TLR4 is a 100 kDa type I transmembrane glycoprotein that belongs to the mammalian Toll-Like Receptor family of pathogen pattern recognition molecules. In the literature molecular weights correspondent to 110 kDa and 130 kDa were reported for TLR4 (1). MD-2, also known as ESOP-1, is a 25 kDa secreted protein that is required for TLR4-mediated responses to bacterial lipopolysaccharide (LPS) (1‑4). The human TLR4 cDNA encodes an 839 amino acid (aa) precursor that contains a 23 aa signal sequence, a 608 aa extracellular domain (ECD), a 21 aa transmembrane segment, and a 187 aa cytoplasmic domain. TLR4 contains 21 leucine rich repeats in its ECD and one cytoplasmic Toll/IL-1 receptor (TIR) domain (5). The ECD of human TLR4 shares approximately 25% aa sequence identity with other TLRs and 60%‑74% aa sequence identity with bovine, equine, feline, mouse, rat, and porcine TLR4. The human MD-2 cDNA encodes a 160 aa precursor with an 18 aa signal sequence (5). Human MD-2 shares 20% aa sequence identity with MD-1 and 62%‑64% aa sequence identity with bovine, mouse, and rat MD-2. MD-2 associates with TLR4 on monocytes, macrophages, dendritic cells, and B cells (6‑8). MD-2 expression is required for cell surface localization of TLR4 and for optimal LPS-induced TLR4 signaling (8, 9). MD-2 also forms soluble disulfide-linked homo-oligomers which can interact with TLR4 (7). Through a domain separate from its TLR4-binding domain, MD-2 extracts LPS from circulating CD14-LPS complexes and carries the LPS into a ternary complex with TLR4 (10‑12). The interaction of MD-2/LPS with TLR4 induces receptor oligomerization and the triggering of an inflammatory response (13). Increased levels of plasma MD-2 in septic shock patients sensitizes MD-2 non-expressing epithelial cells to LPS and promotes widespread tissue inflammation (14).

Key Product Details
Species Reactivity
Validated:
Human
Cited:
Human, Mouse, Rat, Bacteria - Helicobacter pylori
Applications
Validated:
Immunohistochemistry, Neutralization, Flow Cytometry, Immunocytochemistry, CyTOF-ready
Cited:
Immunohistochemistry, Immunohistochemistry-Frozen, Western Blot, Neutralization, Flow Cytometry, Immunocytochemistry, Immunoprecipitation, Functional Assay, Neutralizing
Label
Unconjugated
Antibody Source
Polyclonal Goat IgG
Loading...
Product Specifications
Immunogen
Mouse myeloma cell line NS0-derived recombinant human TLR4
Glu24-Lys631
Accession # O00206
Glu24-Lys631
Accession # O00206
Specificity
Detects human TLR4 in direct ELISAs and Western blots. In direct ELISAs, approximately 10% cross-reactivity with recombinant mouse TLR4 is observed.
Clonality
Polyclonal
Host
Goat
Isotype
IgG
Endotoxin Level
<0.10 EU per 1 μg of the antibody by the LAL method.
Scientific Data Images for Human TLR4 Antibody
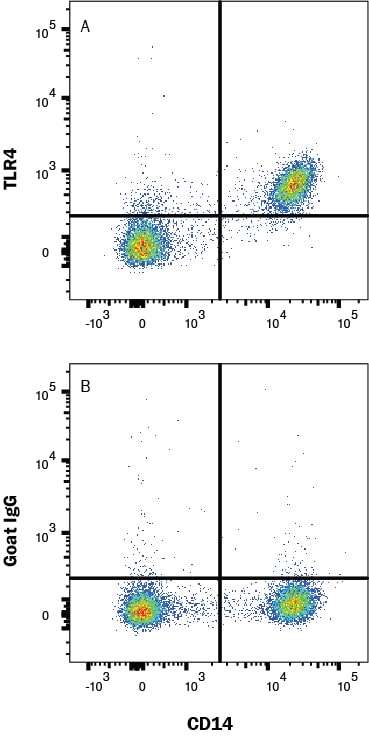
Detection of TLR4 in PBMC's by Flow Cytometry
Human peripheral blood mononuclear cells (PBMCs) were stained with Mouse Anti-Human CD14 PE‑conjugated Monoclonal Antibody (Catalog # FAB3832P) and either (A) Goat Anti-Human TLR4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1478) or (B) isotype control antibody (Catalog # AB-108-C) followed by Allophycocyanin-conjugated Anti-Goat IgG Secondary Antibody (Catalog # F0108). View our protocol for Staining Membrane-associated Proteins.TLR4 in Human Prostate.
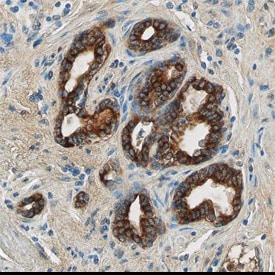
TLR4 was detected in immersion fixed paraffin-embedded sections of human prostate using Goat Anti-Human TLR4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1478) at 10 µg/mL overnight at 4 °C. Before incubation with the primary antibody tissue was subjected to heat-induced epitope retrieval using Antigen Retrieval Reagent-Basic (Catalog # CTS013). Tissue was stained using the Anti-Goat HRP-DAB Cell & Tissue Staining Kit (brown; Catalog # CTS008) and counterstained with hematoxylin (blue). View our protocol for Chromogenic IHC Staining of Paraffin-embedded Tissue Sections.TLR4 in Human PBMCs.
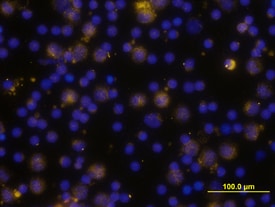
TLR4 was detected in immersion fixed human peripheral blood mononuclear cells (PBMCs) using Goat Anti-Human TLR4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1478) at 10 µg/mL for 3 hours at room temperature. Cells were stained using the NorthernLights™ 557-conjugated Anti-Goat IgG Secondary Antibody (yellow; Catalog # NL001) and counterstained with DAPI (blue). View our protocol for Fluorescent ICC Staining of Non-adherent Cells.IL‑8 Secretion Induced by LPS and Neutralization by Human TLR4 Antibody.
Lipopolysacharide (LPS) stimulates IL-8 secretion in the HEK293 human embryonic kidney cell line co-transfected with human TLR4 and MD-2, in a dose-dependent manner (orange line), as measured by the Human CXCL8/IL-8 Quantikine ELISA Kit (Catalog # D8000C). IL-8 secretion elicited by LPS (75 ng/mL) is neutralized (green line) by increasing concentrations of Goat Anti-Human TLR4 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1478). The ND50 is typically 1.5-7.5 µg/mL.Detection of Human TLR4 by Immunocytochemistry/ Immunofluorescence
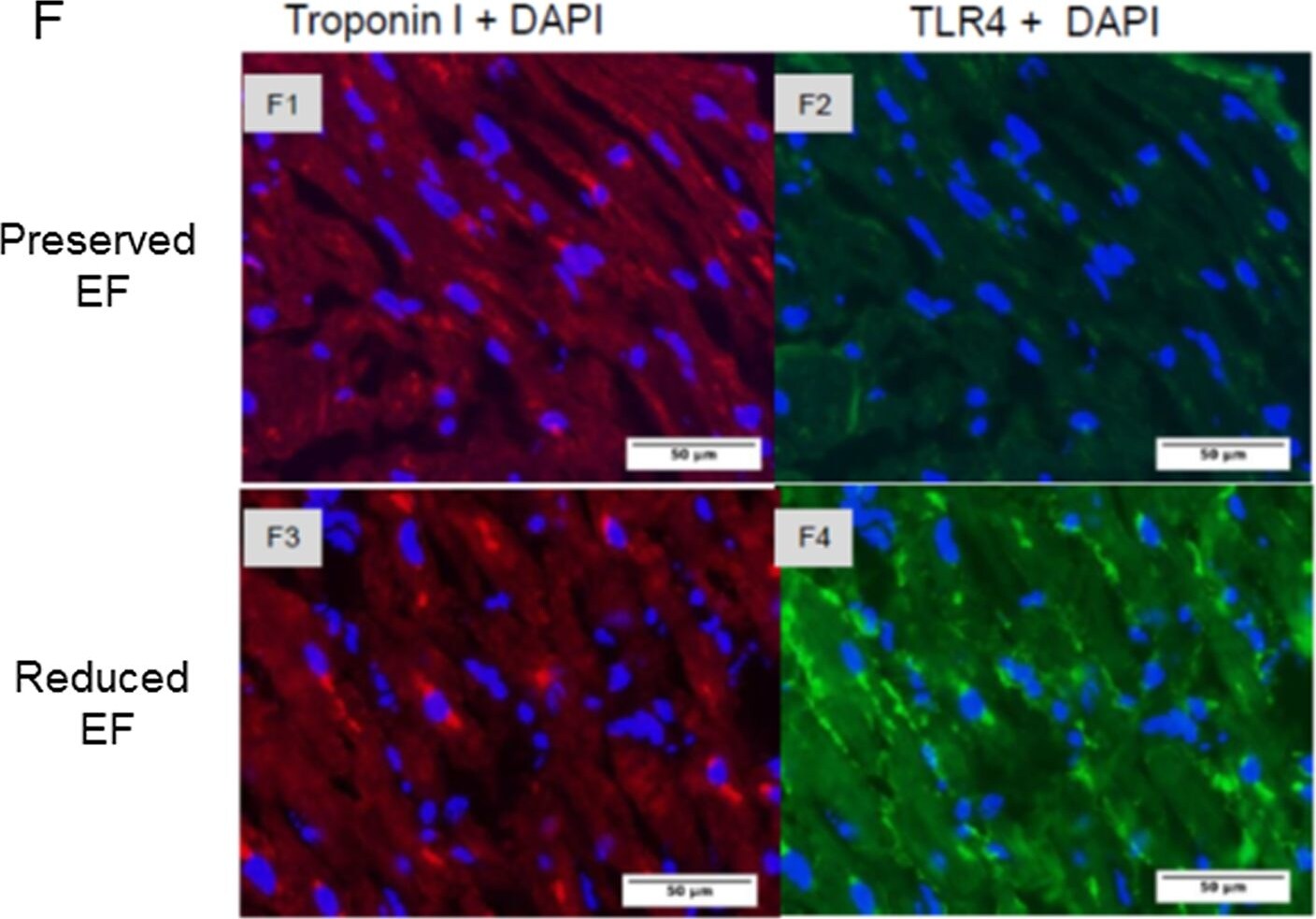
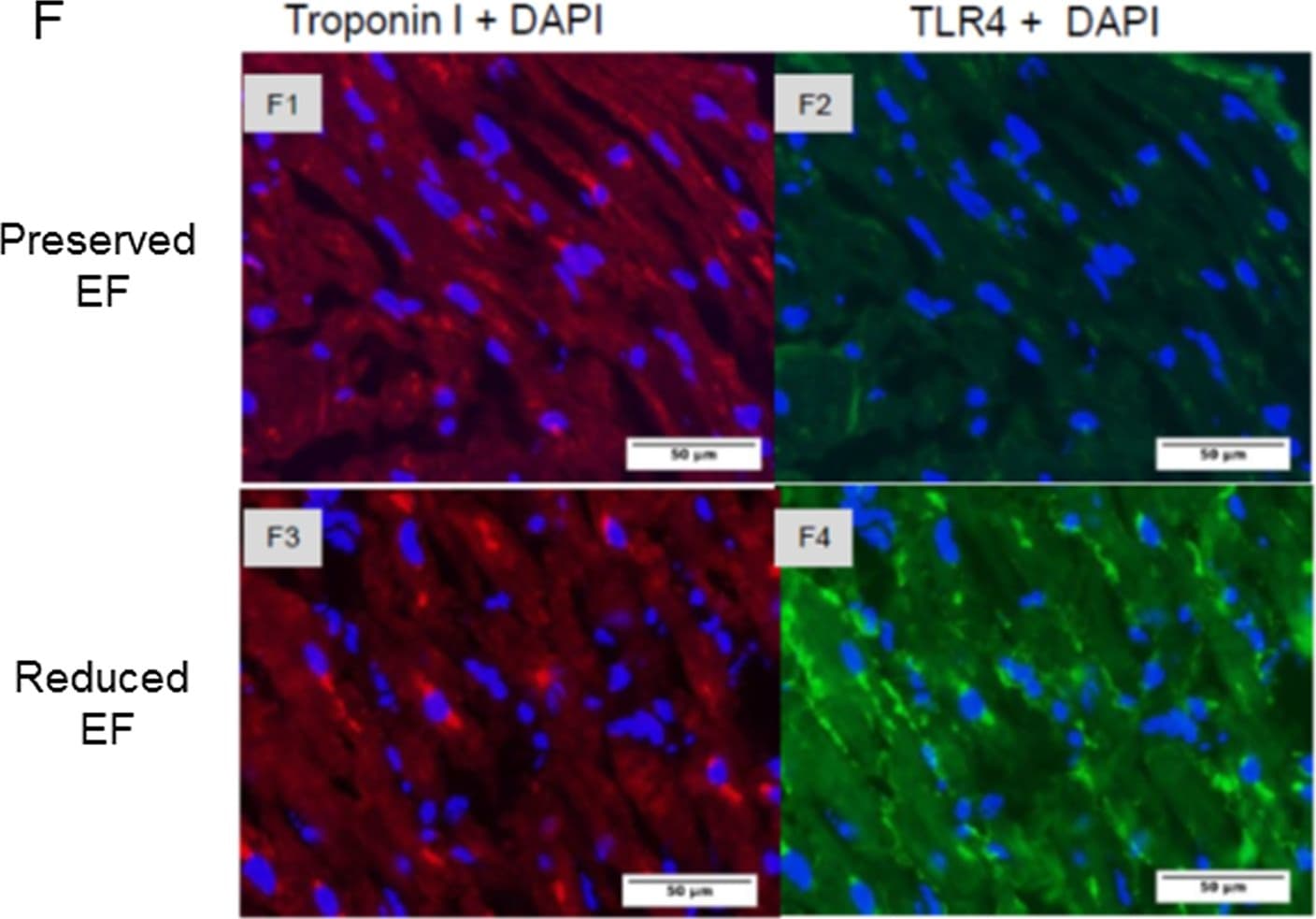
Gene expression of biomarkers of injury and immunostaining of TLR4 in patient auricles.(A-D) TLR4 and NOX4 are activated resulting in elevated TNF-alpha in the auricles. Auricles obtained during CABG surgery presented higher expression of TLR4 (P<0.03), BNP (P<0.05), NOX4 (P<0.03), and TNF-alpha (P = 0.135) in reduced versus ‘preserved EF’. (E) TLR2 expression was similar in both groups. (F1-F2) Representative photographs show double-immunostaining of Troponin I and TLR4 in ‘preserved EF’ auricle. (F3-F4) Representative photographs show double-immunostaining of troponin I and TLR4 in the ‘reduced EF’ auricle. TLR4 staining revealed an apparent upregulation in all ‘reduced EF’ patients examined compared to ‘preserved EF’ patients' tissue. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/26030867), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of Human TLR4 by Immunocytochemistry/ Immunofluorescence
Gene expression of biomarkers of injury and immunostaining of TLR4 in patient auricles.(A-D) TLR4 and NOX4 are activated resulting in elevated TNF-alpha in the auricles. Auricles obtained during CABG surgery presented higher expression of TLR4 (P<0.03), BNP (P<0.05), NOX4 (P<0.03), and TNF-alpha (P = 0.135) in reduced versus ‘preserved EF’. (E) TLR2 expression was similar in both groups. (F1-F2) Representative photographs show double-immunostaining of Troponin I and TLR4 in ‘preserved EF’ auricle. (F3-F4) Representative photographs show double-immunostaining of troponin I and TLR4 in the ‘reduced EF’ auricle. TLR4 staining revealed an apparent upregulation in all ‘reduced EF’ patients examined compared to ‘preserved EF’ patients' tissue. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/26030867), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
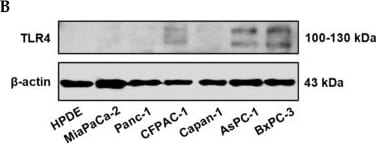
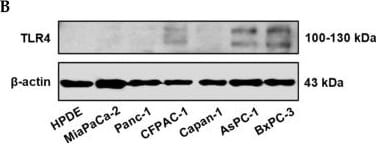
Positive TLR4 expression is detected in six different pancreatic cancer cell lines but not the HPDE normal pancreatic cell line. (A) mRNA expression levels of TLR4 quantified by RT-qPCR from the current study and retrieved from the Cancer Cell Line Encyclopedia (CCLE) Expression 22Q2 Public database. (B) Protein expression level of TLR4 assessed by Western blot analyses, the two bands of TLR4 appeared to be glycosylated (130 kDa) and deglycosulated (100 kDa) TLR4 [9,10]. (C) Relative mRNA levels of TLR1~9 in two representative pancreatic cancer cell lines. The mRNA expression levels were adjusted based on the two cell lines’ TLR4 data from (A). Data are represented as a mean ± SD from triplicates, ** p < 0.01, **** p < 0.0001 (indicating differences between Panc-1 and BxPC-3) were obtained from two-way ANOVA and post-hoc multiple comparisons with Bonferroni correction. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Immunocytochemistry/ Immunofluorescence
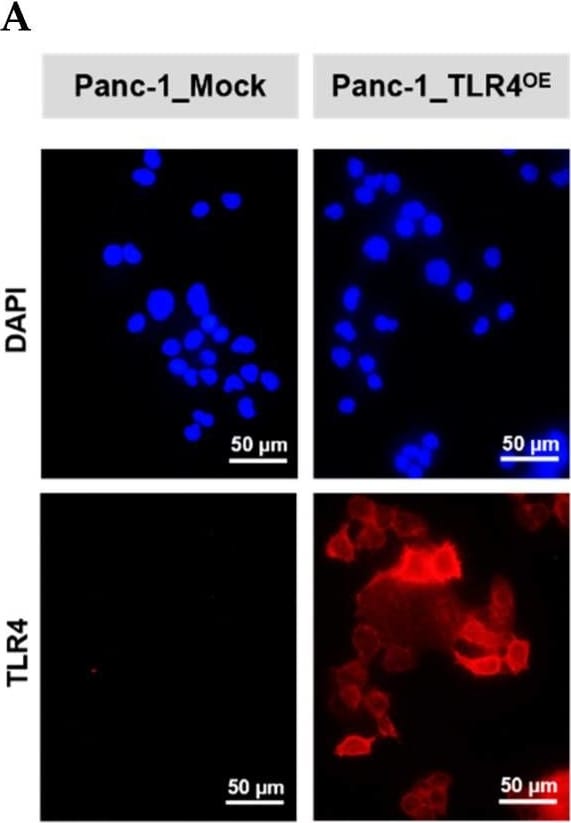
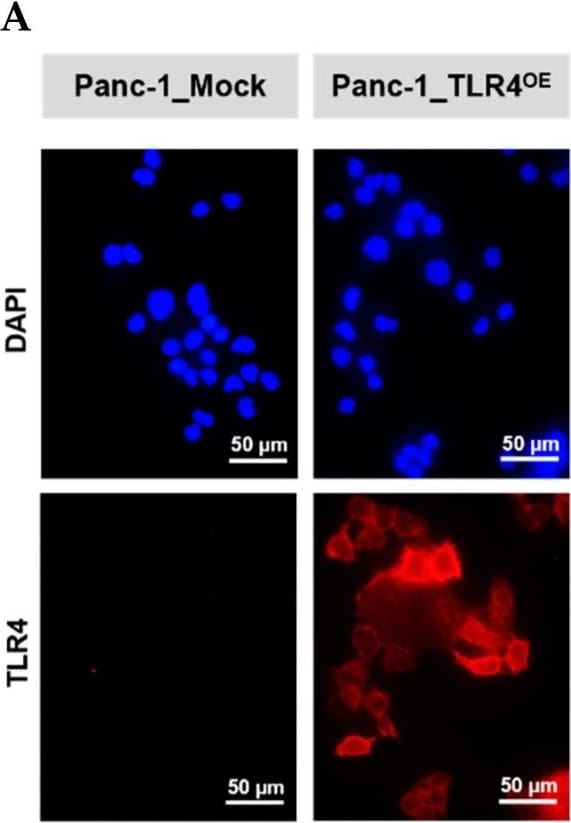
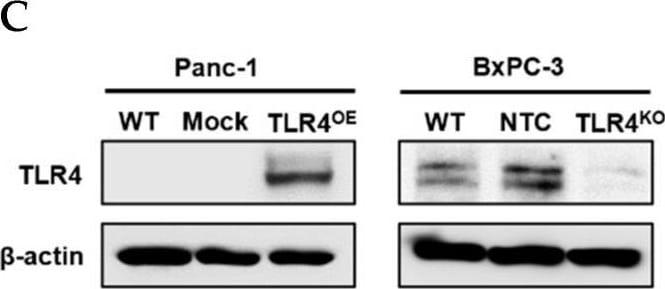
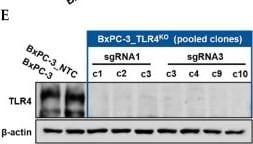
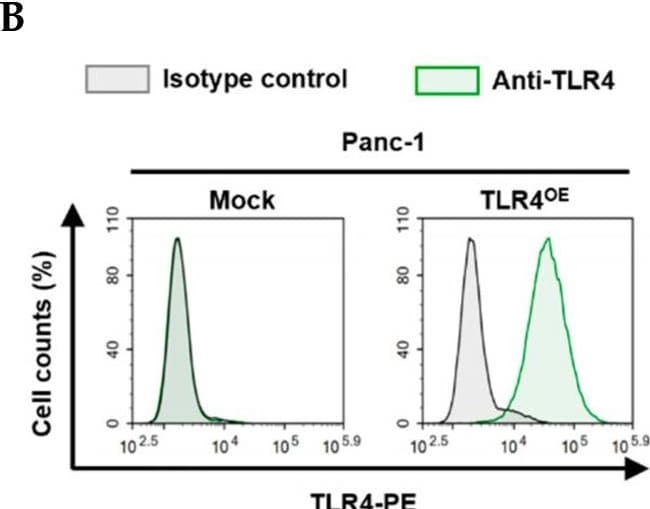
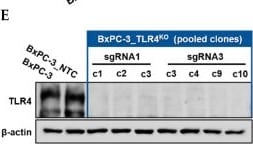
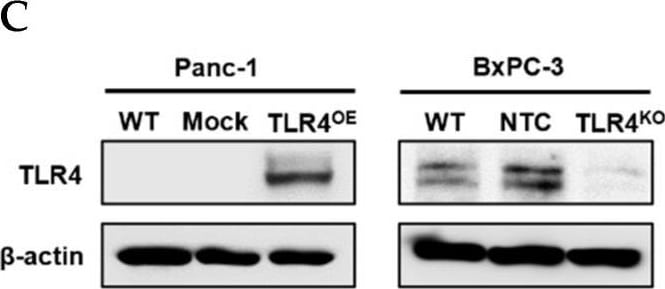
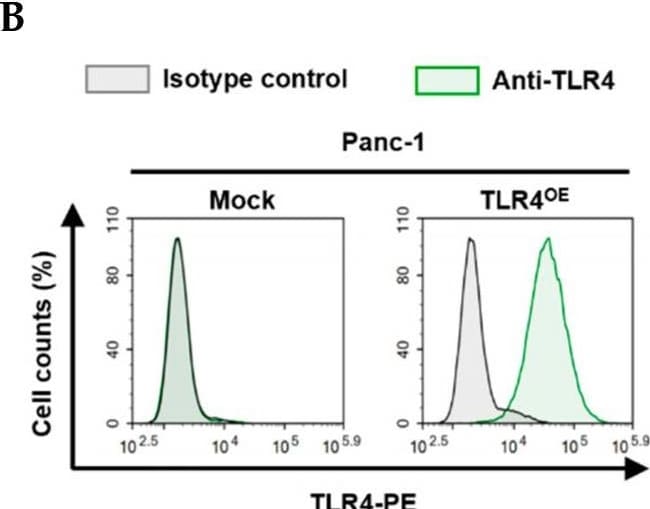
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Immunocytochemistry/ Immunofluorescence
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Flow Cytometry
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
Positive TLR4 expression is detected in six different pancreatic cancer cell lines but not the HPDE normal pancreatic cell line. (A) mRNA expression levels of TLR4 quantified by RT-qPCR from the current study and retrieved from the Cancer Cell Line Encyclopedia (CCLE) Expression 22Q2 Public database. (B) Protein expression level of TLR4 assessed by Western blot analyses, the two bands of TLR4 appeared to be glycosylated (130 kDa) and deglycosulated (100 kDa) TLR4 [9,10]. (C) Relative mRNA levels of TLR1~9 in two representative pancreatic cancer cell lines. The mRNA expression levels were adjusted based on the two cell lines’ TLR4 data from (A). Data are represented as a mean ± SD from triplicates, ** p < 0.01, **** p < 0.0001 (indicating differences between Panc-1 and BxPC-3) were obtained from two-way ANOVA and post-hoc multiple comparisons with Bonferroni correction. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Western Blot
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of TLR4 by Flow Cytometry
Successful generation of Panc-1 TLR4 overexpressed stable cell line (Panc-1_TLR4OE) and BxPC-3 TLR4 knockout stable cell line (BxPC-3_TLR4KO), and the impacts of TLR4 and PAUF expression on each other. Successful overexpression of TLR4 in Panc-1_TLR4OE cell line was confirmed by (A) immunofluorescence, (B) flow cytometry, and (C) Western blot (SDS-PAGE gel: 10%). Successful knockout of TLR4 by CRISPR/Cas9 was confirmed in BxPC-3_TLR4KO cells by (D) Cas9 mRNA expression and (E) Western blot (SDS-PAGE gel: 8%) against TLR4 in seven single clones with loss-of-function TLR4 mutations, which were pooled to form BxPC-3_TLR4KO cells. And the knockout of TLR4 in the pooled cells was confirmed by Western blot and shown in (C). (F) The correlation of TLR4 and PAUF mRNA expression was analyzed using CCLE expression 22Q2 public data by Pearson correlation. (G) PAUF protein concentration in the four cell lines analyzed by sandwich ELISA. (H) Impacts of rPAUF (0, 0.1, 1, and 3 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells. (I) Impacts of lipopolysaccharide (LPS, 0, 1, 5, and 10 μg/mL) on TLR4 mRNA expression in Panc-1 and BxPC-3 cells (LPS was used here as a positive control of PAUF). The dose-dependency of TLR4 mRNA expression on rPAUF/LPS concentration was tested by Jonckheere-Terpstra test, after a significant multiple comparisons test (* p < 0.05, compared to control, obtained from one-way ANOVA and post-hoc multiple comparisons with Dunnett correction). All data are presented as mean ± SD from triplicate independent experiments. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/36232715), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human TLR4 Antibody
Application
Recommended Usage
CyTOF-ready
Ready to be labeled using established conjugation methods. No BSA or other carrier proteins that could interfere with conjugation.
Flow Cytometry
2.5 µg/106 cells
Sample: Human peripheral blood monocytes
Sample: Human peripheral blood monocytes
Immunocytochemistry
5-15 µg/mL
Sample: Immersion fixed human peripheral blood mononuclear cells (PBMCs)
Sample: Immersion fixed human peripheral blood mononuclear cells (PBMCs)
Immunohistochemistry
5-15 µg/mL
Sample: Immersion fixed paraffin-embedded sections of human prostate cancer tissue subjected to Antigen Retrieval Reagent-Basic (Catalog # CTS013)
Sample: Immersion fixed paraffin-embedded sections of human prostate cancer tissue subjected to Antigen Retrieval Reagent-Basic (Catalog # CTS013)
Neutralization
Measured by its ability to neutralize LPS-induced IL‑8 secretion in the HEK293 human embryonic kidney cell line co-transfected with human TLR4 and MD-2. The Neutralization Dose (ND50) is typically 1.5-7.5 µg/mL in the presence of 75 ng/mL Lipopolysacharide (LPS).
Reviewed Applications
Read 1 review rated 5 using AF1478 in the following applications:
Flow Cytometry Panel Builder
Bio-Techne Knows Flow Cytometry
Save time and reduce costly mistakes by quickly finding compatible reagents using the Panel Builder Tool.
Advanced Features
- Spectra Viewer - Custom analysis of spectra from multiple fluorochromes
- Spillover Popups - Visualize the spectra of individual fluorochromes
- Antigen Density Selector - Match fluorochrome brightness with antigen density
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. See Certificate of Analysis for details.
*Small pack size (-SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
*Small pack size (-SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: TLR4
References
- Ohnishi, T. et al. (2003) Clin. Diagn. Lab. Immouol. 10:405.
- Gangloff, M. and N.J. Gay (2004) Trends Biochem. Sci. 29:294.
- Palsson-McDermott, E.M. and L.A. O’Neill (2004) Immunology 113:153.
- Miyake, K. (2004) Semin. Immunol. 16:11.
- Medzhitov, R. et al. (1997) Nature 388:394.
- Shimazu, R. et al. (1999) J. Exp. Med. 189:1777.
- Visintin, A. et al. (2001) Proc. Natl. Acad. Sci. USA 98:12156.
- Akashi, S. et al. (2000) J. Immunol. 164:3471.
- Nagai, Y. et al. (2002) Nat. Immunol. 3:667.
- Re, F. and J.L. Strominger (2003) J. Immunol. 171:5272.
- Kennedy, M.N. et al. (2004) J. Biol. Chem. 279:34698.
- Gioannini, T.L. et al. (2004) Proc. Natl. Acad. Sci. USA 101:4186.
- Saitoh, S. et al. (2004) J. Endotoxin Res. 10:257.
- Pugin, J. et al. (2004) Blood 104:4071.
Long Name
Toll-like Receptor 4
Alternate Names
CD284
Gene Symbol
TLR4
UniProt
Additional TLR4 Products
Product Documents for Human TLR4 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human TLR4 Antibody
For research use only
Citations for Human TLR4 Antibody
Customer Reviews for Human TLR4 Antibody (1)
5 out of 5
1 Customer Rating
Have you used Human TLR4 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Customer Images
Showing
1
-
1 of
1 review
Showing All
Filter By:
-
Application: Immunocytochemistry/ImmunofluorescenceSample Tested: Cell Culture SamplesSpecies: HumanVerified Customer | Posted 07/06/2025
There are no reviews that match your criteria.
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- 7-Amino Actinomycin D (7-AAD) Cell Viability Flow Cytometry Protocol
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Extracellular Membrane Flow Cytometry Protocol
- Flow Cytometry Protocol for Cell Surface Markers
- Flow Cytometry Protocol for Staining Membrane Associated Proteins
- Flow Cytometry Staining Protocols
- Flow Cytometry Troubleshooting Guide
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- ICC Cell Smear Protocol for Suspension Cells
- ICC Immunocytochemistry Protocol Videos
- ICC for Adherent Cells
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunocytochemistry (ICC) Protocol
- Immunocytochemistry Troubleshooting
- Immunofluorescence of Organoids Embedded in Cultrex Basement Membrane Extract
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Intracellular Flow Cytometry Protocol Using Alcohol (Methanol)
- Intracellular Flow Cytometry Protocol Using Detergents
- Intracellular Nuclear Staining Flow Cytometry Protocol Using Detergents
- Intracellular Staining Flow Cytometry Protocol Using Alcohol Permeabilization
- Intracellular Staining Flow Cytometry Protocol Using Detergents to Permeabilize Cells
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Propidium Iodide Cell Viability Flow Cytometry Protocol
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Liperfluo
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Characterization of Human Th22 Cells
- Protocol for the Characterization of Human Th9 Cells
- Protocol for the Fluorescent ICC Staining of Cell Smears - Graphic
- Protocol for the Fluorescent ICC Staining of Cultured Cells on Coverslips - Graphic
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent ICC Staining of Cells on Coverslips
- Protocol for the Preparation and Fluorescent ICC Staining of Non-adherent Cells
- Protocol for the Preparation and Fluorescent ICC Staining of Stem Cells on Coverslips
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- Protocol for the Preparation of a Cell Smear for Non-adherent Cell ICC - Graphic
- Protocol: Annexin V and PI Staining by Flow Cytometry
- Protocol: Annexin V and PI Staining for Apoptosis by Flow Cytometry
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Fluorokine Flow Cytometry Kits
- Troubleshooting Guide: Immunohistochemistry
- View all Protocols, Troubleshooting, Illustrated assays and Webinars