Human IFITM1 Antibody
R&D Systems | Catalog # AF4827


Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human, Mouse
Applications
Validated:
Western Blot, Immunocytochemistry
Cited:
Western Blot, Flow Cytometry, Immunocytochemistry
Label
Unconjugated
Antibody Source
Polyclonal Goat IgG
Loading...
Product Specifications
Immunogen
E. coli-derived recombinant human IFITM1
Met1-His36
Accession # NP_003632
Met1-His36
Accession # NP_003632
Specificity
Detects human IFITM1 in direct ELISAs and Western blots. In direct ELISAs and Western blots, approximately 40% cross-reactivity with recombinant human (rh) IFITM2 and rhIFITM3 is observed.
Clonality
Polyclonal
Host
Goat
Isotype
IgG
Scientific Data Images for Human IFITM1 Antibody
Detection of Human IFITM1 by Western Blot.
Western blot shows lysates of Jurkat human acute T cell leukemia cell line, TF-1 human erythroleukemic cell line, human placenta tissue. PVDF membrane was probed with 1 µg/mL of Goat Anti-Human IFITM1 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF4827) followed by HRP-conjugated Anti-Goat IgG Secondary Antibody (Catalog # HAF019). A specific band was detected for IFITM1 at approximately 17 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 8.IFITM1 in K562 Human Cell Line.
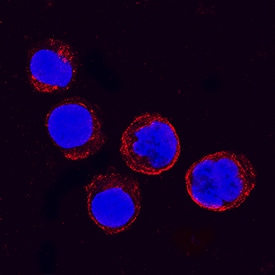
IFITM1 was detected in immersion fixed K562 human chronic myelogenous leukemia cell line using Goat Anti-Human IFITM1 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF4827) at 1.7 µg/mL for 3 hours at room temperature. Cells were stained using the NorthernLights™ 557-conjugated Anti-Goat IgG Secondary Antibody (red; Catalog # NL001) and counterstained with DAPI (blue). Specific staining was localized to plasma membranes and cytoplasm. View our protocol for Fluorescent ICC Staining of Non-adherent Cells.Detection of IFITM1 by Western Blot
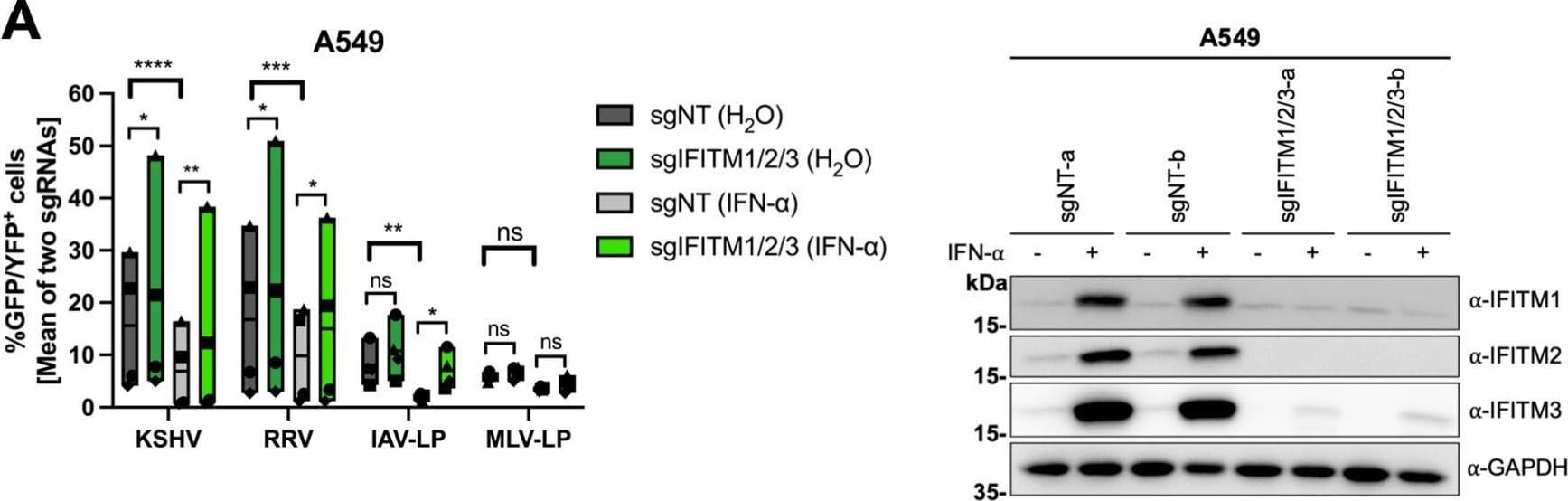
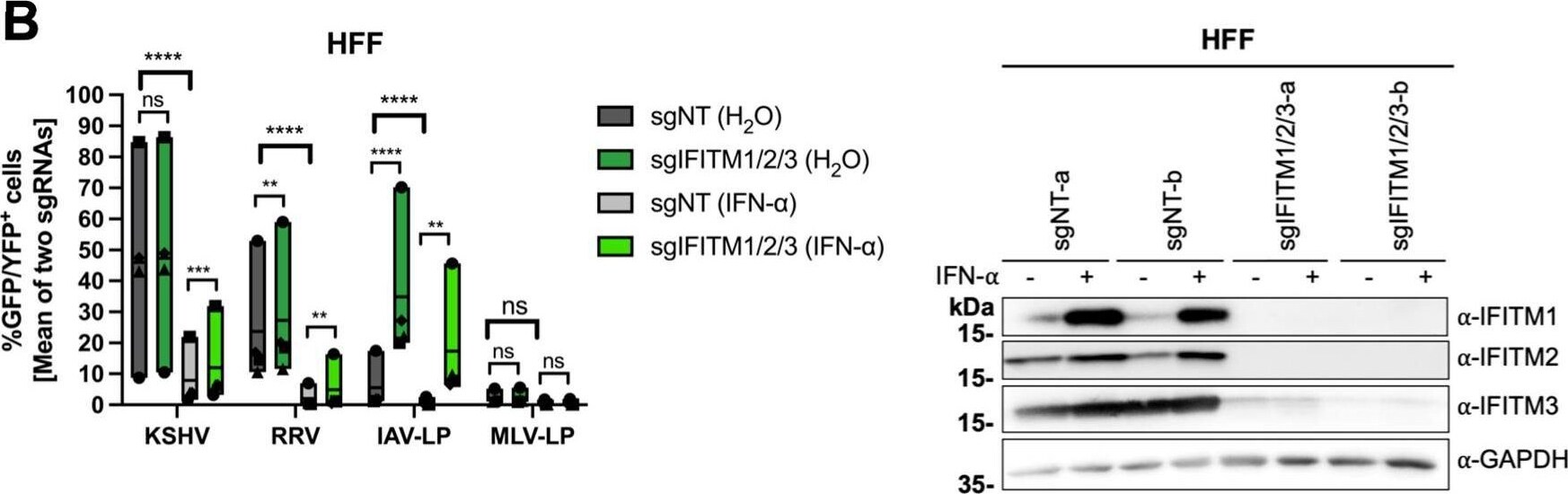
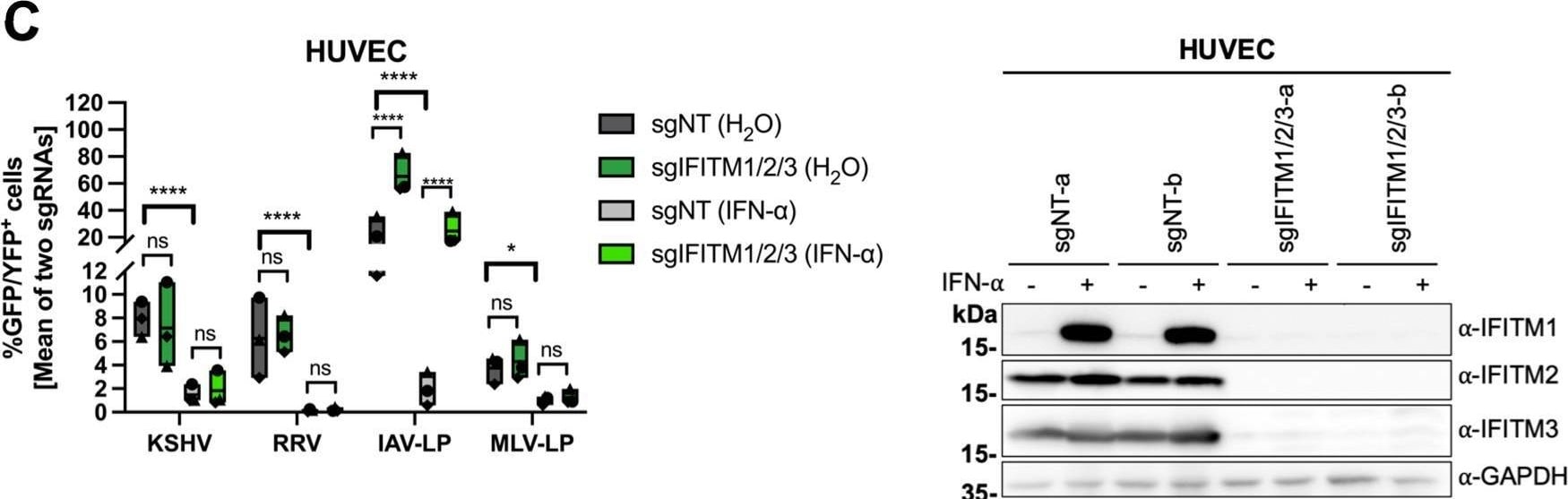
IFITM1/2/3 triple knockout enhances KSHV and RRV infection in A549 cells and HFF. A549 cells (A), HFF (B), and HUVEC (C) were transduced with lentiviral vectors encoding Cas9 and the sgRNAs shown in Fig. 2. (A to C, left panels) IFITM knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) or control (sgNT-a, sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O (control) and infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter gene. The graph shows individual data points representing averaged values for GFP+/YFP+ cells of either two nontargeting (sgNT-a, sgNT-b) or IFITM1/2/3 knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) transduced cells and floating bars representing the mean averaged from results of four independent experiments for A549 cells and HFF (A and B) and three independent experiments for HUVEC (C). Infections for each single experiment were performed in triplicate for each condition. Data points from the same experiment are labeled with identical symbols. The different sgRNAs were treated as biological replicates within each experiment. Statistical significance was determined by two‐way analysis of variance (ANOVA), and P values were corrected for all possible multiple comparisons within one family by Tukey’s method (nonsignificant [ns], P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM knockout (sgIFITM1/2/3-a or sgIFITM1/2/3-b) or control (sgNT-a or sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O. Indicated IFITM expression was detected with antibodies shown in Fig. 1A; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
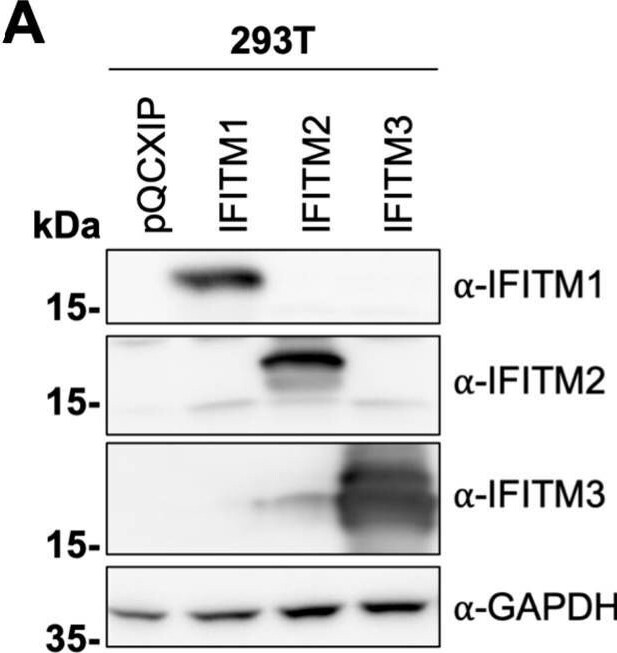
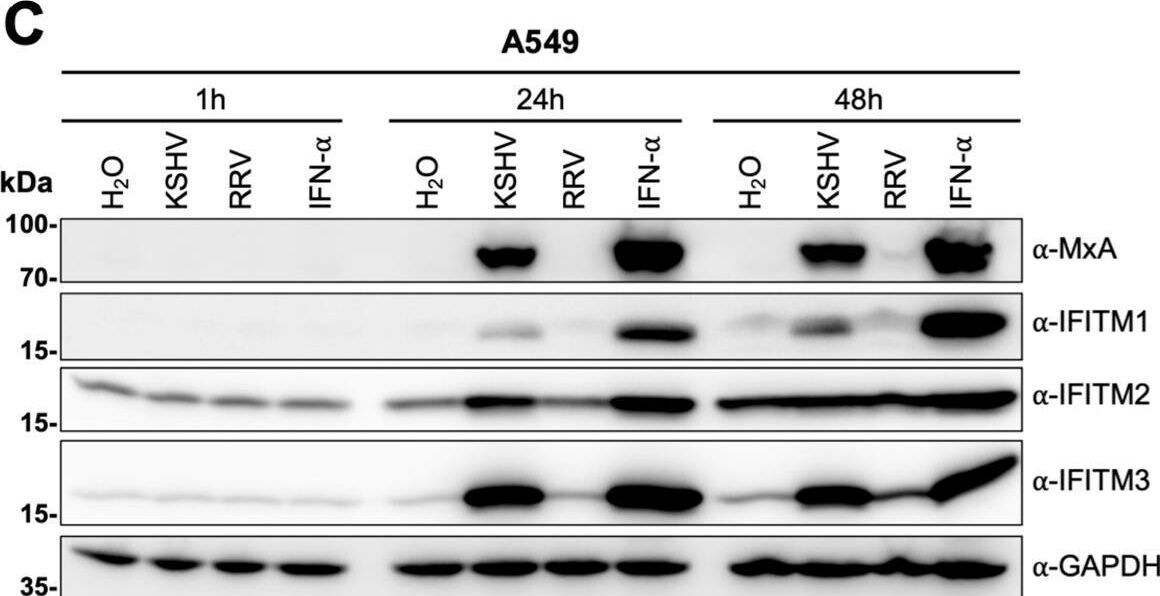
KSHV induces IFITM1, IFITM2, and IFITM3 expression in A549 cells. (A) Western blot of 293T cells transduced with pQCXIP constructs to express IFITM1 to -3 or pQCXIP (empty vector). IFITMs were detected using the respective IFITM antibody, and GAPDH served as a loading control. (B and C) Fluorescence microscopy images (scale bar, 200 μm) (B) and Western blot analysis (C) of A549 cells infected with KSHV-GFP or RRV-YFP or treated with H2O or IFN-alpha (5,000 U/ml) for the indicated time and harvested using SDS sample buffer. IFITM expression was detected with antibodies shown in panel A. MxA served as control for IFN-stimulated gene induction; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
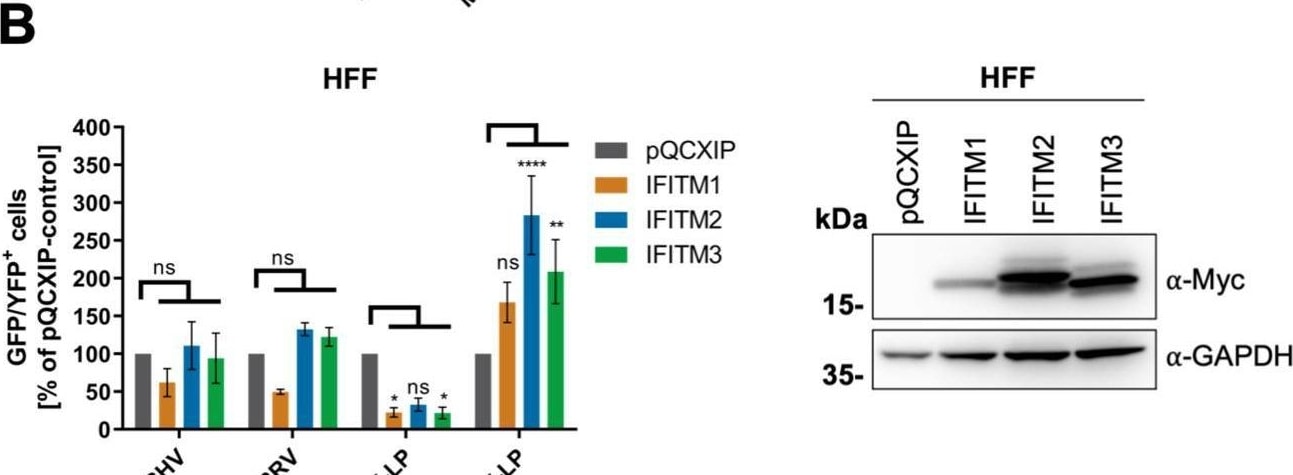
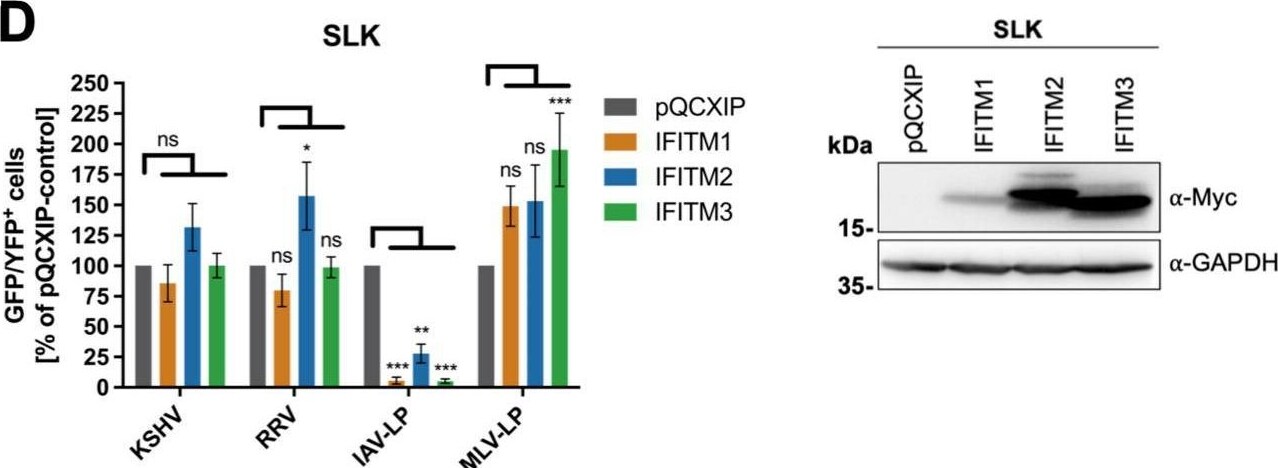
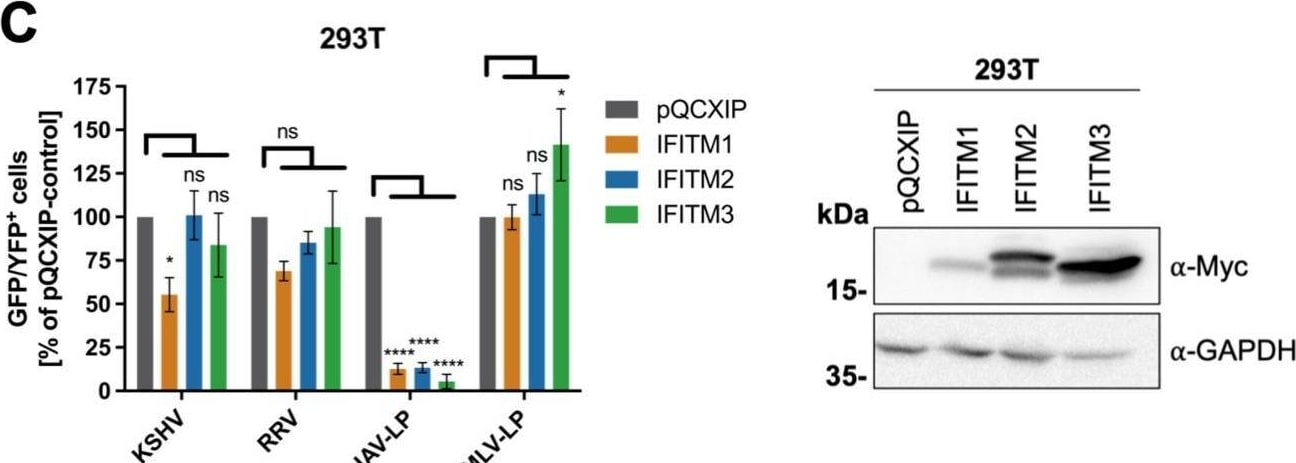
Overexpression of IFITM1 inhibits KSHV and RRV infection in a cell-specific manner. A549 cells (A), HFF (B), 293T cells (C), and SLK cells (D) were transduced with pQCXIP constructs to express IFITM1-3 or pQCXIP (empty vector). (A to D, left panels) IFITM-overexpressing cells were infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter genes. The data show values normalized to pQCXIP empty vector, which was set to 100%, and the error bars represent the standard error of the mean of results of four independent experiments, each performed in triplicate. Statistical significance was determined by ordinary two-way ANOVA, and P values were corrected for multiple comparisons by Dunnett’s method (ns, P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM-overexpressing cells. Expression of myc-tagged IFITMs was determined using anti-myc antibody; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
Overexpression of IFITM1 inhibits KSHV and RRV infection in a cell-specific manner. A549 cells (A), HFF (B), 293T cells (C), and SLK cells (D) were transduced with pQCXIP constructs to express IFITM1-3 or pQCXIP (empty vector). (A to D, left panels) IFITM-overexpressing cells were infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter genes. The data show values normalized to pQCXIP empty vector, which was set to 100%, and the error bars represent the standard error of the mean of results of four independent experiments, each performed in triplicate. Statistical significance was determined by ordinary two-way ANOVA, and P values were corrected for multiple comparisons by Dunnett’s method (ns, P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM-overexpressing cells. Expression of myc-tagged IFITMs was determined using anti-myc antibody; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
IFITM1/2/3 triple knockout enhances KSHV and RRV infection in A549 cells and HFF. A549 cells (A), HFF (B), and HUVEC (C) were transduced with lentiviral vectors encoding Cas9 and the sgRNAs shown in Fig. 2. (A to C, left panels) IFITM knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) or control (sgNT-a, sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O (control) and infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter gene. The graph shows individual data points representing averaged values for GFP+/YFP+ cells of either two nontargeting (sgNT-a, sgNT-b) or IFITM1/2/3 knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) transduced cells and floating bars representing the mean averaged from results of four independent experiments for A549 cells and HFF (A and B) and three independent experiments for HUVEC (C). Infections for each single experiment were performed in triplicate for each condition. Data points from the same experiment are labeled with identical symbols. The different sgRNAs were treated as biological replicates within each experiment. Statistical significance was determined by two‐way analysis of variance (ANOVA), and P values were corrected for all possible multiple comparisons within one family by Tukey’s method (nonsignificant [ns], P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM knockout (sgIFITM1/2/3-a or sgIFITM1/2/3-b) or control (sgNT-a or sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O. Indicated IFITM expression was detected with antibodies shown in Fig. 1A; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
IFITM1/2/3 triple knockout enhances KSHV and RRV infection in A549 cells and HFF. A549 cells (A), HFF (B), and HUVEC (C) were transduced with lentiviral vectors encoding Cas9 and the sgRNAs shown in Fig. 2. (A to C, left panels) IFITM knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) or control (sgNT-a, sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O (control) and infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter gene. The graph shows individual data points representing averaged values for GFP+/YFP+ cells of either two nontargeting (sgNT-a, sgNT-b) or IFITM1/2/3 knockout (sgIFITM1/2/3-a, sgIFITM1/2/3-b) transduced cells and floating bars representing the mean averaged from results of four independent experiments for A549 cells and HFF (A and B) and three independent experiments for HUVEC (C). Infections for each single experiment were performed in triplicate for each condition. Data points from the same experiment are labeled with identical symbols. The different sgRNAs were treated as biological replicates within each experiment. Statistical significance was determined by two‐way analysis of variance (ANOVA), and P values were corrected for all possible multiple comparisons within one family by Tukey’s method (nonsignificant [ns], P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM knockout (sgIFITM1/2/3-a or sgIFITM1/2/3-b) or control (sgNT-a or sgNT-b) cells treated with IFN-alpha (5,000 U/ml) or H2O. Indicated IFITM expression was detected with antibodies shown in Fig. 1A; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
KSHV induces IFITM1, IFITM2, and IFITM3 expression in A549 cells. (A) Western blot of 293T cells transduced with pQCXIP constructs to express IFITM1 to -3 or pQCXIP (empty vector). IFITMs were detected using the respective IFITM antibody, and GAPDH served as a loading control. (B and C) Fluorescence microscopy images (scale bar, 200 μm) (B) and Western blot analysis (C) of A549 cells infected with KSHV-GFP or RRV-YFP or treated with H2O or IFN-alpha (5,000 U/ml) for the indicated time and harvested using SDS sample buffer. IFITM expression was detected with antibodies shown in panel A. MxA served as control for IFN-stimulated gene induction; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of IFITM1 by Western Blot
Overexpression of IFITM1 inhibits KSHV and RRV infection in a cell-specific manner. A549 cells (A), HFF (B), 293T cells (C), and SLK cells (D) were transduced with pQCXIP constructs to express IFITM1-3 or pQCXIP (empty vector). (A to D, left panels) IFITM-overexpressing cells were infected with KSHV-GFP, RRV-YFP, IAV lentiviral pseudotype (IAV-LP), or MLV lentiviral pseudotype (MLV-LP). Infection was measured using flow cytometry to detect expression of the fluorescent reporter genes. The data show values normalized to pQCXIP empty vector, which was set to 100%, and the error bars represent the standard error of the mean of results of four independent experiments, each performed in triplicate. Statistical significance was determined by ordinary two-way ANOVA, and P values were corrected for multiple comparisons by Dunnett’s method (ns, P > 0.05; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001; ****, P ≤ 0.0001). (A to C, right panels) Representative Western blots of IFITM-overexpressing cells. Expression of myc-tagged IFITMs was determined using anti-myc antibody; GAPDH served as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/34933450), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human IFITM1 Antibody
Application
Recommended Usage
Immunocytochemistry
5-15 µg/mL
Sample: Immersion fixed K562 human chronic myelogenous leukemia cell line
Sample: Immersion fixed K562 human chronic myelogenous leukemia cell line
Western Blot
1 µg/mL
Sample: Jurkat human acute T cell leukemia cell line, TF‑1 human erythroleukemic cell line, human placenta tissue
Sample: Jurkat human acute T cell leukemia cell line, TF‑1 human erythroleukemic cell line, human placenta tissue
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: IFITM1
Long Name
Interferon-induced Transmembrane Protein 1
Alternate Names
CD225, IFI17, Leu-13
Gene Symbol
IFITM1
UniProt
Additional IFITM1 Products
Product Documents for Human IFITM1 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human IFITM1 Antibody
For research use only
Related Research Areas
Citations for Human IFITM1 Antibody
Customer Reviews for Human IFITM1 Antibody
There are currently no reviews for this product. Be the first to review Human IFITM1 Antibody and earn rewards!
Have you used Human IFITM1 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...