SARS-CoV-2 Spike S1 Subunit Antibody
R&D Systems | Catalog # MAB105805

Key Product Details
Species Reactivity
Applications
Label
Antibody Source
Product Specifications
Immunogen
Accession # YP_009724390.1
Specificity
Clonality
Host
Isotype
Scientific Data Images for SARS-CoV-2 Spike S1 Subunit Antibody
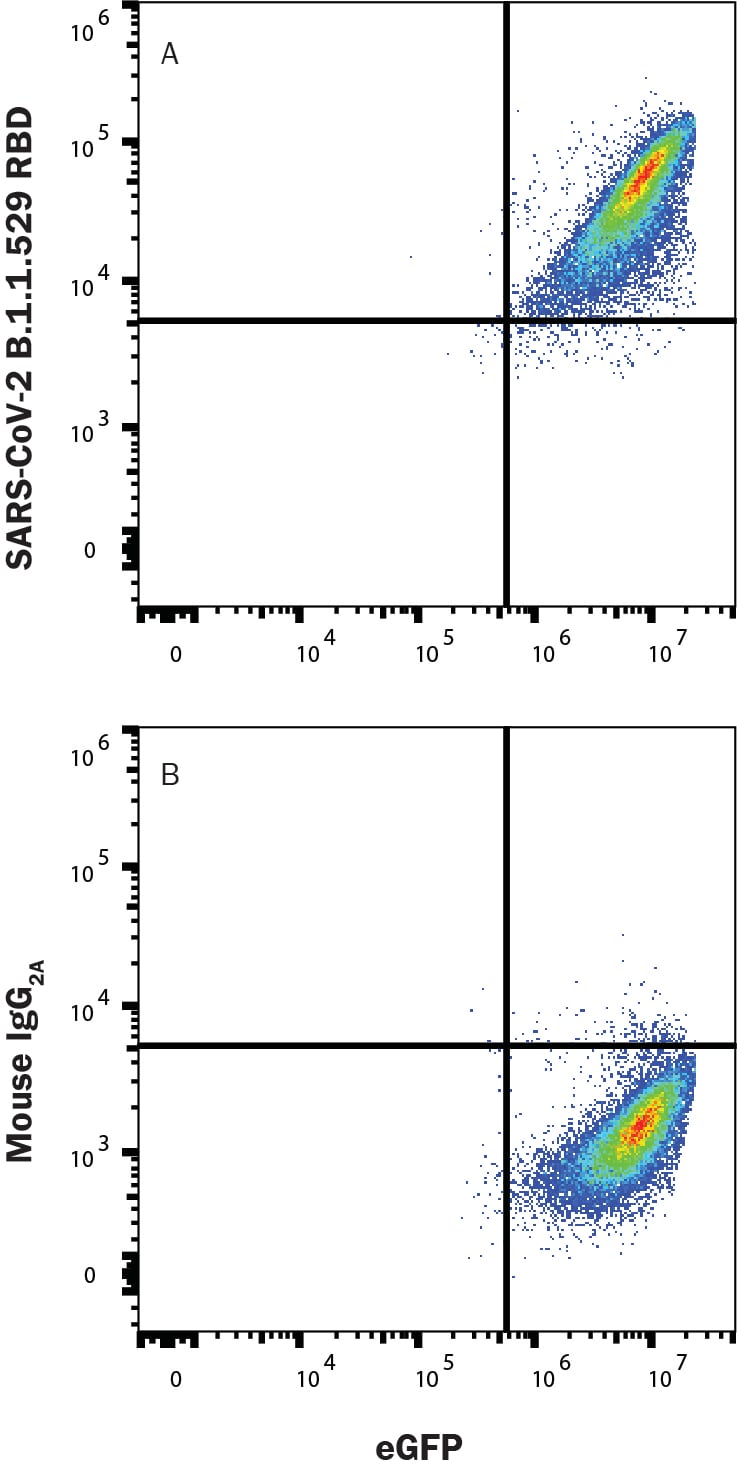
Detection of SARS-CoV-2 Omicron protein (B.1.1.529 variant) bound to ACE-2 in HEK293 Human Cell Line Transfected with Human ACE-2 and eGFP by Flow Cytometry.
HEK293 human embryonic kidney cell line transfected with human ACE-2 and eGFP was incubated with Recombinant SARS-CoV-2 Omicron His-Tag protein (B.1.1.529 variant; 11056-CV), then stained with (A) Mouse Anti-SARS-CoV-2 Spike S1 Monoclonal Antibody (Catalog # MAB105805) or (B) Mouse IgG2A Isotype Control Antibody (MAB003) followed by Allophycocyanin-conjugated Anti-Mouse IgG Secondary Antibody (F0101B). Staining was performed using our Staining Membrane-associated Proteins protocol.Detection of SARS-CoV-2 Spike S1 protein bound to ACE-2 in HEK293 Human Cell Line Transfected with Human ACE-2 and eGFP by Flow Cytometry.
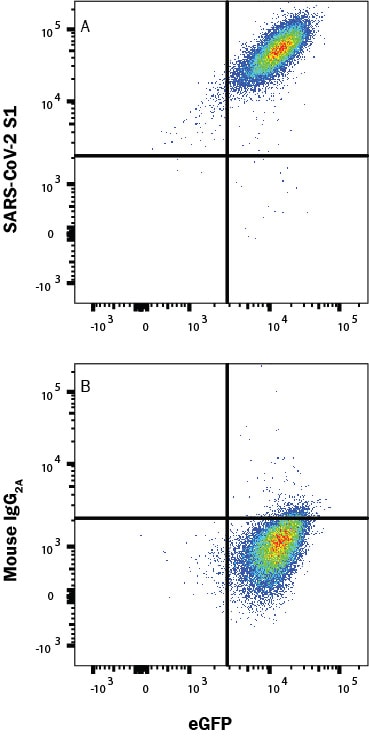
HEK293 human embryonic kidney cell line transfected with human ACE-2 and eGFP was incubated with Recombinant SARS-CoV-2 Spike S1 Subunit His-Tag protein (10522-CV), then stained with (A) Mouse Anti-SARS-CoV-2 Spike S1 Monoclonal Antibody (Catalog # MAB105805) or (B) Mouse IgG2A Isotype Control Antibody (MAB003) followed by Allophycocyanin-conjugated Anti-Mouse IgG Secondary Antibody (F0101B). Staining was performed using our Staining Membrane-associated Proteins protocol.Applications for SARS-CoV-2 Spike S1 Subunit Antibody
CyTOF-ready
Flow Cytometry
Sample: SARS-CoV-2 Spike S1 protein (Catalog # 10522-CV) and SARS-CoV-2 B.1.1.529 S RBD (Catalog 11056-CV) bound to ACE-2 in HEK293 Human Cell Line Transfected with Human ACE-2 and eGFP.
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Reconstitution
Reconstitute at 0.5 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Formulation
Shipping
Stability & Storage
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: Spike S1 Subunit
References
- Wu, F. et al. (2020) Nature 579:265.
- Tortorici, M.A. and D. Veesler (2019). Adv. Virus Res. 105:93.
- Bosch, B.J. et al. (2003) J. Virol. 77:8801.
- Belouzard, S. et al. (2009) Proc. Natl. Acad. Sci. 106:5871.
- Millet, J.K. and G. R. Whittaker (2015) Virus Res. 202:120.
- Yuan, Y. et al. (2017) Nat. Commun. 8:15092.
- Walls, A.C. et al. (2010) Cell 180:281.
- Jiang, S. et al. (2020) Trends. Immunol. https://doi.org/10.1016/j.it.2020.03.007.
- Ortega, J.T. et al. (2020) EXCLI J. 19:410.
- Wrapp, D. et al. (2020) Science 367:1260.
- Tai, W. et al. (2020) Cell. Mol. Immunol. https://doi.org/10.1016/j.it.2020.03.007.
- Okba, N. M. A. et al. (2020). Emerg. Infect. Dis. https://doi.org/10.3201/eid2607.200841.
- Wang, X. et al. (2020) https://doi.org/10.1038/s41423-020-0424-9.
- Wang, K. et al. (2020) bioRxiv https://www.biorxiv.org/content/10.1101/2020.03.14.988345v1.
Long Name
Alternate Names
UniProt
Additional Spike S1 Subunit Products
Product Documents for SARS-CoV-2 Spike S1 Subunit Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for SARS-CoV-2 Spike S1 Subunit Antibody
For research use only
Related Research Areas
Customer Reviews for SARS-CoV-2 Spike S1 Subunit Antibody
There are currently no reviews for this product. Be the first to review SARS-CoV-2 Spike S1 Subunit Antibody and earn rewards!
Have you used SARS-CoV-2 Spike S1 Subunit Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Liperfluo
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- View all Protocols, Troubleshooting, Illustrated assays and Webinars