MDM2 is a key regulator of p53 tumor suppressor protein activity and stability. MDM2 binds to and inhibits the transactivation domain of p53. In addition, MDM2 controls p53 stability by functioning as its E3 ligase in ubiquitination and by shuttling p53 from the nucleus to the cytoplasm for subsequent degradation. The importance of the p53/MDM2 relationship is underscored by the existence of an autoregulatory feedback loop whereby activated p53 transcriptionally up‑regulates the expression of its own inhibitor, MDM2.
Human/Mouse/Rat MDM2/HDM2 Antibody
R&D Systems | Catalog # AF1244

Key Product Details
Validated by
Biological Validation
Species Reactivity
Validated:
Human, Mouse, Rat
Cited:
Human, Mouse, Rat, Transgenic Mouse
Applications
Validated:
Immunohistochemistry, Western Blot
Cited:
Immunohistochemistry, Immunohistochemistry-Paraffin, Western Blot
Label
Unconjugated
Antibody Source
Polyclonal Rabbit IgG
Loading...
Product Specifications
Immunogen
E. coli-derived recombinant human MDM2
Asn3-Pro491
Accession # Q00987
Asn3-Pro491
Accession # Q00987
Specificity
Detects human, mouse, and rat MDM2.
Clonality
Polyclonal
Host
Rabbit
Isotype
IgG
Scientific Data Images for Human/Mouse/Rat MDM2/HDM2 Antibody
MDM2/HDM2 in Human Kidney.
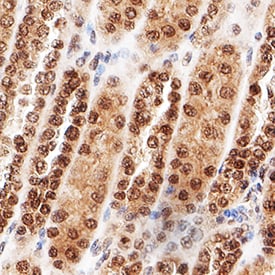
MDM2/HDM2 was detected in immersion fixed paraffin-embedded sections of human kidney using Rabbit Anti-Human/Mouse/Rat MDM2/HDM2 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1244) at 3 µg/mL for 1 hour at room temperature followed by incubation with the Anti-Rabbit IgG VisUCyte™ HRP Polymer Antibody (VC003). Before incubation with the primary antibody, tissue was subjected to heat-induced epitope retrieval using Antigen Retrieval Reagent-Basic (CTS013). Tissue was stained using DAB (brown) and counterstained with hematoxylin (blue). Specific staining was localized to cytoplasm and nuclei for epithelial cells in convoluted tubules. Staining was performed using our protocol for IHC Staining with VisUCyte HRP Polymer Detection Reagents.MDM2 in Mouse Kidney.
MDM2 was detected in perfusion fixed frozen sections of mouse kidney using Human/Mouse/Rat MDM2 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1244) at 5 µg/mL overnight at 4 °C. Tissue was stained using the Anti-Rabbit HRP-DAB Cell & Tissue Staining Kit (brown; Catalog # CTS005) and counterstained with hematoxylin (blue). View our protocol for Chromogenic IHC Staining of Frozen Tissue Sections.Detection of Human MDM2 by Western Blot.
Western blot shows lysates of U2OS human osteosarcoma cell line, MCF-7 human breast cancer cell line, and CEM human T-lymphoblastoid cell line untreated (-) or treated (+) with 5 µM LLnV for 1 hour. PVDF membrane was probed with 1 µg/mL of Human/Mouse/Rat MDM2 Antigen Affinity-purified Polyclonal Antibody (Catalog # AF1244), followed by HRP-conjugated Anti-Rabbit IgG Secondary Antibody (Catalog # HAF008). A specific band was detected for MDM2 at approximately 95 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 1.Detection of MDM2/HDM2 by Western Blot
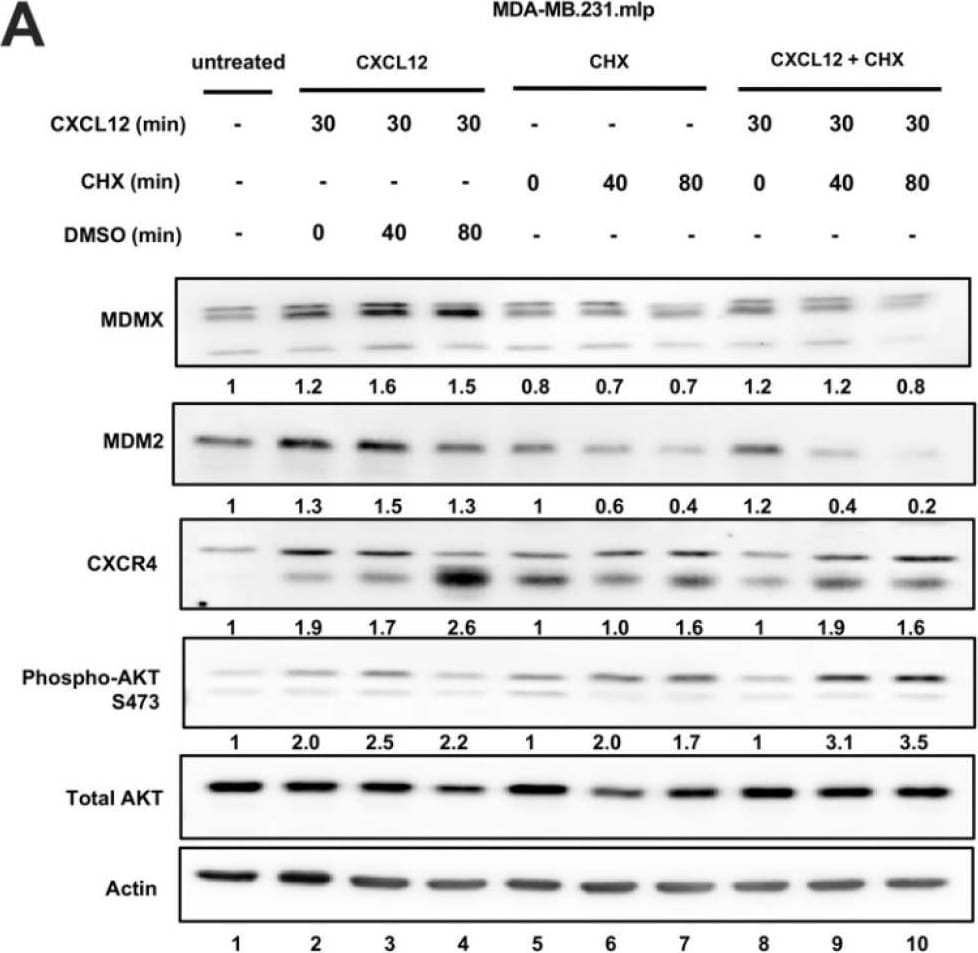
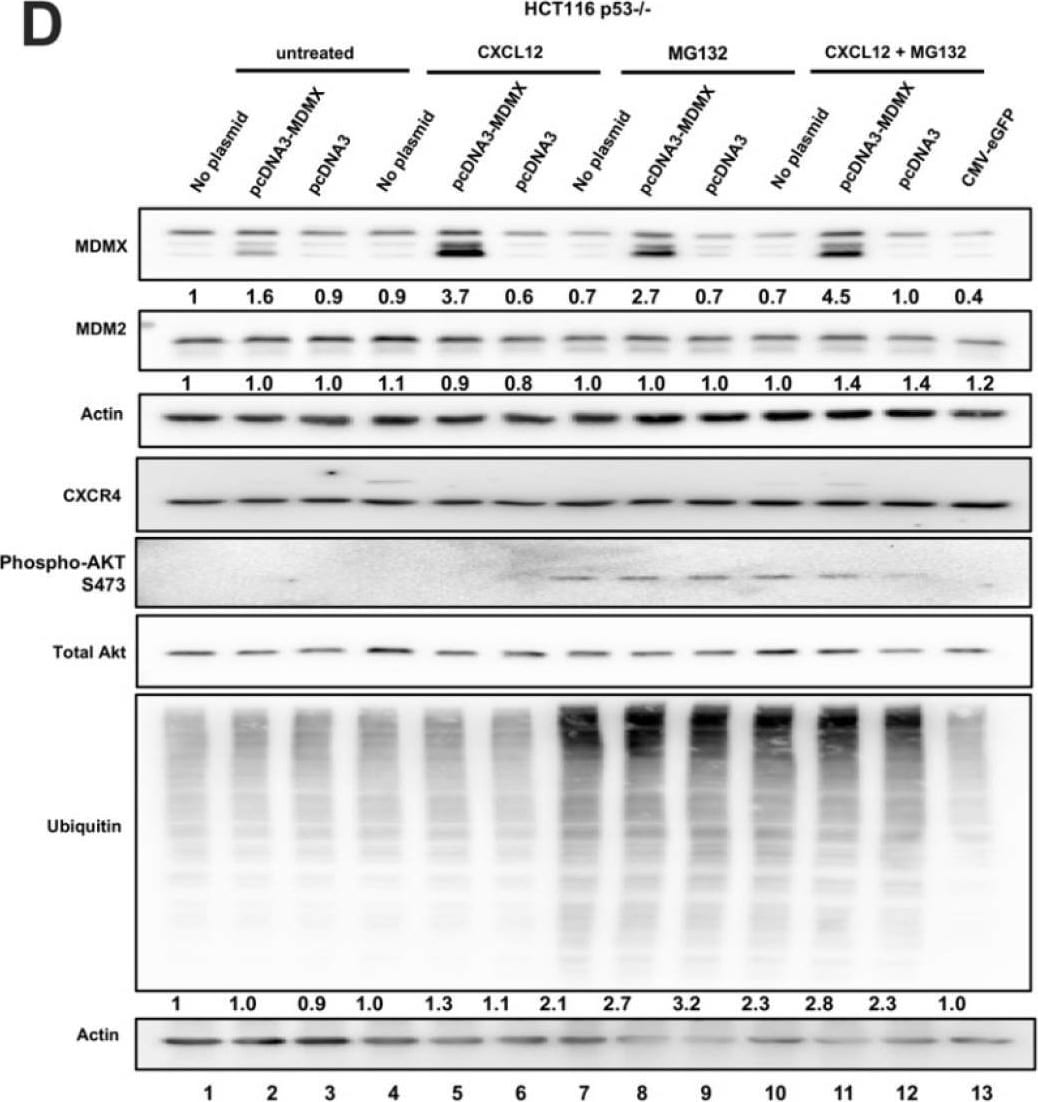
CXCL12 addition does not increase MDMX protein half-life following cycloheximide or MG132 treatment. (A) Immunoblot analysis of lysates from MDA-MB-231 cells treated with CXCL12 (50 ng/mL) for 30 min followed by cycloheximide (CHX) or DMSO for 40 or 80 min. Cells were harvested and lysed in CHAPS lysis buffer and subjected to immunoblotting to probe for MDM2 and MDMX. Actin was probed as a loading control. (B,C) Evaluation of Western blot band density was carried out using ImageJ and Prism software. Error bars represent SD. * p < 0.05, NS = non-significant. (D) HCT116 p53-/- cells were transfected with pcDNA3-MDMX for 24 h and then treated with CXCL12 (50 ng/mL) for 30 min followed by MG132 or DMSO for 40 or 80 min. Cells were harvested and lysed in CHAPS lysis buffer and subjected to immunoblotting to probe for MDM2, MDMX, and Ubiquitin. Actin was probed as a loading control. (E,F) Evaluation of Western blot band density was carried out using ImageJ and Prism 10 software. Error bars represent SD. * p < 0.05, NS = non-significant. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
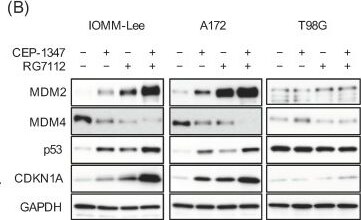
The MDM2 antagonist RG7112 concomitant with CEP-1347 effectively activates p53 and inhibits the growth of malignant brain tumor cells expressing wild-type p53. (A) IMR90 normal human fibroblasts, IOMM-Lee cells, and A172 cells treated with the indicated drugs (CEP-1347, 250 nM; RG7112, 500 nM) for 3 days were subjected to the trypan blue dye exclusion assay. (B) IOMM-Lee, A172, and T98G cells treated with the indicated drugs (CEP-1347, 250 nM; RG7112, 500 nM) for 2 days were subjected to a Western blot analysis. (C) Cells treated with the indicated drugs (CEP-1347, 250 nM; RG7112, 500 nM) for 3 days were cultured for another 5 days in the absence of any drugs for the colony formation assay. Representative images (left panels) and the number of colonies (right graphs) are shown. Values represent means + SD from triplicate samples of a representative experiment. * p < 0.05. † p < 0.05 vs. cells treated without any drugs. Similar results were obtained from more than two independent biological replicates. Original blot images can be found in Figure S4. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37686602), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
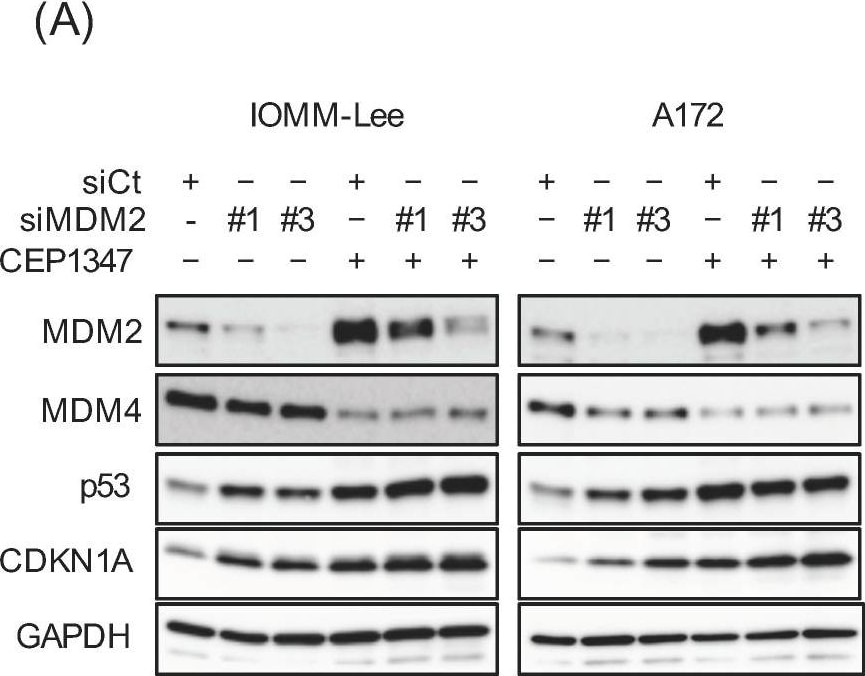
CEP-1347-induced MDM2 overexpression counteracts the CEP-1347-induced activation of p53. IOMM-Lee and A172 cells were transiently transfected with the indicated siRNA against MDM2 (#1 and #3) or with control siRNA (siCt). After being cultured for 1 day, cells treated without or with 250 nM CEP-1347 for 2 and 3 additional days were subjected to a Western blot analysis (A) and the trypan blue dye exclusion assay (B), respectively. Values represent the means + SD of triplicate samples of a representative experiment. * p < 0.05. † p < 0.05 vs. siCt-transfected cells treated without CEP-1347. Similar results were obtained from more than three independent biological replicates. Original blot images can be found in Figure S3. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37686602), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
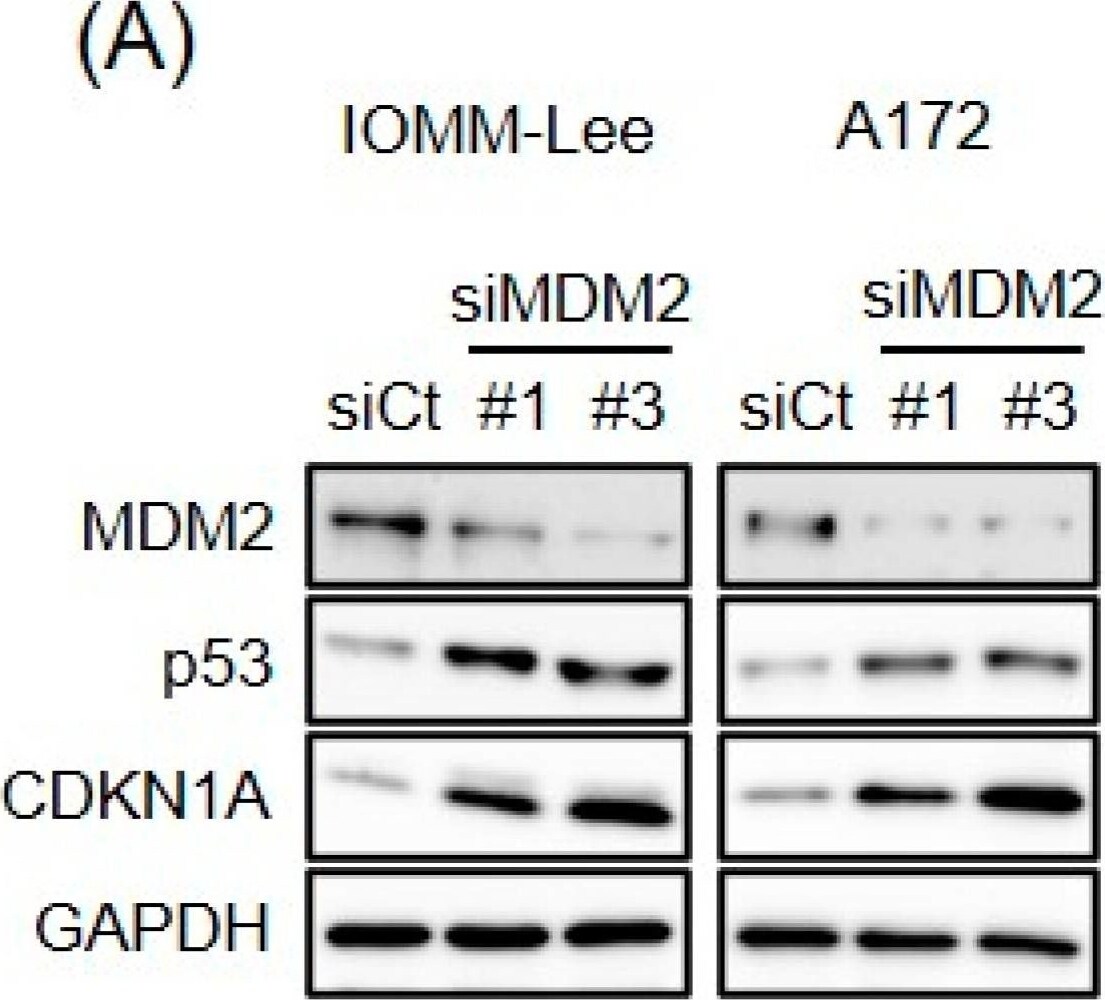
Endogenous expression of MDM2 contributes to p53 inactivation. (A) IOMM-Lee and A172 cells were transiently transfected with the indicated siRNA against MDM2 (#1 and #3) or with control siRNA (siCt). After being cultured for 3 days, cells were subjected to a Western blot analysis. (B) Cells were transiently transfected as in (A). After 3 days, transfected cells were subjected to the trypan blue dye exclusion assay to measure the number of viable cells. Values represent means + SD from triplicate samples of a representative experiment. * p < 0.05 vs. siCt-transfected cells. Similar results were obtained from more than two independent biological replicates. Original blot images can be found in Figure S2. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/37686602), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
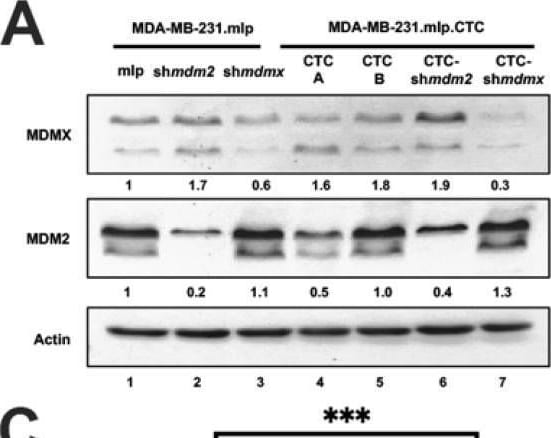
MDA-MB-231.mlp.CTC lines compared to MDA-MB-231.shmdm2.CTC and MDA-MB-231.shmdmx.CTC maintain increased migratory compacity but have reduced response to CXCL12. (A) Immunoblot of whole cell lysates from MDA-MB-231.mlp, MDA-MB-231.shmdm2, and MDA-MB-231.shmdmx (lanes 1–3) and MDA-MB-231.mlp.CTC A and B (lanes 4 and 5), MDA-MB-231.shmdm2.CTC (lane 6), and MDA-MB-231.shmdmx.CTC (lane 7) cell lines probed for MDMX (top) and MDM2 (middle). The loading control was actin (bottom) (B) Representative images of MDA-MB-231-derived CTC lines in wound healing assay. Black lines denote the borders of the scratch made. (C) Graph of % wound closure at 12 h time point. Error bars represent SD. *** p < 0.001, NS = nonsignificant. The p value was calculated using two-tailed unpaired t tests on Prism software (D) Immunoblot of whole cell lysates from MDA-MB-231.mlp.CTC lines treated with CXCL12 at a final concentration of 50 ng/mL for 30 and 60 min. (E) MDMX and MDM2 protein expression was compared using ImageJ quantitation relative to actin, respectively, as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
MDA-MB-231.mlp.CTC lines compared to MDA-MB-231.shmdm2.CTC and MDA-MB-231.shmdmx.CTC maintain increased migratory compacity but have reduced response to CXCL12. (A) Immunoblot of whole cell lysates from MDA-MB-231.mlp, MDA-MB-231.shmdm2, and MDA-MB-231.shmdmx (lanes 1–3) and MDA-MB-231.mlp.CTC A and B (lanes 4 and 5), MDA-MB-231.shmdm2.CTC (lane 6), and MDA-MB-231.shmdmx.CTC (lane 7) cell lines probed for MDMX (top) and MDM2 (middle). The loading control was actin (bottom) (B) Representative images of MDA-MB-231-derived CTC lines in wound healing assay. Black lines denote the borders of the scratch made. (C) Graph of % wound closure at 12 h time point. Error bars represent SD. *** p < 0.001, NS = nonsignificant. The p value was calculated using two-tailed unpaired t tests on Prism software (D) Immunoblot of whole cell lysates from MDA-MB-231.mlp.CTC lines treated with CXCL12 at a final concentration of 50 ng/mL for 30 and 60 min. (E) MDMX and MDM2 protein expression was compared using ImageJ quantitation relative to actin, respectively, as a loading control. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
CXCL12 addition does not increase MDMX protein half-life following cycloheximide or MG132 treatment. (A) Immunoblot analysis of lysates from MDA-MB-231 cells treated with CXCL12 (50 ng/mL) for 30 min followed by cycloheximide (CHX) or DMSO for 40 or 80 min. Cells were harvested and lysed in CHAPS lysis buffer and subjected to immunoblotting to probe for MDM2 and MDMX. Actin was probed as a loading control. (B,C) Evaluation of Western blot band density was carried out using ImageJ and Prism software. Error bars represent SD. * p < 0.05, NS = non-significant. (D) HCT116 p53-/- cells were transfected with pcDNA3-MDMX for 24 h and then treated with CXCL12 (50 ng/mL) for 30 min followed by MG132 or DMSO for 40 or 80 min. Cells were harvested and lysed in CHAPS lysis buffer and subjected to immunoblotting to probe for MDM2, MDMX, and Ubiquitin. Actin was probed as a loading control. (E,F) Evaluation of Western blot band density was carried out using ImageJ and Prism 10 software. Error bars represent SD. * p < 0.05, NS = non-significant. Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
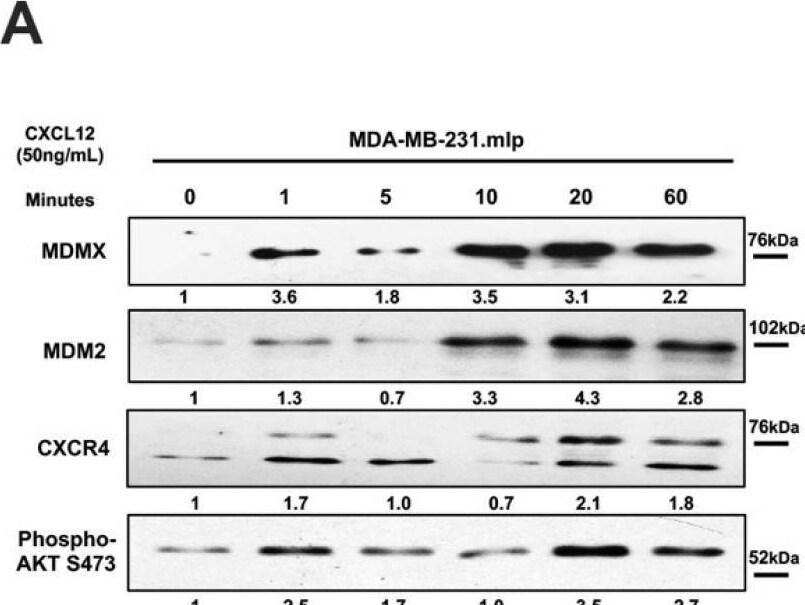
Chemokine CXCL12 addition to cell culture increases CXCR4, activates PI3K/AKT, and MDMX and MDM2 protein levels. (A) Panels show immunoblot analysis for MDMX, MDM2, CXCR4, and phospho-AKT S473, total AKT, and actin in MDA-MB-231.mlp cells after treatment for up to 60 min with CXCL12 at 50 ng/mL for 0, 1, 5, 10, 20, and 60 min, lanes 1–6. Total AKT and actin were used as loading controls. The proteins were derived from the same samples run on different gels/membranes at the same time and their molecular weights are shown. (B) Untreated cells and those following the 20 min treatments were compared for the MDMX and MDM2 protein levels evaluated (using actin as a normalizer control for loading) using ImageJ and graphs were created with Prism 10 software, with the untreated value set as 1 and the ratio reported for CXCL12-treated samples (20 min) used to report the fold change. (C) Untreated and 20 min treatments were compared for CXCR4 and phospho-AKT S473 protein levels quantified via ImageJ relative to total AKT as a loading control with the untreated value set as 1 and the ratio reported for CXCL12-treated samples (20 min) used to report the fold change. Images were analyzed using ImageJ and graphs were created with Prism software. Error bars represent SD. * p < 0.05, ** p < 0.01, NS = non-significant (N = 4 biological replicates). Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Detection of MDM2/HDM2 by Western Blot
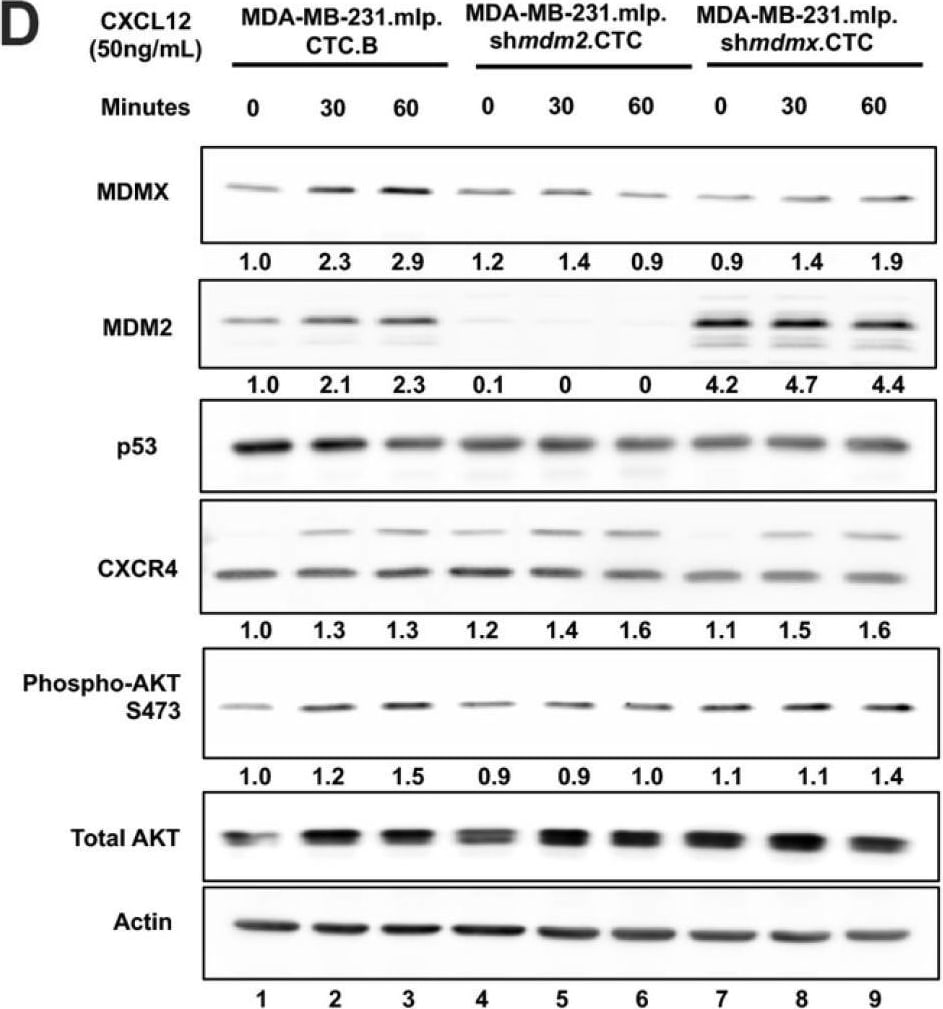
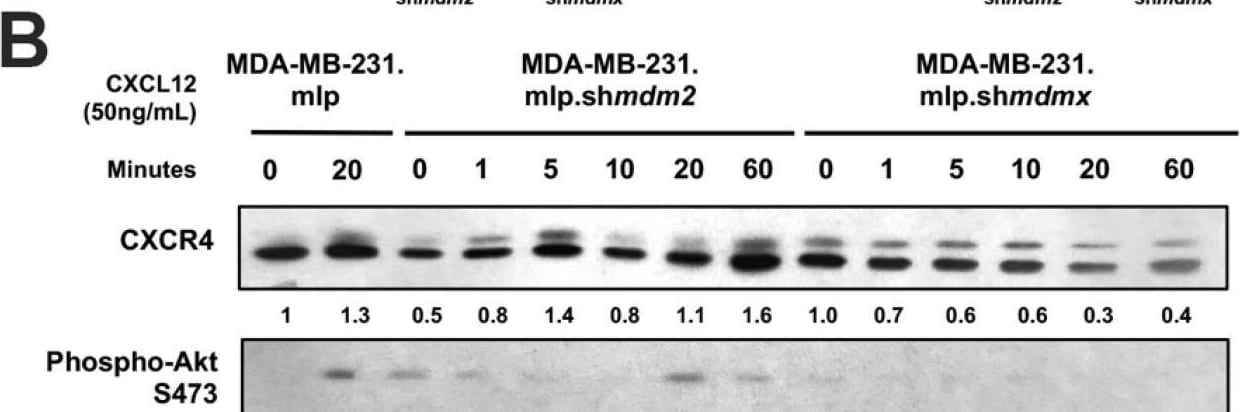
Knockdown of MDMX in MDA-MB-231 cells disrupts CXCL12 signaling to upregulate CXCR4, MDM2, and AKT activation. MDA-MB-231.mlp, MDA-MB-231.mlp.shmdm2, and MDA-MB-231.mlp.shmdmx cells treated in cell culture with the addition of CXCL12 at a final concentration of 50 ng/mL in cell culture for up to 60 min. (A) CXCR4 and phospho-AKT S473 protein levels were semi-quantified at 20 min via ImageJ relative to actin as a loading control. Protein level analysis was carried out using Western blot results using Image J and Prism software and densitometries were measured as a ratio relative to the actin band density. Fold change was calculated relative to protein levels in the untreated 231.mlp vector control cells. Error bars represent SD. * p < 0.05, NS = non-significant (N = 3 biological replicates). (B) Immunoblot analysis for CXCR4, phospho-AKT, MDMX, and MDM2 protein levels in MDA-MB-231 or knockdown cells after the addition of CXCL12. (C) MDMX and MDM2 protein levels were semi-quantified at 20 min post addition of CXCL12 to the cell culture. Protein levels were normalized to actin and fold change was calculated relative to untreated 231.mlp vector control cells. MDM2 or MDMX knockdown were confirmed for each respective cell line. Protein level analysis was carried out from Western blot results using Image J and Prism software and expression scores were normalized to actin. Error bars represent SD. * p < 0.05, NS = non-significant (N = 3 biological replicates). Image collected and cropped by CiteAb from the following open publication (https://pubmed.ncbi.nlm.nih.gov/39766093), licensed under a CC-BY license. Not internally tested by R&D Systems.Applications for Human/Mouse/Rat MDM2/HDM2 Antibody
Application
Recommended Usage
Immunohistochemistry
3-15 µg/mL
Sample: Immersion fixed paraffin-embedded sections of human kidney and perfusion fixed frozen sections of mouse kidney
Sample: Immersion fixed paraffin-embedded sections of human kidney and perfusion fixed frozen sections of mouse kidney
Western Blot
1 µg/mL
Sample: MCF‑7 human breast cancer cell line
Sample: MCF‑7 human breast cancer cell line
Formulation, Preparation, and Storage
Purification
Antigen Affinity-purified
Reconstitution
Reconstitute at 0.2 mg/mL in sterile PBS. For liquid material, refer to CoA for concentration.
Loading...
Formulation
Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. *Small pack size (SP) is supplied either lyophilized or as a 0.2 µm filtered solution in PBS.
Shipping
Lyophilized product is shipped at ambient temperature. Liquid small pack size (-SP) is shipped with polar packs. Upon receipt, store immediately at the temperature recommended below.
Stability & Storage
Use a manual defrost freezer and avoid repeated freeze-thaw cycles.
- 12 months from date of receipt, -20 to -70 °C as supplied.
- 1 month, 2 to 8 °C under sterile conditions after reconstitution.
- 6 months, -20 to -70 °C under sterile conditions after reconstitution.
Calculators
Background: MDM2/HDM2
Long Name
Double Minute 2 Protein
Alternate Names
HDM2
Gene Symbol
MDM2
UniProt
Additional MDM2/HDM2 Products
Product Documents for Human/Mouse/Rat MDM2/HDM2 Antibody
Certificate of Analysis
To download a Certificate of Analysis, please enter a lot or batch number in the search box below.
Note: Certificate of Analysis not available for kit components.
Product Specific Notices for Human/Mouse/Rat MDM2/HDM2 Antibody
For research use only
Related Research Areas
Citations for Human/Mouse/Rat MDM2/HDM2 Antibody
Customer Reviews for Human/Mouse/Rat MDM2/HDM2 Antibody
There are currently no reviews for this product. Be the first to review Human/Mouse/Rat MDM2/HDM2 Antibody and earn rewards!
Have you used Human/Mouse/Rat MDM2/HDM2 Antibody?
Submit a review and receive an Amazon gift card!
$25/€18/£15/$25CAN/¥2500 Yen for a review with an image
$10/€7/£6/$10CAN/¥1110 Yen for a review without an image
Submit a review
Protocols
Find general support by application which include: protocols, troubleshooting, illustrated assays, videos and webinars.
- Antigen Retrieval Protocol (PIER)
- Antigen Retrieval for Frozen Sections Protocol
- Appropriate Fixation of IHC/ICC Samples
- Cellular Response to Hypoxia Protocols
- Chromogenic IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Chromogenic Immunohistochemistry Staining of Frozen Tissue
- ClariTSA™ Fluorophore Kits
- Detection & Visualization of Antibody Binding
- Fluorescent IHC Staining of Frozen Tissue Protocol
- Graphic Protocol for Heat-induced Epitope Retrieval
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Graphic Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Graphic Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- IHC Sample Preparation (Frozen sections vs Paraffin)
- Immunofluorescent IHC Staining of Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Protocol
- Immunohistochemistry (IHC) and Immunocytochemistry (ICC) Protocols
- Immunohistochemistry Frozen Troubleshooting
- Immunohistochemistry Paraffin Troubleshooting
- Preparing Samples for IHC/ICC Experiments
- Preventing Non-Specific Staining (Non-Specific Binding)
- Primary Antibody Selection & Optimization
- Protocol for Heat-Induced Epitope Retrieval (HIER)
- Protocol for Making a 4% Formaldehyde Solution in PBS
- Protocol for VisUCyte™ HRP Polymer Detection Reagent
- Protocol for the Preparation & Fixation of Cells on Coverslips
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Frozen Tissue Sections - Graphic
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation and Chromogenic IHC Staining of Paraffin-embedded Tissue Sections - Graphic
- Protocol for the Preparation and Fluorescent IHC Staining of Frozen Tissue Sections
- Protocol for the Preparation and Fluorescent IHC Staining of Paraffin-embedded Tissue Sections
- Protocol for the Preparation of Gelatin-coated Slides for Histological Tissue Sections
- R&D Systems Quality Control Western Blot Protocol
- TUNEL and Active Caspase-3 Detection by IHC/ICC Protocol
- The Importance of IHC/ICC Controls
- Troubleshooting Guide: Immunohistochemistry
- Troubleshooting Guide: Western Blot Figures
- Western Blot Conditions
- Western Blot Protocol
- Western Blot Protocol for Cell Lysates
- Western Blot Troubleshooting
- Western Blot Troubleshooting Guide
- View all Protocols, Troubleshooting, Illustrated assays and Webinars
Loading...
Associated Pathways
 Ubiquitination Cascade Pathway
Ubiquitination Cascade Pathway