Matrix metalloproteinases are a family of zinc and calcium dependent endopeptidases with the combined ability to degrade all the components of the extracellular matrix. MMP-13 (Collagenase-3) has been demonstrated to degrade a range of extracellular matrix proteins, including collagen types I, II, III, IV, IX, X and XIV, gelatin, aggrecan, perlecan and fibronectin. MMP-13 is distinguished from the other human collagenases by its effecient degradation of type II collagen. MMP-13 is expressed by fibroblasts, chrondrocytes and squamous epithelial cells. Structurally, MMP-13 may be divided into several distinct domains; a pro-domain which is cleaved upon activation; a catalytic domain containing the zinc binding site; a short hinge region and a carboxyl terminal (hemopexin-like) domain.


Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human, Rabbit
Applications
Validated:
Immunohistochemistry, Western Blot, Immunoprecipitation
Cited:
Immunohistochemistry, Western Blot, ELISA Development
Label
Unconjugated
Antibody Source
Polyclonal Goat IgG
Loading...
Product Specifications
Immunogen
Mouse myeloma cell line NS0-derived recombinant human MMP-13
Leu20-Cys471
Accession # P45452
Leu20-Cys471
Accession # P45452
Specificity
Detects human MMP-13 in Western blots. In direct ELISAs, less than 5% cross-reactivity with recombinant human (rh) MMP-3 and rhMMP-8 is observed.
Clonality
Polyclonal
Host
Goat
Isotype
IgG
Scientific Data Images for Human MMP‑13 Antibody
MMP‑13 in Human Ovarian Cancer Tissue.
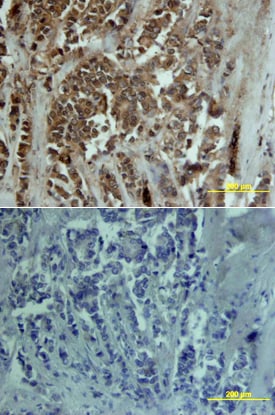
MMP-13 was detected in immersion fixed paraffin-embedded sections of human ovarian cancer tissue using 15 µg/mL Goat Anti-Human MMP-13 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF511) overnight at 4 °C. Tissue was stained with the Anti-Goat HRP-DAB Cell & Tissue Staining Kit (brown; Catalog # CTS008) and counterstained with hematoxylin (blue). View our protocol for Chromogenic IHC Staining of Paraffin-embedded Tissue Sections.MMP‑13 in Human Breast.
MMP-13 was detected in immersion fixed paraffin-embedded sections of human breast array using Goat Anti-Human MMP-13 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF511) at 15 µg/mL overnight at 4 °C. Tissue was stained using the Anti-Goat HRP-DAB Cell & Tissue Staining Kit (brown; Catalog # CTS008) and counterstained with hematoxylin (blue). Lower panel shows a lack of labeling if primary antibodies are omitted and tissue is stained only with secondary antibody followed by incubation with detection reagents. View our protocol for Chromogenic IHC Staining of Paraffin-embedded Tissue Sections.Detection of MMP-13 by Western Blot
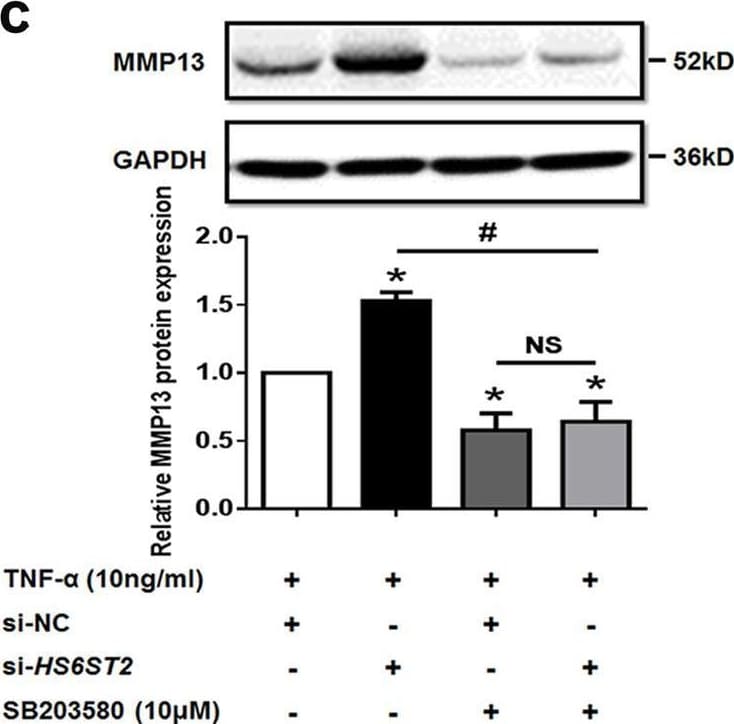
HS6ST2 could regulate the matrix degradation depending on the activity of p38 Malka SW1353 cells were transfected by si-HS6ST2 or negative control (si-NC) with or without TNF-alpha for 24 h, and the protein expression of p-p38 and p38 was detected by western blotting. Asterisk (*): compared with the si-NC group. b Under stimulation with TNF-alpha, SW1353 cells were transfected by empty vector (FLAG-Ctrl) or pcDNA3.1-FLAG-HS6ST2 vector (FLAG-HS6ST2) under treatment with mimic miR-23b-3p, and p-p38 and total p38 were determined by western blotting. Asterisk (*): compared with the mimic NC group. c SW1353 cells were transfected with HS6ST2 siRNA with or without p38 MAPK inhibitor SB203580 (10 μM) after treatment of TNF-alpha, and the protein expression of MMP13 was determined by western blotting. Each relative expression of phosphorylation form was normalized by the total form. d Schematic representation of miR-23b-3p–HS6ST2 axis-mediated catabolic effects in human chondrocyte. GAPDH was used as internal controls in western blotting detection. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05. NS stands for not significant Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
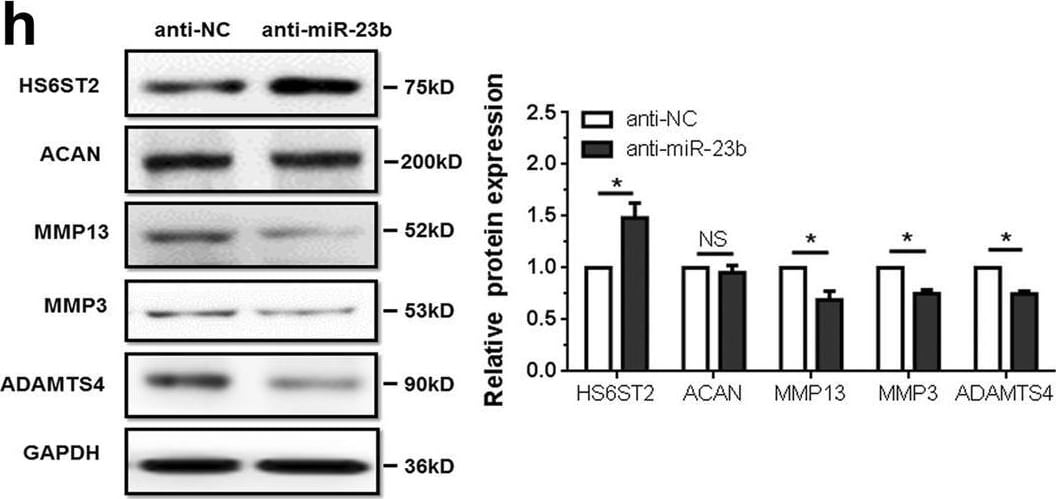
MiR-23b-3p inhibits HS6ST2 expression by targeting HS6ST2 mRNA and effects specific gene expression in chondrocytes. A, b Dual luciferase reporter assay to validate target relationship between HS6ST2 and miR-23b-3p. SW1353 cells were transfected with mimic miR-23b-3p (a) or anti-miR-23b-3p (b) by control vector, pmirGLO-HS6ST2 wild-type 3′UTR vector or pmirGLO-HS6ST2 mutant 3′UTR vector, respectively, for 48 h. c, d Stem-loop RT-qPCR and western blotting results in SW1353 cells (c) or C28/I2 cells (d) transfected with 10 nM mimic miR-23b-3p (left panel) or 50 nM anti-miR-23b-3p sequence (right panel). e, f RT-qPCR (e) and western blotting (f) results of cartilage-specific gene expression in SW1353 cells transfected with mimic miR-23b-3p or negative control. g, h RT-qPCR (g) and western blotting (h) results of cartilage-specific gene expression in SW1353 cells transfected with anti-miR-23b-3p or negative control. RNA was harvested at 24 h, while protein was isolated at 48 h after transfection. U6 snRNA was used as internal controls in miRNA stem-loop RT-qPCR detection and GAPDH was used as internal controls in mRNA RT-qPCR and western blotting detection. Bars represent standard error of the mean (SEM) from three independent experiments. Mann–Whitney U test was used to identify statistical differences between two groups. *P value < 0.05 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
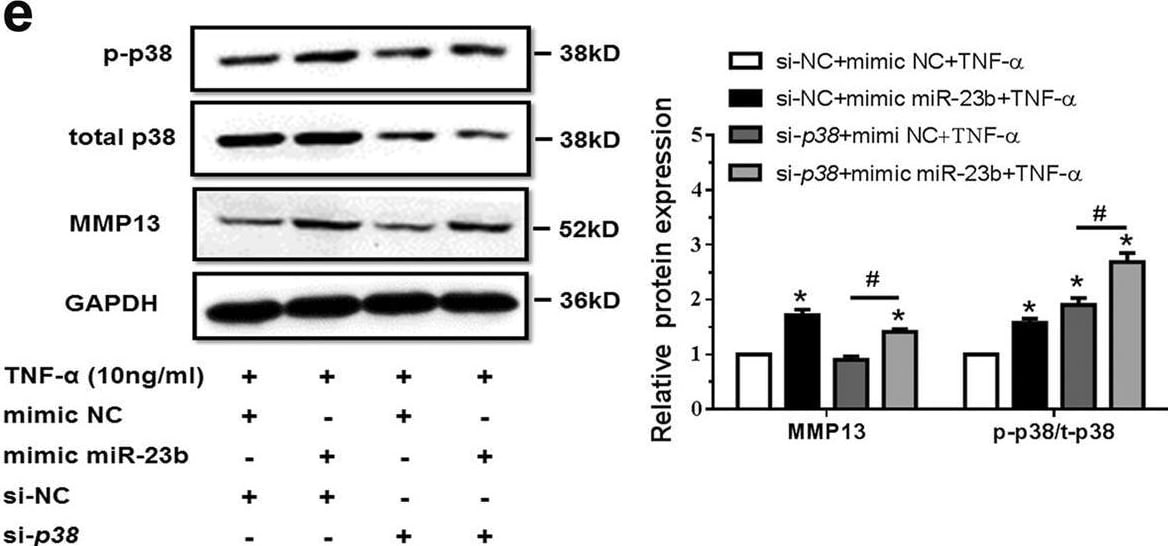
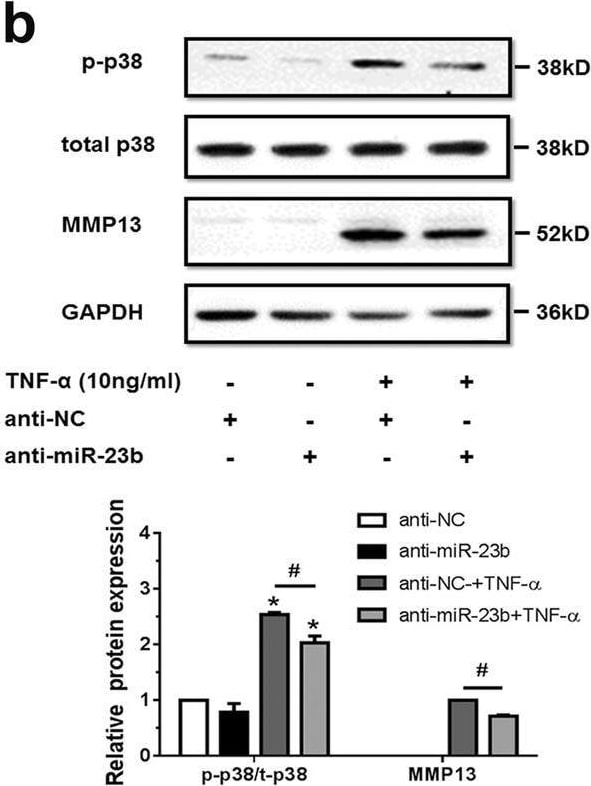
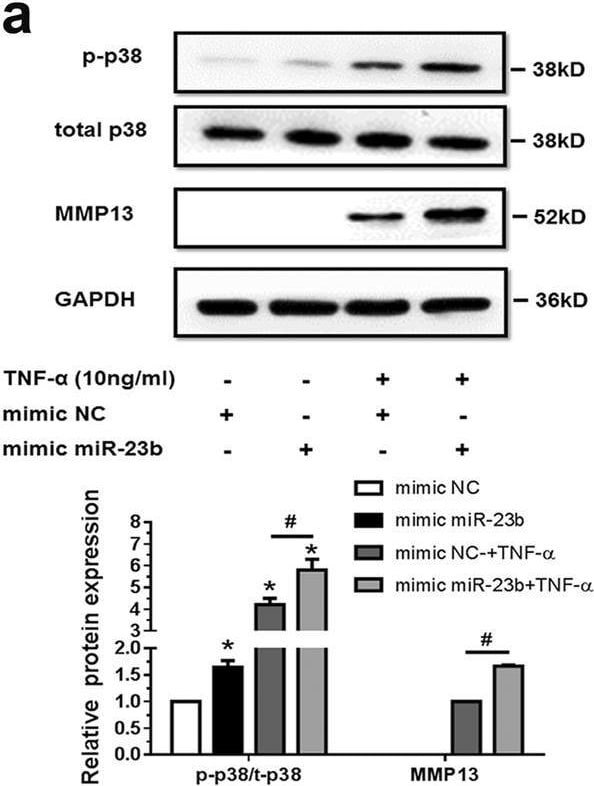
MiR-23b-3p could enhance matrix degradation by means of regulating activity of p38 MAPK in human chondrocytes under TNF-alpha treatment. A, b SW1353 cells were transfected by mimic miR-23b-3p (a) or anti-miR-23b-3p sequence (b) with or without TNF-alpha for 24 h, and the protein expression of phosphorylation form of p38 MAPK (p-p38), total p38 MAPK, and MMP13 was detected by western blotting. Asterisk (*): compared with the mimic NC (a) or anti-NC group (b). c SW1353 cells were treated with mimic miR-23b-3p with or without p38 MAPK inhibitor SB203580 (10 μM) under stimulation of TNF-alpha. The protein expression of MMP13 was determined by western blotting. Asterisk (*): compared with the mimic NC group. d Interfering efficiency of siRNAs against p38 MAPK was determined by western blotting under 50 nM p38 siRNA, 50 nM negative control (NC), and Mock (transfection regent only) transfection for 48 h. e Under stimulation with TNF-alpha, SW1353 cells were transfected by mimic miR-23b-3p under treatment with si-p38 mixture (containing three siRNA target sequences), and p-p38, total p38, and MMP13 were determined by western blotting. Asterisk (*): compared with mimic NC or si-NC. Each relative expression of phosphorylation form was normalized by the total form. GAPDH was used as internal controls in western blotting detection. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05. NS stands for not significant Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
MiR-23b-3p could enhance matrix degradation by means of regulating activity of p38 MAPK in human chondrocytes under TNF-alpha treatment. A, b SW1353 cells were transfected by mimic miR-23b-3p (a) or anti-miR-23b-3p sequence (b) with or without TNF-alpha for 24 h, and the protein expression of phosphorylation form of p38 MAPK (p-p38), total p38 MAPK, and MMP13 was detected by western blotting. Asterisk (*): compared with the mimic NC (a) or anti-NC group (b). c SW1353 cells were treated with mimic miR-23b-3p with or without p38 MAPK inhibitor SB203580 (10 μM) under stimulation of TNF-alpha. The protein expression of MMP13 was determined by western blotting. Asterisk (*): compared with the mimic NC group. d Interfering efficiency of siRNAs against p38 MAPK was determined by western blotting under 50 nM p38 siRNA, 50 nM negative control (NC), and Mock (transfection regent only) transfection for 48 h. e Under stimulation with TNF-alpha, SW1353 cells were transfected by mimic miR-23b-3p under treatment with si-p38 mixture (containing three siRNA target sequences), and p-p38, total p38, and MMP13 were determined by western blotting. Asterisk (*): compared with mimic NC or si-NC. Each relative expression of phosphorylation form was normalized by the total form. GAPDH was used as internal controls in western blotting detection. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05. NS stands for not significant Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
MiR-23b-3p could enhance matrix degradation by means of regulating activity of p38 MAPK in human chondrocytes under TNF-alpha treatment. A, b SW1353 cells were transfected by mimic miR-23b-3p (a) or anti-miR-23b-3p sequence (b) with or without TNF-alpha for 24 h, and the protein expression of phosphorylation form of p38 MAPK (p-p38), total p38 MAPK, and MMP13 was detected by western blotting. Asterisk (*): compared with the mimic NC (a) or anti-NC group (b). c SW1353 cells were treated with mimic miR-23b-3p with or without p38 MAPK inhibitor SB203580 (10 μM) under stimulation of TNF-alpha. The protein expression of MMP13 was determined by western blotting. Asterisk (*): compared with the mimic NC group. d Interfering efficiency of siRNAs against p38 MAPK was determined by western blotting under 50 nM p38 siRNA, 50 nM negative control (NC), and Mock (transfection regent only) transfection for 48 h. e Under stimulation with TNF-alpha, SW1353 cells were transfected by mimic miR-23b-3p under treatment with si-p38 mixture (containing three siRNA target sequences), and p-p38, total p38, and MMP13 were determined by western blotting. Asterisk (*): compared with mimic NC or si-NC. Each relative expression of phosphorylation form was normalized by the total form. GAPDH was used as internal controls in western blotting detection. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05. NS stands for not significant Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
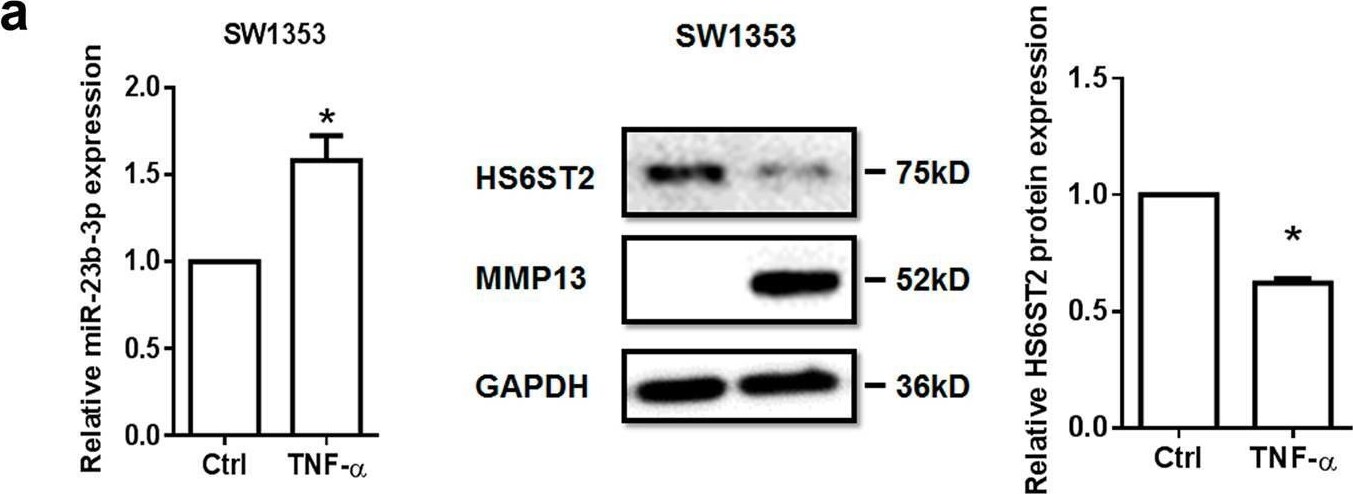
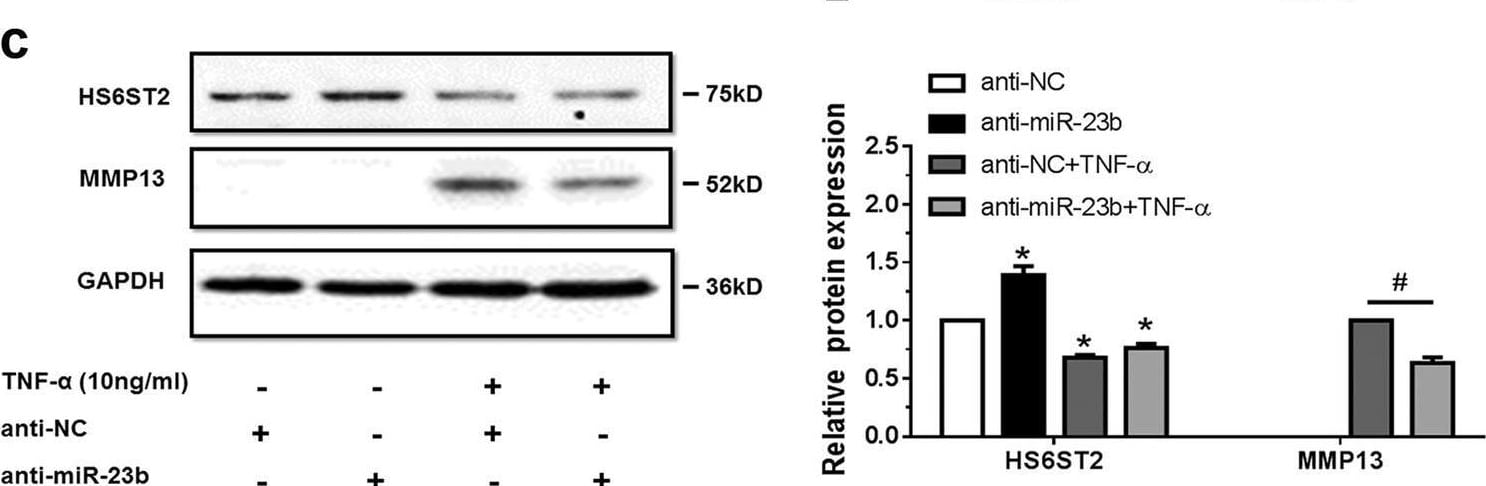
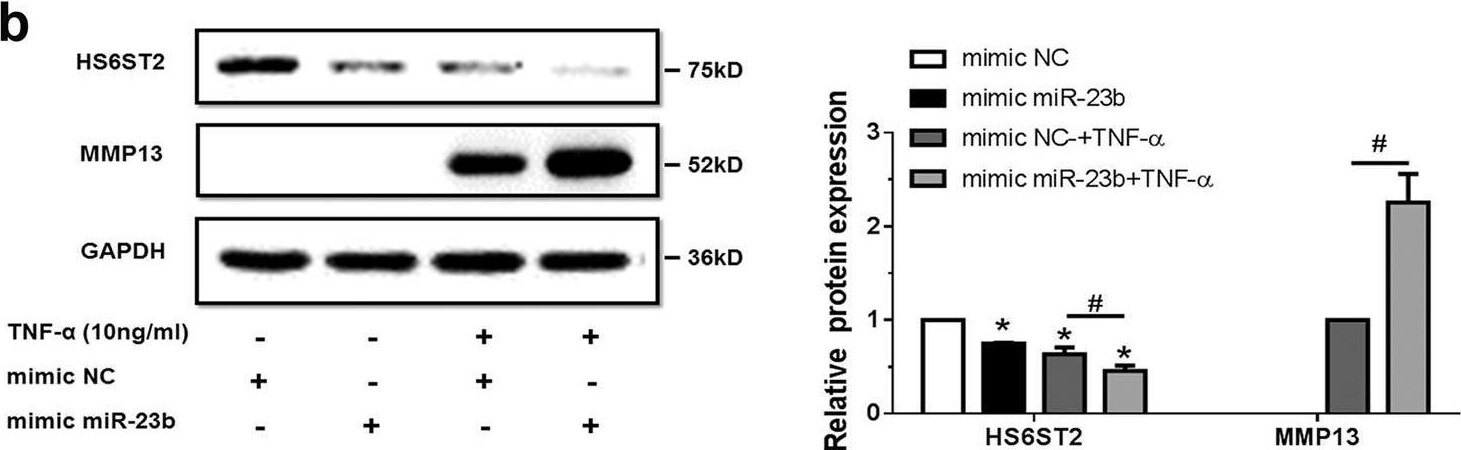
TNF-alpha -treated chondrocytes increases matrix degradation regulated by miR-23b-3p.a Stem-loop RT-qPCR result of miR-23b-3p (left panel) and protein levels of HS6ST2 and MMP13 (right panel) were assayed in SW1353 cells under stimulation by 10 ng/ml TNF-alpha for 24 h. b, c Western blotting result of HS6ST2 and MMP13 protein in SW1353 cells transfected with 10 nM mimic miR-23b-3p (b) or 50 nM anti-miR-23b-3p sequence (c) and stimulated by 10 ng/ml TNF-alpha for 24 h. d, e Toluidine blue staining results of SW1353 cells transfected with 10 nM mimic miR-23b-3p (d) or 50 nM anti-miR-23b-3p sequence (e) and stimulated by 10 ng/ml TNF-alpha for 24 h. Scale bar, 100 μm. Lower panel, statistical analysis of average optical density of matrix staining of toluidine blue. Asterisk (*): compared with mimic NC or anti-NC group. U6 snRNA and GAPDH were used as internal controls in RT-qPCR for miRNA and western blotting detection, respectively. Bars represent standard error of the mean (SEM) from three independent experiments. One representative result and quantitative data from three independent western blotting and toluidine blue staining. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
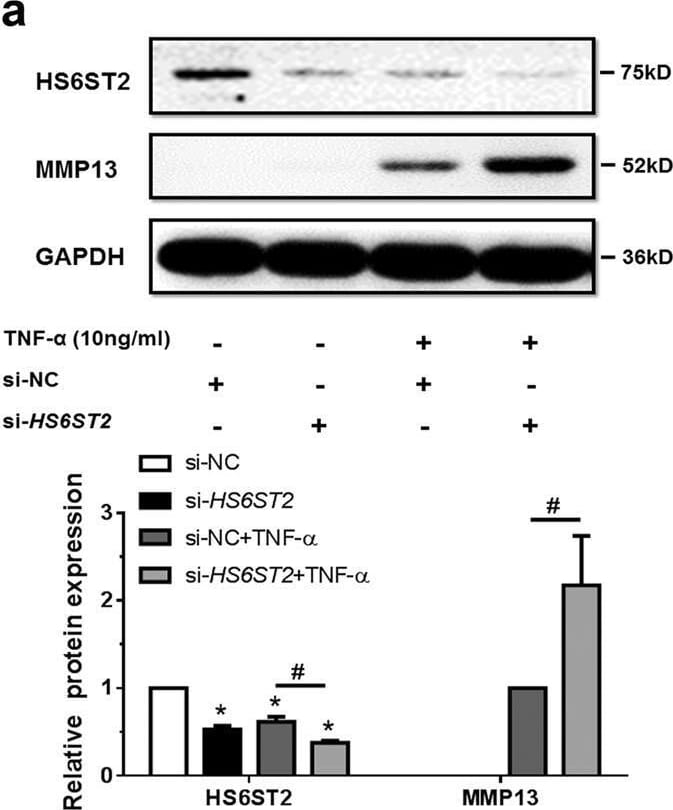
MiR-23b-3p could enhance matrix degradation in human chondrocytes via regulating HS6ST2.a, b SW1353 cells transfected with 50 nM HS6ST2 siRNA mixture (containing three target sequences, si-HS6ST2) or negative control (si-NC) were stimulated by 10 ng/ml TNF-alpha for 24 h. Protein level of HS6ST2 and MMP13 were assayed by western blotting (a) and matrix content of chondrocytes was determined by toluidine blue staining (b). Asterisk (*): compared with the si-NC group. c, d SW1353 cells transfected with empty vector (FLAG-Ctrl) or pcDNA3.1-FLAG-HS6ST2 vector (FLAG-HS6ST2) for 24 h were stimulated by TNF-alpha for another 24 h. The protein expression of MMP13 was assayed by western blotting (c) and matrix content of chondrocytes was determined by toluidine blue staining (d). Asterisk (*): compared with the FLAG-Ctrl group. e, f Under stimulation with TNF-alpha, SW1353 cells were treated with mimic NC or mimic miR-23b-3p for 24 h and then transfected with empty vector (FLAG-Ctrl) or pcDNA3.1-FLAG-HS6ST2 vector (FLAG-HS6ST2) for another 24 h to observe the rescuing effect of mimic miR-23b-3p in MMP13 expression (e) and matrix content (f). Asterisk (*): compared with the mimic NC group. GAPDH was used as internal controls in western blotting detection. In toluidine blue staining results, scale bar, 100 μm. Lower panel, statistical analysis of average optical density of matrix staining of toluidine blue. Bars represent standard error of the mean (SEM) from three independent experiments. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05. NS stands for not significant Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
TNF-alpha -treated chondrocytes increases matrix degradation regulated by miR-23b-3p.a Stem-loop RT-qPCR result of miR-23b-3p (left panel) and protein levels of HS6ST2 and MMP13 (right panel) were assayed in SW1353 cells under stimulation by 10 ng/ml TNF-alpha for 24 h. b, c Western blotting result of HS6ST2 and MMP13 protein in SW1353 cells transfected with 10 nM mimic miR-23b-3p (b) or 50 nM anti-miR-23b-3p sequence (c) and stimulated by 10 ng/ml TNF-alpha for 24 h. d, e Toluidine blue staining results of SW1353 cells transfected with 10 nM mimic miR-23b-3p (d) or 50 nM anti-miR-23b-3p sequence (e) and stimulated by 10 ng/ml TNF-alpha for 24 h. Scale bar, 100 μm. Lower panel, statistical analysis of average optical density of matrix staining of toluidine blue. Asterisk (*): compared with mimic NC or anti-NC group. U6 snRNA and GAPDH were used as internal controls in RT-qPCR for miRNA and western blotting detection, respectively. Bars represent standard error of the mean (SEM) from three independent experiments. One representative result and quantitative data from three independent western blotting and toluidine blue staining. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MMP-13 by Western Blot
TNF-alpha -treated chondrocytes increases matrix degradation regulated by miR-23b-3p.a Stem-loop RT-qPCR result of miR-23b-3p (left panel) and protein levels of HS6ST2 and MMP13 (right panel) were assayed in SW1353 cells under stimulation by 10 ng/ml TNF-alpha for 24 h. b, c Western blotting result of HS6ST2 and MMP13 protein in SW1353 cells transfected with 10 nM mimic miR-23b-3p (b) or 50 nM anti-miR-23b-3p sequence (c) and stimulated by 10 ng/ml TNF-alpha for 24 h. d, e Toluidine blue staining results of SW1353 cells transfected with 10 nM mimic miR-23b-3p (d) or 50 nM anti-miR-23b-3p sequence (e) and stimulated by 10 ng/ml TNF-alpha for 24 h. Scale bar, 100 μm. Lower panel, statistical analysis of average optical density of matrix staining of toluidine blue. Asterisk (*): compared with mimic NC or anti-NC group. U6 snRNA and GAPDH were used as internal controls in RT-qPCR for miRNA and western blotting detection, respectively. Bars represent standard error of the mean (SEM) from three independent experiments. One representative result and quantitative data from three independent western blotting and toluidine blue staining. Mann–Whitney U test was used to identify statistical differences between two groups. Asterisk (*) or hash (#) stands for P value <0.05 Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/29899528), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human MMP‑13 Antibody
Application
Recommended Usage
Immunohistochemistry
5-15 µg/mL
Sample: Immersion fixed paraffin-embedded sections of human ovarian cancer tissue and human breast array
Sample: Immersion fixed paraffin-embedded sections of human ovarian cancer tissue and human breast array
Immunoprecipitation
25 µg/mL
Sample: Conditioned cell culture medium spiked with Recombinant Human MMP‑13 (Catalog # 511‑MM), see our available Western blot detection antibodies
Sample: Conditioned cell culture medium spiked with Recombinant Human MMP‑13 (Catalog # 511‑MM), see our available Western blot detection antibodies
Western Blot
0.1 µg/mL
Sample: Recombinant Human MMP‑13 Western Blot Standard (Catalog # WBC020)
Sample: Recombinant Human MMP‑13 Western Blot Standard (Catalog # WBC020)
Reviewed Applications
Read 3 reviews rated 4 using AF511 in the following applications:
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: MMP-13
References
- Jeffery, J.J. (1998) in Collagenase 3. A.J. Barrett, et al. (eds): Handbook of Proteolytic Enzymes, San Diego: Academic Press, p. 1167.
Long Name
Matrix Metalloproteinase 13
Alternate Names
MMP13
Entrez Gene IDs
4322 (Human)
Gene Symbol
MMP13
UniProt
Additional MMP-13 Products
Product Documents for Human MMP‑13 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human MMP‑13 Antibody
For research use only
Related Research Areas
Citations for Human MMP‑13 Antibody
Customer Reviews for Human MMP‑13 Antibody (3)
4 out of 5
3 Customer Ratings
Have you used Human MMP‑13 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
3 of
3 reviews
Showing All
Filter By:
-
Application: ELISASample Tested: Serum and PlasmaSpecies: HumanVerified Customer | Posted 08/20/2021
-
Application: ELISASample Tested: Serum and PlasmaSpecies: HumanVerified Customer | Posted 11/10/2017The R&D antibody MAB511 was used as the capture antibody in conjunction with this antibody (AF511) to build an ELISA. The ELISA was used to measure MMP-13 in human plasma samples.
-
Application: Western BlotSample Tested: See PMID 22521401Species: HumanVerified Customer | Posted 01/08/2015
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Immunoprecipitation Protocol
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...