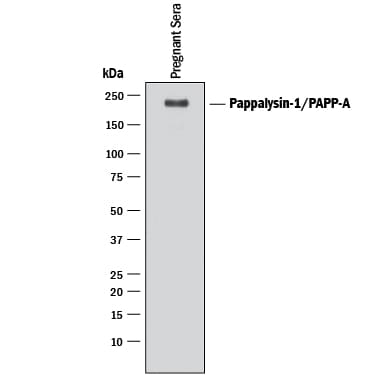
Detection of Human Pappalysin‑1/ PAPP‑A by Western Blot.
Western blot shows lysate of human pregnant sera. PVDF membrane was probed with 0.1 µg/mL of Goat Anti-Human Pappalysin-1/PAPP-A Antigen Affinity-purified Polyclonal Antibody (Catalog # AF2487) followed by HRP-conjugated Anti-Goat IgG Secondary Antibody (Catalog # HAF017). A specific band was detected for Pappalysin-1/PAPP-A at approximately 200 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.Detection of Human Pappalysin-1/PAPP-A by Western Blot
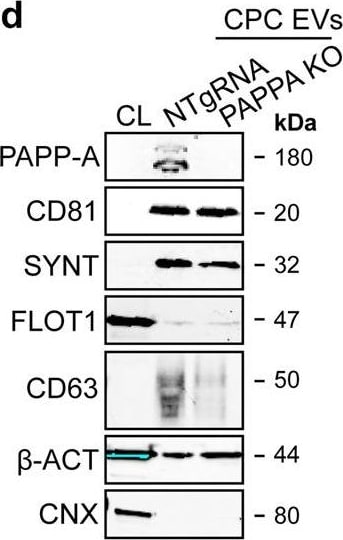
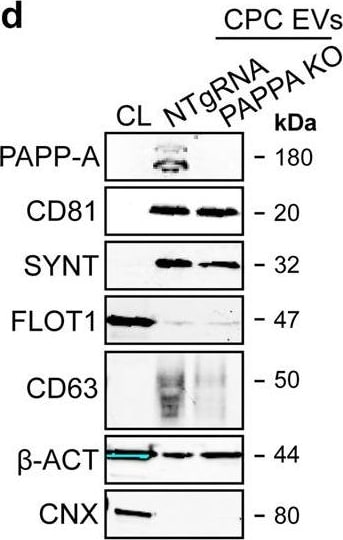
PAPPA KO-EVs were generated using CRISPR/Cas9.a Schematic depicting hypothesized mechanism of (intra)cellular signalling activated by EV-associated PAPP-A, based on identified proteins and significantly altered phosphosites measured in HMEC-1 upon veh-EV stimulation by (phopho)proteomic analysis. Detected proteins in HMEC-1 are displayed in grey, while significantly changing phosphosites present in cluster C1 (see Fig. 3d) are displayed in brown. b Representative western blot analysis of phosphorylated AKT (pAKT), total AKT (tAKT), phosphorylated ERK1/2 (pERK1/2) and total ERK1/2 (tERK1/2) in HMEC-1 treated with 6 × 1010 or 2 × 1010 CPC-EVs, or with 200 ng/mL free IGF-1 after pre-incubation with different doses of picropodophyllin (PPP). beta -actin ( beta -ACT) was included as housekeeping protein (I = phosphorylated protein blot, II = total protein blot). Biological replicates of (b) are displayed in Supplementary Fig. 9g, h. c Sanger sequencing results confirming 1 bp insertion in exon 3 of PAPPA at the CRISPR/Cas9 target site of the PAPPA KO-CPC clone, compared with the NTgRNA polyclonal CPC line. d Western blot analysis showing the absence of PAPP-A in PAPPA KO-EVs, compared with NTgRNA-EVs; the presence of CD81, CD63, Syntenin-1 (SYNT), Flotillin (FLOT1), beta -ACT, and absence of Calnexin (CNX) in both EV populations. FLOT1, beta -ACT and CNX were present in CPC lysate (CL). e Representative NTA plot showing the size distribution and particle concentration of PAPPA KO- and NTgRNA-CPC-EVs. f Protein content per 1 × 1010 PAPPA KO- and NTgRNA-EVs of two representative experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human Pappalysin-1/PAPP-A by Western Blot
PAPPA KO-EVs were generated using CRISPR/Cas9.a Schematic depicting hypothesized mechanism of (intra)cellular signalling activated by EV-associated PAPP-A, based on identified proteins and significantly altered phosphosites measured in HMEC-1 upon veh-EV stimulation by (phopho)proteomic analysis. Detected proteins in HMEC-1 are displayed in grey, while significantly changing phosphosites present in cluster C1 (see Fig. 3d) are displayed in brown. b Representative western blot analysis of phosphorylated AKT (pAKT), total AKT (tAKT), phosphorylated ERK1/2 (pERK1/2) and total ERK1/2 (tERK1/2) in HMEC-1 treated with 6 × 1010 or 2 × 1010 CPC-EVs, or with 200 ng/mL free IGF-1 after pre-incubation with different doses of picropodophyllin (PPP). beta -actin ( beta -ACT) was included as housekeeping protein (I = phosphorylated protein blot, II = total protein blot). Biological replicates of (b) are displayed in Supplementary Fig. 9g, h. c Sanger sequencing results confirming 1 bp insertion in exon 3 of PAPPA at the CRISPR/Cas9 target site of the PAPPA KO-CPC clone, compared with the NTgRNA polyclonal CPC line. d Western blot analysis showing the absence of PAPP-A in PAPPA KO-EVs, compared with NTgRNA-EVs; the presence of CD81, CD63, Syntenin-1 (SYNT), Flotillin (FLOT1), beta -ACT, and absence of Calnexin (CNX) in both EV populations. FLOT1, beta -ACT and CNX were present in CPC lysate (CL). e Representative NTA plot showing the size distribution and particle concentration of PAPPA KO- and NTgRNA-CPC-EVs. f Protein content per 1 × 1010 PAPPA KO- and NTgRNA-EVs of two representative experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human Pappalysin-1/PAPP-A by Western Blot
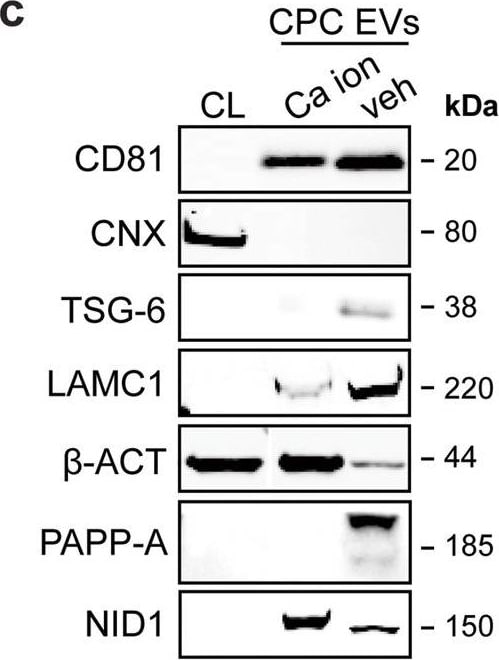
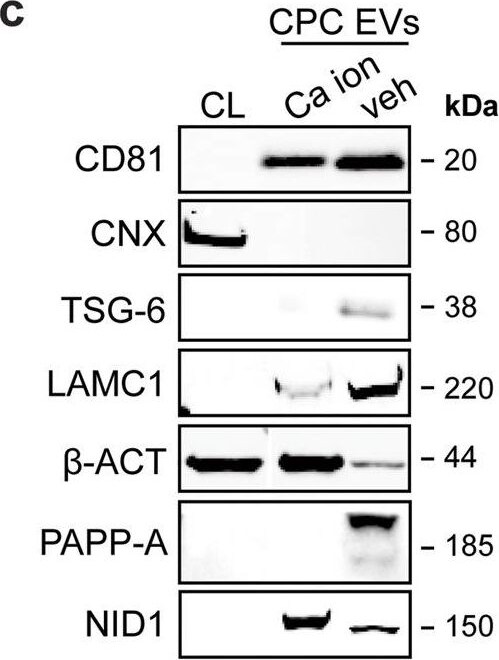
LC–MS/MS identified enriched proteins in veh-EVs compared with Ca ion- and SKOV-3-EVs.a Heat map of protein abundance (log2) of proteins identified in each biological replicate (veh-, Ca ion- and SKOV-3-EVs), as identified by LC–MS/MS. b Volcano plots showing average fold changes for protein abundance (log2) of proteins identified in veh-EVs compared to (i) Ca ion-EVs and (ii) SKOV-3-EVs. P-values were calculated using student’s T-test, and significantly changing proteins (p-value ≤ 0.05 and fold change >2) in veh-EVs are highlighted in red, while significantly changing proteins in Ca ion- and SKOV-3-EVs are highlighted in blue. c Western blot analysis confirming the enrichment of MS-identified proteins NID1, TSG-6, LAMC1, PAPP-A, CD81 and beta -actin ( beta -ACT) in veh-EVs compared with Ca ion-EVs. CNX was solely present in CPC lysate (CL). Complete blots of beta -ACT, PAPP-A and NID1 are displayed in Supplementary Fig. 10. d Venn diagram showing number of proteins with >2-fold significant enrichment (p ≤ 0.05) in veh-EVs compared to Ca ion- and SKOV-3-EVs, and overlap between those two populations. e Gene ontology analysis using PANTHER of enriched biological processes for the 105 overlapping proteins, depicting number of identified proteins in each group, ranked on smallest corrected p-value (−log10(FDR)). Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Pappalysin-1/PAPP-A by Western Blot
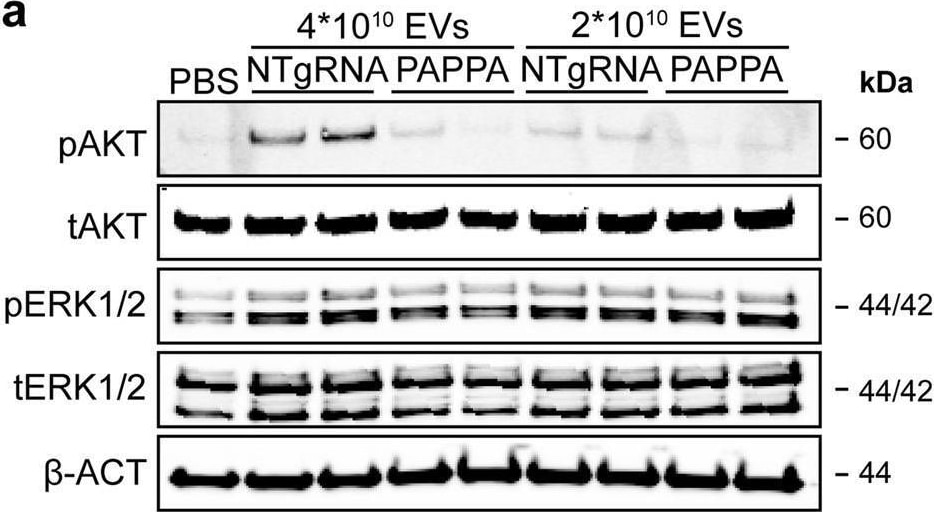
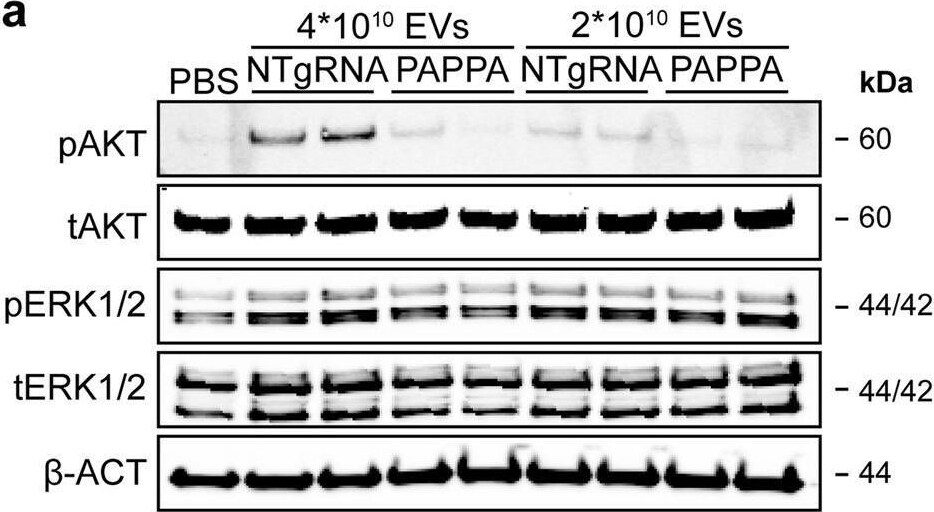
PAPPA KO-EVs showed reduced activation of intracellular signalling and activation of HMEC-1 migration and sprout formation.a–d Representative western blot analyses of pAKT, tAKT, pERK1/2 and tERK1/2 in HMEC-1 treated with PAPPA KO- and NTgRNA-EVs normalized on two doses of a, b total particle numbers or c, d total protein content. beta -ACT was included as housekeeping protein. b, d Quantification of pAKT, tAKT, pERK1/2 and tERK1/2 expression levels using densitometry expressed as pAKT/AKT and pERK/ERK ratios (n = 3). Biological replicates of (a, c) are also displayed in Supplementary Fig. 9i–k. e Wound healing assay showing effects of 1 µg and 2 × 1010 NTgRNA- and PAPPA KO-EVs on HMEC-1 migration, analysed both as % wound closure and absolute migration distance (n = 3, technical replicates. Data are representative of three biologically independent experiments). f, g Sprouting assay showing NTgRNA- and PAPPA KO-EV-induced HMEC-1 sprout formation on beads, analysed both as (g) mean length per sprout and total sprout length per bead (n = 3, technical replicates. Data are representative of two biologically independent experiments). Data are presented as mean ± SD. *p < 0.033, **p < 0.0021, ***p < 0.0002. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human Pappalysin-1/PAPP-A by Western Blot
LC–MS/MS identified enriched proteins in veh-EVs compared with Ca ion- and SKOV-3-EVs.a Heat map of protein abundance (log2) of proteins identified in each biological replicate (veh-, Ca ion- and SKOV-3-EVs), as identified by LC–MS/MS. b Volcano plots showing average fold changes for protein abundance (log2) of proteins identified in veh-EVs compared to (i) Ca ion-EVs and (ii) SKOV-3-EVs. P-values were calculated using student’s T-test, and significantly changing proteins (p-value ≤ 0.05 and fold change >2) in veh-EVs are highlighted in red, while significantly changing proteins in Ca ion- and SKOV-3-EVs are highlighted in blue. c Western blot analysis confirming the enrichment of MS-identified proteins NID1, TSG-6, LAMC1, PAPP-A, CD81 and beta -actin ( beta -ACT) in veh-EVs compared with Ca ion-EVs. CNX was solely present in CPC lysate (CL). Complete blots of beta -ACT, PAPP-A and NID1 are displayed in Supplementary Fig. 10. d Venn diagram showing number of proteins with >2-fold significant enrichment (p ≤ 0.05) in veh-EVs compared to Ca ion- and SKOV-3-EVs, and overlap between those two populations. e Gene ontology analysis using PANTHER of enriched biological processes for the 105 overlapping proteins, depicting number of identified proteins in each group, ranked on smallest corrected p-value (−log10(FDR)). Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Pappalysin-1/PAPP-A by Western Blot
PAPPA KO-EVs showed reduced activation of intracellular signalling and activation of HMEC-1 migration and sprout formation.a–d Representative western blot analyses of pAKT, tAKT, pERK1/2 and tERK1/2 in HMEC-1 treated with PAPPA KO- and NTgRNA-EVs normalized on two doses of a, b total particle numbers or c, d total protein content. beta -ACT was included as housekeeping protein. b, d Quantification of pAKT, tAKT, pERK1/2 and tERK1/2 expression levels using densitometry expressed as pAKT/AKT and pERK/ERK ratios (n = 3). Biological replicates of (a, c) are also displayed in Supplementary Fig. 9i–k. e Wound healing assay showing effects of 1 µg and 2 × 1010 NTgRNA- and PAPPA KO-EVs on HMEC-1 migration, analysed both as % wound closure and absolute migration distance (n = 3, technical replicates. Data are representative of three biologically independent experiments). f, g Sprouting assay showing NTgRNA- and PAPPA KO-EV-induced HMEC-1 sprout formation on beads, analysed both as (g) mean length per sprout and total sprout length per bead (n = 3, technical replicates. Data are representative of two biologically independent experiments). Data are presented as mean ± SD. *p < 0.033, **p < 0.0021, ***p < 0.0002. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37528162), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human Pappalysin‑1/PAPP‑A Antibody
Application
Recommended Usage
Immunoprecipitation
25 µg/mL
Sample: Conditioned cell culture medium spiked with Recombinant Human Pappalysin‑1/PAPP-A (Catalog # 2487-ZN), see our available Western blot detection antibodies
Sample: Conditioned cell culture medium spiked with Recombinant Human Pappalysin‑1/PAPP-A (Catalog # 2487-ZN), see our available Western blot detection antibodies
Western Blot
0.1 µg/mL
Sample: Human pregnant sera
Sample: Human pregnant sera
Reviewed Applications
Read 1 review rated 5 using AF2487 in the following applications:
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: Pappalysin-1/PAPP-A
References
- Boldt, H.B. et al. (2001) Biochem. J. 358:359.
- Fialova L. and I.M. Malbohan (2002) Bratisl. Lek. Listy 103:194.
- Laursen, L.S. et al. (2001) FEBS Lett. 504:36.
- Conover, C.A. et al. (2004) Development 131:1187.
- Boldt, H.B. et al. (2004) J. Biol. Chem. 279:38525.
Long Name
Pregnancy-associated Plasma Protein 1
Alternate Names
ASBABP2, DIPLA1, IGFBP-4ase, PAPA, PAPP-A1, PAPPA, Pappalysin1
Gene Symbol
PAPPA
UniProt
Additional Pappalysin-1/PAPP-A Products
Product Documents for Human Pappalysin‑1/PAPP‑A Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human Pappalysin‑1/PAPP‑A Antibody
For research use only
Related Research Areas
Citations for Human Pappalysin‑1/PAPP‑A Antibody
Customer Reviews for Human Pappalysin‑1/PAPP‑A Antibody (1)
5 out of 5
1 Customer Rating
Have you used Human Pappalysin‑1/PAPP‑A Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
1 of
1 review
Showing All
Filter By:
-
Application: Western BlotSample Tested: Cell LysatesSpecies: HumanVerified Customer | Posted 08/04/2020
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Cellular Response to Hypoxia Protocols
- Immunoprecipitation Protocol
- R&D Systems Quality Control Western Blot Protocol
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...